Abstract
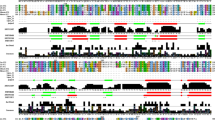
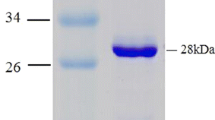
A thl gene encoding the thiolase (EC 2.3.1.9) of Clostridium pasteurianum was cloned by thermal asymmetric interlaced (TAIL) PCR. It consists of 1179 bp with 36.8% GC content and encodes 392 amino acids with a deduced molecular mass of 40,954 Da and shows 77% identity and 88% similarity to that of Clostridium tetani E88 and should be classified as a biosynthetic thiolase with three conserved residues Cys89, Cys382 and His352. The gene was over-expressed in Escherichia coli and the thiolase was purified with Ni-NTA agarose column to homogeneity. The K m of this thiolase for acetoacetyl-CoA is 0.13 mM with 0.06 mM CoASH at pH 8.2, 25°C and a V max value of 46 µmol min−1 mg−1.
Similar content being viewed by others
References
Berndt H, Schlegel HG (1975) Kinetics and properties of beta-ketothiolase from Clostridium pasteurianum. Arch Microbiol 103:21–30
Biebl H (2001) Fermentation of glycerol by Clostridium pasteurianum – batch and continuous culture studies. J Ind Microbiol Biotechnol 27:18–26
Durre P, Fischer RJ, Kuhn A, Lorenz K, Schreiber W, Sturzenhofecker B, Ullmann S, Winzer K, Sauer U (1995) Solventogenic enzymes of Clostridium acetobutylicum: catalytic properties, genetic organization, and transcriptional regulation. FEMS Microbiol Rev 17:251–262
Jenkins LS, Nunn WD (1987) Genetic and molecular characterization of the genes involved in short-chain fatty acid degradation in Escherichia coli: the ato system. J Bacteriol 169:42–52
Kursula P, Ojala J, Lambeir AM, Wierenga RK (2002) The catalytic cycle of biosynthetic thiolase: a conformational journey of an acetyl group through four binding modes and two oxyanion holes. Biochemistry 41:15543–15556
Leaf TA, Srienc F (1998) Metabolic modeling of polyhydroxybutyrate biosynthesis. Biotechnol Bioeng 57:557–570
Li JL, Burris RH (1983) Influence of pN2 and pD2 on HD formation by various nitrogenases. Biochemistry 22:4472–4480
Liu YG, Whittier RF (1995) Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 25:674–681
Mazzei Y, Negrel R, Ailhaud G (1970) Purification and some properties of thiolase from Escherichia coli. Biochim Biophys Acta 220:129–131
Modis Y, Wierenga RK (1999) A biosynthetic thiolase in complex with a reaction intermediate: the crystal structure provides new insights into the catalytic mechanism. Structure Fold Des 7:1279–1290
Modis Y, Wierenga RK (2000) Crystallographic analysis of the reaction pathway of Zoogloea ramigera biosynthetic thiolase. J Mol Biol 297:1171–1182
Palmer MA, Differding E, Gamboni R, Williams SF, Peoples OP, Walsh CT, Sinskey AJ, Masamune S (1991) Biosynthetic thiolase from Zoogloea ramigera. Evidence for a mechanism involving Cys-378 as the active site base. J Biol Chem 266:8369–8375
Peoples OP, Sinskey AJ (1989) Poly-beta-hydroxybutyrate biosynthesis in Alcaligenes eutrophus H16. Characterization of the genes encoding beta-ketothiolase and acetoacetyl-CoA reductase. J Biol Chem 264:15293–15297
Petersen DJ, Bennett GN (1991) Cloning of the Clostridium acetobutylicum ATCC 824 acetyl coenzyme A acetyltransferase (thiolase; EC 2.3.1.9) gene. Appl Environ Microbiol 57:2735–2741
Stewart PR, Rudney H (1966) The biosynthesis of beta-hydroxy-beta-methylglutaryl coenzyme A in yeast. 3. Purification and properties of the condensing enzyme thiolase system. J Biol Chem 241:1212–1221
Thompson S, Mayerl F, Peoples OP, Masamune S, Sinskey AJ, Walsh CT (1989) Mechanistic studies on beta-ketoacyl thiolase from Zoogloea ramigera: identification of the active-site nucleophile as Cys89, its mutation to Ser89, and kinetic and thermodynamic characterization of wild-type and mutant enzymes. Biochemistry 28:5735–5742
Williams SF, Palmer MA, Peoples OP, Walsh CT, Sinskey AJ, Masamune S (1992) Biosynthetic thiolase from Zoogloea ramigera. Mutagenesis of the putative active-site base Cys-378 to Ser-378 changes the partitioning of the acetyl S-enzyme intermediate. J Biol Chem 267:16041–16043
Winzer K, Lorenz K, Zickner B, Durre P (2000) Differential regulation of two thiolase genes from Clostridium acetobutylicum DSM 792. J Mol Microbiol Biotechnol 2:531–541
Zeng J, Li D (2004) Expression and purification of His-tagged rat mitochondrial 3-ketoacyl-CoA thiolase wild-type and His352 mutant proteins. Protein Expr Purif 35:320–326
Acknowledgements
We thank Dr. Jiesheng Tian, Xiaoyu Zhou, Jiangning Liu, Nining Hong, Miao Pan and Feng Guan for helpful discussions. Thanks also to Dr. Yaoping Zhang for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Meng, Y., Li, J. Cloning, Expression and Characterization of a Thiolase Gene from Clostridium pasteurianum . Biotechnol Lett 28, 1227–1232 (2006). https://doi.org/10.1007/s10529-006-9089-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-006-9089-4