Abstract
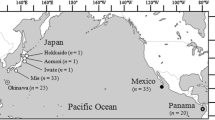
The demographic history and population genetic structure of the blackfin flounder (Glyptocephalus stelleri) along coastal regions of Japan were investigated. Genetic variation in DNA sequences was examined from the first hypervariable region of the mitochondrial DNA control region. A high level of haplotypic diversity (h = 0.99 ± 0.004) was detected, indicating a high level of intrapopulation genetic diversity. The starburst structure of the minimum spanning tree suggested a very recent origin for most haplotypes. The demographic history of G. stelleri was examined using neutrality tests and mismatch distribution analysis, which also indicated a Pleistocene population expansion at about 124,100–413,400 years ago. Hierarchical molecular variance analysis and conventional population Fst comparisons revealed no significant genetic differentiation throughout the range examined.



Similar content being viewed by others
References
Aris-Brosou S, Excoffier L (1996) The impact of population expansion and mutation rate heterogeneity on DNA sequence polymorphism. Mol Biol Evol 13:494–504
Avise JC (1998) The history and preview of phylogeography: a personal reflection. Mol Ecol 7:371–379
Avise J (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Benzie JA (1999) Genetic structure of coral reef organisms ghosts of dispersal past. Am Zool 39:131–145
Chang Y, Huang F, Lo T (1994) The complete nucleotide sequence and gene organization of carp (Cyprinus carpio) mitochondrial DNA gene. J Mol Evol 38:138–155
Excoffier L (1993) MINSPNET: a programme for producing minimum spanning network. Department of Anthropology and Ecology, University of Geneva, Switzerland
Excoffier L (2004) Patterns of DNA sequence diversity and genetic structure after a range expansion: lessons from the infinite-island model. Mol Ecol 13:853–864
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Fauvelot C, Planes S (2002) Understanding origins of present-day genetic structure in marine fish: biologically or historically driven patterns? Mar Biol 141:773–788
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Fujita S (1965) Early development and rearing of two common flatfishes, Eopsetta grigorjewi (Herzenstein) and Tanakius kitaharai (Jordan et starks). Bull Japan Soc Sci Fish 31:258–262
Grant SW, Bowen BW (1998) Shallow population histories in deep evolutionary lineages of marine fishes: insight from sardines and anchovies and lessons for conservation. J Hered 89:415–426
Hamai I, Ishito Y (1958) Distribution of the flatfish, Glyptocephalus stelleri (Schmidt) along the Pacific coast of Tohoku and Hokkaido regions. Bull Tohoku Reg Fish Res Lab 11:1–37
Hansen HJ, Nielsen EE, Grønkjaer P (2007) Evolutionary mechanisms shaping the genetic population structure of marine fishes; lessons from the European flounder (Platichthys flesus L.). Mol Ecol 16:3104–3118
Harpending RC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 66:591–600
Harpending H, Rogers A (2000) Genetic perspectives on human origins and differentiation. Ann Rev Genomics Hum Genet 1:361–385
Hashimoto R (1953) Studies on the age of Glyptocephalus stelleri (Schmidt). Bull Tohoku Reg Fish Res Lab 2:49–55
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Imbrie J, Boyle EA, Clemens SC, Duffy A, Howard WR, Kukla G, Kutzbach J, Martinson DG, McIntyre A, Mix AC, Molfino B, Morley JJ, Peterson LC, Pisias NG, Prell WL, Raymo ME, Shackleton NJ, Toggweiler JR (1992) On the structure and origin of major glaciation cycles, 1. Linear responses to Milankovitch forcing. Paleoceanography 7:701–738
Imron JB, Hale P, Degnan MB, Degnan MS (2007) Pleistocene isolation and recent gene flow in Haliotis asinina, an Indo-Pacific vetigastropod with limited dispersal capacity. Mol Ecol 16:289–304
Ishida R, Kitakata M (1953) Studies on the age determination of flatfish in Hokkaido Glyptocephalus stelleri (Schmidt). Bull Hokkaido Reg Fish Res Lab 8:63–84
Kitamura A, Matsui H, Oda M (1999) Change in the thickness of the warm Tsushima current at the initiation of its flow into the Sea of Japan. Palaeogeogr Palaeoclimatol Palaeoecol 152:305–318
Kitamura A, Takano O, Takata H, Omote H (2001) Late Pliocene-early Pleistocene paleoceanographic evolution of the Japan Sea. Palaeogeogr Palaeoclimatol Palaeoecol 172:81–98
Kocher TD, Thomas WK, Meyer A, Edwards SV (1989) Dynamic of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci USA 86:6196–6200
Liu JX, Gao TX, Yokogawa KJ, Zhang YP (2006a) Differential population structuring and demographic history of two closely related fish species, Japanese sea bass (Lateolabrax japonicus) and spotted sea bass (Lateolabrax maculatus) in Northwestern Pacific. Mol Phylogenet Evol 39:799–811
Liu JX, Gao TX, Zhuang ZM, Jin XS, Yokogawa KJ, Zhang YP (2006b) Late Pleistocene divergence and subsequent population expansion of two closely related fish species, Japanese anchovy (Engraulis japonicus) and Australian anchovy (Engraulis australis). Mol Phylogenet Evol 40:712–723
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nielsen EE, Hansen MM, Ruzzante DE, Meldrup D, Gronkjaer P (2003) Evidence of a hybrid-zone in Atlantic cod (Gadus morhua) in the Baltic and the Danish Belt Sea revealed by individual admixture analysis. Mol Ecol 12:1497–1508
Okiyama M (1963) Larvae and young of the witch flounder, Glyptocephalus stelleri (Schmidt) at metamorphosis stages. Bull Jap Sea Reg Fish Res Lab 11:101–108
Ōuchi A (1954) Breeding of some species of flatfish in Japan Sea. Ann Rep Japan Sea Reg Fish Res Lab 1:17–26
Planes S, Doherty PJ, Bernardi G (2001) Strong genetic divergence among populations of a marine fish with limited dispersal, Acanthochromis polyacanthus, within the great barrier reef and the coral sea. Evolution 55:2263–2273
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Ray N, Currant M, Excoffier L (2003) Intra-deme molecular diversity in spatially expanding populations. Mol Biol Evol 20(1):76–86
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Robards MD, Willson MF, Armstrong RH, Piatt JF (1999) Sand lance: A review of biology and predator relations and annotated bibliography. Exxon Valdez oil spill restoration project 99346
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic difference. Mol Biol Evol 9:552–569
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Saitou N, Nei M (1987) The neighbor-joining methods: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schmidt PJ (1904) Fishes of the eastern seas of the Russian Empire. Scientific results of the Korea–Sakhalin Expedition of the Emperor Russian Geographical Society 1900–1901. St. Petersburg Fish East Seas Russian Empire, St. Petersburg
Schneider S, Excoffier L (1999) Estimation of past demographic parameters from the distribution of pairwise differences when the mutation rates vary among sites: application to human mitochondrial DNA. Genetics 152:1079–1089
Schneider S, Roessli D, Excoffier L (2000) Arlequin, version 2,000: a software of population genetic data analysis. University of Geneva, Geneva
Schulte PM (2001) Environmental adaptations as windows on molecular evolution. Comp Biochem Physiol B 128:597–611
Seeb LW, Seeb JE, Polovina JJ (1990) Genetic variation in highly exploited spiny lobster Panulirus marginatus populations from the Hawaiian Archipelago. Fish Bull 88:713–718
Slatkin M (1993) Isolation by distance in equilibrium and non-equilibrium populations. Evolution 47:264–279
Slatkin M, Hudson RH (1991) Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations. Genetics 129:555–562
Stepien CA (1999) Phylogeographical structure of the Dover sole Microstomus pacificus: the larval retention hypothesis and genetic divergence along the deep continental slope of the northeastern Pacific Ocean. Mol Ecol 8:923–939
Swofford DL (2002) PAUP: Phylogenetic Analysis Using Parsimony (and other methods), version 4. Sinauer Associates, Sunderland
Tajima F (1983) Evolutionary relationship of DNA sequence in finite populations. Genetics 105:437–467
Tajima F (1989) Statistical-method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamaki K, Honza E (1991) Global tectonics and formation of marginal basins: role of the western Pacific. Episodes 14(3):224–230
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Tang WQ, Hu XL, Yang JQ (2007) Species validities of Coilia brachygnathus and C. nasus taihuensis based on sequence variation of complete mtDNA control region. Biodiv Sci 15:224–231
Tudela S, Garca-Marn JL, Pla C (1999) Genetic structure of the European anchovy, Engraulis encrasicolus, in the northwest Mediterranean. J Exp Mar Biol Ecol 234:95–109
Wang PX (1999) Response of Western Pacific marginal seas to glacial cycles: paleoceanographic and sedimentological features. Mar Geol 156:5–39
Xia YZ, Sheng Y, Chen YY (2006) DNA sequence variation in the mitochondrial control region of lenok (Branchymystax lenok) populations in China. Biodiv Sci 14:48–54
Xiao YS, Takahashi M, Yanagimoto T, Zhang Y, Gao TX, Yabe M, Sakurai Y (2008) Genetic variation and population structure of willowy flounder Tanakius kitaharai collected from Aomori, Ibaraki and Niigata in Northern Japan. Afr J Biotechnol 7(21):3836–3844
Acknowledgments
We are grateful to Prof. Syuiti Abe for checking the content of the article. Thanks to Mr. Nwafili Sylvanus and Dr. Jinxian Liu for checking the English. The study was supported by State 863 High-Technology R&D Project of China (No. 2006AA09Z418), the National Key Basic Research Program from the Ministry of Science and Technology, P.R. China (No. 2005CB422306).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xiao, Y., Gao, T., Zhang, Y. et al. Demographic History and Population Structure of Blackfin Flounder (Glyptocephalus stelleri) in Japan Revealed by Mitochondrial Control Region Sequences. Biochem Genet 48, 402–417 (2010). https://doi.org/10.1007/s10528-009-9321-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-009-9321-8