Abstract
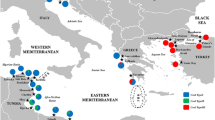
In this study, the genetic structure and phylogeny of the European flounder (Platichthys flesus L.) was investigated through the analysis of three mitochondrial gene sequences: COI, 16S rRNA, and cyt-b with specimens collected from the Black Sea and the Baltic Sea. In addition, available mtDNA sequences of P. flesus with known geographic information (from the Baltic Sea, the North Sea, the White Sea, the Atlantic Ocean, and the Mediterranean Sea) were retrieved from the GenBank database and included in phylogenetic analysis to increase resolution. The Black Sea is comprised of a geographically isolated European flounder population. Among analyzed mtDNA sequences, cyt-b sequence appeared to be more informative for detecting genetic differentiation at population level whereas 16S rRNA sequence was the less informative. The maximum likelihood trees generated based on COI, 16S rRNA, and cyt-b sequences suggested the presence of distinct European flounder population in the Black Sea, separating it from the European flounder populations from the rest of the geographic regions. Similarly, the highest genetic distances were obtained between the Black Sea population and the rest of the populations. The low salinity level of the Black Sea and narrow straits that connect the Black Sea to the Mediterranean Sea may function as an effective barrier that restrict gene flow and larval dispersal. We suggest designation of the Black Sea European flounder population as an evolutionary significant unit.

Similar content being viewed by others
Availability of Data and Materials
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Albaina A, Fox CJ, Taylor N, Hunter E, Maillard M, Taylor MI (2010) A TaqMan real-time PCR based assay targeting plaice (Pleuronectes platessa L.) DNA to detect predation by the brown shrimp (Crangon crangon L.) and the shore crab (Carcinus maenas L.)-Assay development and validation. J Exp Mar Bio Ecol 391:178–189. https://doi.org/10.1016/j.jembe.2010.06.029
Aydın I, Ak O, Kucuk E, Polat H, Ceylan B (2012) Optimum temperature and growth performance of hatchery reared Black Sea flounder (Platichthys flesus luscus Pallas, 1814). Turk J Vet Anim Sci 36:101–106
Aydin I, Firidin Ş, Cebeci A, ÖZTÜRK RÇ, Terzi Y, Polat H, Küçük E (2023) Gonad development of black sea european flounder, platichthys flesus, under culture conditions. Thalassas: Int J Mar Sci 1–6. https://doi.org/10.1007/s41208-023-00538-5
Aydin I, Kucuk E, Sahin T, Kumlu M (2015) Effect of Temperature on Reversed Asymmetry in Hatchery-Reared Flounder (Platichthys flesus luscus Pallas, 1811). Turkish J Fish Aquat Sci 15:737–740. https://doi.org/10.4194/1303-2712-v15_3_17
Aydın İ, Şahin T, Polat H, Küçük E (2011) Kuluçkahanede yetiştirilen pisi balığı (Platichthys flesus luscus PALLAS, 1814)’nin spermatolojik özellikleri. J Fish 5:270–278. https://doi.org/10.3153/jfscom.2011031
Berendzen PB, Dimmick WW (2002) Phylogenetic relationships of pleuronectiformes based on molecular evidence. BioOne Complet 3:642–652. https://doi.org/10.1643/0045-8511
Bergs LS (1932) Revision des formes de Pleuronectes flesus. Notas y Resumenes Inst Esp Ocenografia 58:1–7
Borsa P, Blanquer A, Berrebi P (1997) Genetic structure of the flounders Platichthys flesus and P. stellatus at different geographic scales. Mar Biol 129:233–246. https://doi.org/10.1007/s002270050164
Calvès I, Lavergne E, Meistertzheim A, Charrier G, Cabral H, Guinand B, Quiniou L, Laroche J (2013) Genetic structure of European flounder Platichthys flesus: effects of both the southern limit of the species’ range and chemical stress. Mar Ecol Prog Ser 472:257–273. https://doi.org/10.3354/meps09797
Chapleau F (1993) Pleuronectiform Relationships: A Cladistic Reassessment. Bull Mar Sci 52:516–540
Chapleau F, Keast A (1988) A phylogenetic reassessment of the monophyletic status of the family Soleidae, with comments on the suborder Soleoidei (Pisces; Pleuronectiformes). Can J Zool 66:2797–2810. https://doi.org/10.1139/z88-408
Costa FO, Landi M, Martins R, Costa MH, Costa ME, Carneiro M, Alves MJ, Steinke D, Carvalho GR (2012) A ranking system for reference libraries of DNA barcodes: application to marine fish species from Portugal. PLoS ONE 7:1–9. https://doi.org/10.1371/journal.pone.0035858
Do Prado F, Vera M, Hermida M, Bouza C, Pardo BG, Vilas R, Blanco A, Fernandez C, Maroso F, Maes GE, Turan C, Volckaert FAM, Taggart JB, Carr A, Ogden R, Nielsen EE, Martiner P (2018) Parallel evolution and adaptation to environmental factors in a marine flatfish: Implications for fisheries and aquaculture management of the turbot (Scophthalmus maximus). Evol Appl 11(8):1322–1341. https://doi.org/10.1111/eva.12628
Engell-Sørensen K, Støttrup JG, Holmstrup M (2004) Rearing of flounder (Platichthys flesus) juveniles in semiextensive systems. Aquaculture 230:475–491. https://doi.org/10.1016/S0044-8486(03)00437-X
Ericson GP, Zuccon D, Edmark VN (2020) A DNA key to Swedish vertebrates.
Espiñeira M, González-Lavín N, Vieites JM, Santaclara FJ (2008) Development of a method for the genetic identification of flatfish species on the basis of mitochondrial DNA sequences. J Agric Food Chem 56:8954–8961. https://doi.org/10.1021/jf800570r
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Firidin S, Öztürk RÇ, Alemdag M, Eroglu O, Terzi Y, Kutlu I, Duzgunes ZD, Cakmak E, Aydin I (2020) Population Genetic Structure of Turbot ( Scophthalmus maximus L., 1758) in the Black Sea. J Fish Biol 97:1154–1164. https://doi.org/10.1111/jfb.14487
Florin AB, Höglund J (2008) Population structure of flounder (Platichthys flesus) in the Baltic Sea: Differences among demersal and pelagic spawners. Heredity (edinb) 101:27–38. https://doi.org/10.1038/hdy.2008.22
Galleguillos RA, Ward RD (1982) Genetic and morphological divergence between populations of the flatfish Platichthys flesus (L.) (Pleuronectidae). Biol J Linn Soc 17:395–408. https://doi.org/10.1111/j.1095-8312.1982.tb02029.x
Geiger MF, Herder F, Monaghan MT, Almada V, Barbieri R, Bariche M, Berrebi P, Bohlen J, Casal-Lopez M, Delmastro GB, Denys GPJ, Dettai A, Doadrio I, Kalogianni E, Kärst H, Kottelat M, Kovačić M, Laporte M, Lorenzoni M, Marčić Z, Özuluğ M, Perdices A, Perea S, Persat H, Porcelotti S, Puzzi C, Robalo J, Šanda R, Schneider M, Šlechtová V, Stoumboudi M, Walter S, Freyhof J (2014) Spatial heterogeneity in the Mediterranean Biodiversity Hotspot affects barcoding accuracy of its freshwater fishes M. Mol Ecol Resour 14:1210–1221. https://doi.org/10.1111/1755-0998.12257
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
He S, Mork J, Larsen WB, Møller PR, Berumen ML (2020) Morphology and genetic investigation of flatfish interspecies hybrids (Pleuronectes platessa X Platichthys flesus) from the Baltic Sea. Fish Res 225:105498. https://doi.org/10.1016/j.fishres.2020.105498
Hemmer-Hansen J, Nielsen EE, Frydenberg J, Loeschcke V (2007a) Adaptive divergence in a high gene flow environment: Hsc70 variation in the European flounder (Platichthys flesus L.). Heredity (edinb) 99:592–600. https://doi.org/10.1038/sj.hdy.6801055
Hemmer-Hansen J, Nielsen EE, Grønkjær P, Loeschcke V (2007b) Evolutionary mechanisms shaping the genetic population structure of marine fishes; lessons from the European flounder (Platichthys flesus L.). Mol Ecol 16:3104–3118. https://doi.org/10.1111/j.1365-294X.2007.03367.x
Jokinen H, Momigliano P, Merilä J (2019) From ecology to genetics and back: the tale of two flounder species in the Baltic Sea. ICES J Mar Sci 76(7):2267–2275. https://doi.org/10.1093/icesjms/fsz151
Keskin E, Atar HH (2013) DNA barcoding commercially important fish species of Turkey. Mol Ecol Resour 13:788–797. https://doi.org/10.1111/1755-0998.12120
Kijewska A, Burzyński A, Wenne R (2009) Molecular identification of European flounder (Platichthys flesus) and its hybrids with European plaice (Pleuronectes platessa). ICES J Mar Sci 66:902–906. https://doi.org/10.1093/icesjms/fsp110
Knebelsberger T, Dunz AR, Neumann D, Geiger MF (2014a) Molecular diversity of Germany’s freshwater fishes and lampreys assessed by DNA barcoding. Mol Ecol Resour 15:562–572. https://doi.org/10.1111/1755-0998.12322
Knebelsberger T, Landi M, Neumann H, Kloppmann M, Sell AF, Campbell PD, Laakmann S, Raupach MJ, Carvalho GR, Costa FO (2014b) A reliable DNA barcode reference library for the identification of the North European shelf fish fauna. Mol Ecol Resour 14:1060–1071. https://doi.org/10.1111/1755-0998.12238
Kocher TD, Thomas WK, Meyer A, Edwards SV, Paabo S, Villablanca FX, Wilson AC (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci 86:6196–6200. https://doi.org/10.1073/pnas.86.16.6196
Kochzius M, Seidel C, Antoniou A, Botla SK, Campo D, Cariani A, Vazquez EG, Hauschild J, Hervet C, Hjörleifsdottir S, Hreggvidsson G, Kappel K, Landi M, Magoulas A, Marteinsson V, Nölte M, Planes S, Tinti F, Turan C, Venugopal MN, Weber H, Blohm D (2010a) Identifying fishes through DNA barcodes and microarrays. PLoS ONE 5:1–15. https://doi.org/10.1371/journal.pone.0012620
Kochzius M, Seidel C, Antoniou A, Botla SK, Campo D, Cariani A, Vazquez EG, Hauschild J, Hervet C, Hjörleifsdottir S, Hreggvidsson G, Kappel K, Landi M, Magoulas A, Marteinsson V, Nölte M, Planes S, Tinti F, Turan C, Venugopal MN, Weber H, Blohm D (2010b) Identifying Fishes through DNA Barcodes and Microarrays. PLoS ONE 5:e12620. https://doi.org/10.1371/journal.pone.0012620
Kuciński M, Jakubowska-Lehrmann M, Góra A, Mirny Z, Nadolna-Ałtyn K, Szlinder-Richert J, Ocalewicz K (2023) Population Genetic Study on the European Flounder (Platichthys flesus) from the Southern Baltic Sea Using SNPs and Microsatellite Markers. Animals 13:1448. https://doi.org/10.3390/ani13091448
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kume M, Lavergne E, Ahn H, Terashima Y, Kadowaki K, Ye F, Kameyama S, Kai Y, Henmi Y, Yamashita Y, Kasai A (2021) Factors structuring estuarine and coastal fish communities across Japan using environmental DNA metabarcoding. Ecol Ind 121:107216. https://doi.org/10.1016/j.ecolind.2020.107216
Laakkonen HM, Hardman M, Strelkov P, Väinölä R (2021) Cycles of trans-Arctic dispersal and vicariance, and diversification of the amphi-boreal marine fauna. J Evol Biol 34:73–96. https://doi.org/10.1111/jeb.13674
Landi M, Dimech M, Arculeo M, Biondo G, Martins R, Carneiro M, Carvalho GR, Brutto SL, Costa FO (2014) DNA barcoding for species assignment: The case of Mediterranean marine fishes. PLoS ONE 9:e106135. https://doi.org/10.1371/journal.pone.0106135
Larsen PF, Nielsen EE, Williams TD, Hemmer-Hansen J, Chipman JK, Kruhøffer M, Grønkjær P, George SG, Dyrskjøt L, Loeschcke V (2007) Adaptive differences in gene expression in European flounder (Platichthys flesus). Mol Ecol 16:4674–4683. https://doi.org/10.1111/j.1365-294X.2007.03530.x
Leigh JW, Bryant D (2015) POPART: full-feature software for haplo- type network construction. Methods Ecol Evol 6:1110–1116
Librado P, Rozas J (2009) DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Luo Z, Yang C, Wang L, Liu Y, Shan B, Liu M, Chen C, Guo T, Sun D (2023) Relationships between fish community structure and environmental factors in the nearshore waters of hainan island. South China Diversity 15:901. https://doi.org/10.3390/d15080901
Marko PB, Hart MW (2018) Genetic analysis of larval dispersal, gene flow, and connectivity. In Evolutionary Ecology of Marine Invertebrate Larvae. Edited by Tyler J. Carrier, Adam M. Reitzel, and Andreas Heyland: Oxford University Press. https://doi.org/10.1093/oso/9780198786962.003.0012
Macpherson E, Raventos N. (2006) Relationship between pelagic larval duration and geographic distribution of Mediterranean littoral fishes. Mar Ecol Prog Ser 327:257–265. https://doi.org/10.3354/meps327257
Momigliano P, Denys GP, Jokinen H, Merila J (2018) Platichthys solemdali sp. nov. (Actinopterygii, Pleuronectiformes): A New Flounder Species From the Baltic Sea. Front Mar Sci 5:225. https://doi.org/10.3389/fmars.2018.00225
Morais P, Dias E, Babaluk J, Antunes C (2011) The migration patterns of the European flounder Platichthys flesus (Linnaeus, 1758) (Pleuronectidae, Pisces) at the southern limit of its distribution range: Ecological implications and fishery management. J Sea Res 65:235–246. https://doi.org/10.1016/j.seares.2010.11.001
Nissling A, Dahlman G (2010) Fecundity of flounder, Pleuronectes flesus, in the Baltic Sea—reproductive strategies in two sympatric populations. J Sea Res 64:190–198. https://doi.org/10.1016/j.seares.2010.02.001
Nissling A, Larsson R (2018) Population specific sperm production in European flounder Platichthys flesus : Adaptation to salinity at spawning. J Fish Biol 93:47–52. https://doi.org/10.1111/jfb.13667
Nissling A, Nyberg S, Petereit C (2017) Egg buoyancy of flounder, Platichthys flesus, in the Baltic Sea—adaptation to salinity and implications for egg survival. Fish Res 191:179–189. https://doi.org/10.1016/j.fishres.2017.02.020
Öztürk RÇ (2023) Genetic identification of ichthyoplankton in the Black Sea and their abundance and community assemblages. Turkish J Fish Aquat Sci 23(10):TRJFAS23514. https://doi.org/10.4194/TRJFAS23514
Öztürk RÇ, Altınok İ (2021) Genetic population structure of the striped venus clam Chamelea gallina across its range. Fish Res 234:105758. https://doi.org/10.1016/j.fishres.2020.105758
Palumbi SR (1996) Nucleic acids II: the polymerase chainreaction. In: Hillis DM, Moritz C, Mable BK (eds) Molecular Systematics. Sinauer Associates, Sunderland, pp 205–247
Pardo BG, Machordom A, Foresti F, Porto-Foresti F, Azevedo MFC, Bañon R, Sánchez L, Martínez P (2005) Phylogenetic analysis of flatfish (order pleuronectiformes) based on mitochondrial 16s rDNA sequences. Sci Mar 69:531–543. https://doi.org/10.3989/scimar.2005.69n4531
Pascual M, Rives B, Schunter C, Macpherson E (2017) Impact of life history traits on gene flow: A multispecies systematic review across oceanographic barriers in the Mediterranean Sea. PLoS ONE 12(5):e0176419. https://doi.org/10.1371/journal.pone.0176419
Pereira R, Rodrigues SM, Silva DM, Ramos S (2023) Assessing Environmental Control on Temporal and Spatial Patterns of Larval Fish Assemblages in a Marine Protected Area. Ecologies 4:288–309. https://doi.org/10.3390/ecologies4020019
Posada D (2008) jModelTest: Phylogenetic Model Averaging. Mol Biol Evol 25:1253–1256. https://doi.org/10.1093/molbev/msn083
Reiss H, Hoarau G, Dickey-Collas M, Wolff W (2009) Genetic population structure of marine fish: Mismatch between biological and fisheries management units. Fish Fish 10(4):361–395. https://doi.org/10.1111/j.1467-2979.2008.00324.x
Reis-Santos P, Tanner SE, Aboim MA, Vasconcelos RP, Laroche J, Charrier G, Pérez M, Presa P, Gillanders BM, Cabral HN (2018) Reconciling differences in natural tags to infer demographic and genetic connectivity in marine fish populations. Sci Rep 8:1–12. https://doi.org/10.1038/s41598-018-28701-6
Şahin T, Güneş E, Aydın İ, Polat H (2008) Reproductive characteristics and egg development in flounder (Pleuronectes flesus luscus) in the southern Black Sea. Isr J Aquac – Bamidgeh, 60(1): 20–26.
Şahin T, Güneş E, Aydın İ, Kurtoğlu İZ (2012) Quantitative characteristics of wild European flounder (Platychthys flesus luscus) sperm. Isr J Aquac-Bamidgeh, 64, 1–5. https://doi.org/10.46989/001c.20610
Semushin AV, Fuks GV, Shilova NA (2015) Flatfishes of the white sea: new data on the biology of the arctic flounder liopsetta glacialis, european flounder platichthys flesus, and common dab limanda limand. J Ichthyol 55:527–539
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Tinti F, Piccinetti C, Tommasini S, Vallisneri M (2000) Mitochondrial DNA variation, phylogenetic relationships, and evolution of four Mediterranean genera of soles (soleidae, pleuronectiformes). Mar Biotechnol 2:274–284. https://doi.org/10.1007/s101269900035
Veliz D, Rojas-Hernández N, Fibla P, Dewitte B, Cornejo-Guzmán S, Parada C (2021) High levels of connectivity over large distances in the diadematid sea urchin Centrostephanus sylviae. PLoS ONE 16(11):e0259595. https://doi.org/10.1371/journal.pone.0259595.PMID:34735545;PMCID:PMC8568165
Verrez-Bagnis VS, Jerome M, Mouchel O, Iglesias S, Pruvost P (2017) FishTrace: a genetic catalogue of European fishes. Database 1–11. https://doi.org/10.1093/database/bax075
Vinnikov KA, Thomson RC, Munroe TA (2018) Revised classification of the righteye flounders (Teleostei: Pleuronectidae) based on multilocus phylogeny with complete taxon sampling. Mol Phylogenet Evol 125:147–162. https://doi.org/10.1016/j.ympev.2018.03.014
Walsh HJ, Richardson DE, Marancik KE, Hare JA (2015) Long-Term Changes in the Distributions of Larval and Adult Fish in the Northeast U.S. Shelf Ecosystem. Plos ONE 10 (9), e0137382. https://doi.org/10.1371/journal.pone.0137382
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN (2005) DNA barcoding Australia’s fish species. Philos Trans r Soc B Biol Sci 360:1847–1857. https://doi.org/10.1098/rstb.2005.1716
Funding
This work was partially supported by the Ministry of Agriculture and Rural Affairs, General Directorate of Agricultural Research (TAGEM), [grant number TAGEM/HAYSÜD/2006/].
Author information
Authors and Affiliations
Contributions
İ.A. administered the project and received the funding. Ş.F. performed the investigation and resources. R. Ç. Ö. Conceptualized the paper, performed investigation and formal analysis, and wrote the main manuscript text. M.A. performed the investigation. Y.T. performed visualization and revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethical Approval
The procedures applied in this study were performed in accordance with the ethical standards of the Agricultural Research and Policy General Directorate of the Ministry of Agriculture and Forestry of the Republic of Türkiye.
Competing Interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aydin, İ., Fi̇ri̇di̇n, Ş., Öztürk, R. et al. Genetically Distinct European Flounder (Platichthys Flesus, L.) Matriline in the Black Sea. Thalassas 40, 115–123 (2024). https://doi.org/10.1007/s41208-023-00634-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41208-023-00634-6