Abstract
Using baseline (ages 9–10) and two-year follow-up (ages 11–12) data from monozygotic and dizygotic twins enrolled in the longitudinal Adolescent Brain Cognitive DevelopmentSM Study, we investigated the genetic and environmental contributions to microstructure and volume of nine subcortical gray matter regions. Microstructure was assessed using diffusion MRI data analyzed using restriction spectrum imaging (RSI) and diffusion tensor imaging (DTI) models. The highest heritability estimates (estimate [95% confidence interval]) for microstructure were found using the RSI model in the pallidum (baseline: 0.859 [0.818, 0.889], follow-up: 0.835 [0.787, 0.871]), putamen (baseline: 0.859 [0.819, 0.889], follow-up: 0.874 [0.838, 0.902]), and thalamus (baseline: 0.855 [0.814, 0.887], follow-up: 0.819 [0.769, 0.857]). For volumes the corresponding regions were the caudate (baseline: 0.831 [0.688, 0.992], follow-up: 0.848 [0.701, 1.011]) and putamen (baseline: 0.906 [0.875, 0.914], follow-up: 0.906 [0.885, 0.923]). The subcortical regions displayed high genetic stability (rA = 0.743–1.000) across time and exhibited unique environmental correlations (rE = 0.194–0.610). Individual differences in both gray matter microstructure and volumes can be largely explained by additive genetic effects in this sample.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Understanding how genetic variability influences brain structure in adolescence may provide insights into the pathophysiology and biological constructs that impact normal brain development, cognition, and the development of neuropsychiatric disorders. Brain structures vary in heritability, suggesting that different genes may influence the structural development of different areas (Bis et al. 2012; den Braber et al. 2013; Stein et al. 2012) and the genetic influences may vary over time (Brouwer et al. 2017). Subcortical regions of the brain are associated with neuropsychiatric disorders including depression (Koolschijn et al. 2009), anxiety (Holzschneider and Mulert 2011), and schizophrenia (Shepherd et al. 2012), as well as intellectual functioning (Bohlken et al. 2014).
Studies, predominantly in adult populations, have investigated the heritability of subcortical volumes of anatomic structures in the brain (den Braber et al. 2013; Kremen et al. 2010). Hippocampal and intracranial volume have been associated with genes located on chromosome 12q24 (Bis et al. 2012; Stein et al. 2012; Yoon et al. 2011). A meta-analysis of twin studies reported high heritability from 0.52 for the right thalamus to 0.82 for the right putamen volume, but with a relatively small sample size and wide confidence intervals of 0.4–0.8 for the right thalamus (Blokland et al. 2012). Another twin study that looked at subcortical volumes over a 5 year period also found high heritability for the thalamus, caudate, and putamen (den Braber et al. 2013). Other studies on adult twins have demonstrated high heritability of total intracranial brain volume (Posthuma et al. 2000; Thompson et al. 2001).
The brain undergoes rapid development in the first two years of life and nears adult size by early childhood at around age six (Jansen et al. 2015), but more dynamic changes in cortical and subcortical regions involving gray and white matter continue to occur into adolescence and young adulthood (Lenroot and Giedd 2006). Prior work in neurodevelopment has suggested greater genetic heritability earlier in childhood for primary cortices (motor, somatosensory), and more heritability in later adolescence for structures unique to the human species such as the dorsolateral prefrontal cortex (Lenroot and Giedd 2006).
There are few reports on adolescents with respect to heritability of subcortical volumes. A study of twins scanned at ages 9 and 12 showed high heritability for intracranial volume, gray matter, and white matter volume from 0.77 to 0.91 (Peper et al. 2009). In another study of 8 year-olds, heritability ranged from 0.57 to 0.79 for gray matter, white matter, corpus callosum, and total cerebrum, but with higher heritability in the left cerebral hemisphere (Yoon et al. 2011). A larger study of 326 twins resulted in heritability measures of 0.65 for corpus callosum, 0.64 for basal ganglia, and 0.42 for the thalamus, but lower for the ventricles (0.17) and cerebellum (0.24) (Schmitt et al. 2007).
Phenotypic neurodevelopmental trajectory research shows that there are significant patterns of age-related change in subcortical volumes across adolescence (Herting et al. 2018); however, it is unclear which etiological influences underlying the phenotypes remain stable. Longitudinal twin research allows for testing whether the genetic and environmental influences on subcortical structure are stable in adolescence, or if there are new genetic influences that emerge. Anomalies in brain structure are a robust correlate to adolescent and adult psychopathology (Gurholt et al. 2022; Guyer 2020), and understanding the stable and emerging influences in brain development may provide insights into the relationship between genetics, brain development and mental health.
One previous longitudinal twin study found that subcortical volumes (thalamus, hippocampus, amygdala, putamen, caudate, pallidum, and nucleus accumbens) were highly heritable and entirely genetically stable from ages 9 to 12; however the sample size was limited (48 MZ pairs; 64 DZ pairs)(Swagerman et al. 2014).
ABCD study
While several studies have considered the heritability of the volumes of subcortical structures, very few have investigated their microstructure (Gillespie et al. 2017), which can be probed non-invasively using diffusion MRI. In this study, we leverage the Adolescent Brain Cognitive DevelopmentSM Study (ABCD Study→) to investigate the heritability of both subcortical volume and microstructure in adolescents.
The ABCD study is a longitudinal study of brain development that includes extensive cognitive, behavioral, genetic and imaging data (Casey et al. 2018). The study is enriched for same sex twin pairs (Iacono et al. 2018), enabling the assessment of the genetic and environmental influences on a wide variety of traits over development. We use twin data to estimate the genetic and environmental influences on deep gray matter volume, assessed using conventional structural MRI, and microstructure, assessed using diffusion MRI.
The imaging battery included in ABCD, and repeated every two years, includes structural and diffusion MRI. Structural MRI scans have high (1 mm) spatial resolution, and sufficient gray/white matter contrast to enable segmentation of deep gray matter structures, and determination of their volumes. Diffusion MRI uses the motion of water molecules in the brain to provide information on the microscopic environment, and is sensitive to developmental changes related to, for example, myelination and cell density (Watts et al. 2003).
To our knowledge, this is the largest twin sample used to assess subcortical volume and microstructure heritability, as well as the first to longitudinally test the genetic and environmental contributions to subcortical microstructure in adolescence.
Data and methods
Subject selection
The ABCD data used in this report came from Curated Data Release 4.0. To maximize data consistency for the technically demanding diffusion acquisition, only data from twins scanned using a single MRI vendor (Siemens Healthineers AG) were included in the diffusion analysis. To maximize sample size, volumetric data for twins scanned on both Siemens and Philips (Philips HealthTech) scanners was included. Twin pairs were excluded if they had incomplete data, or if the structural and diffusion imaging data did not pass ABCD-recommended quality control, including a diffusion mean motion threshold of 1.5 mm. Subjects with a history of brain injury, cerebral palsy, muscular dystrophy, multiple sclerosis, or substance abuse were excluded. Demographic information for the two groups is given in Table 1.
MRI data acquisition
The MRI acquisition is described in more detail elsewhere (Casey et al. 2018). Briefly, structural images were acquired using a T1-weighted 3D MP-RAGE sequence with 1 mm isotropic spatial resolution, TE/TI/TR = 2.88/1060/2500ms, flip angle = 8°, sagittal field of view of 256 × 256 × 176 mm3. Real-time motion detection and correction using volumetric navigators was used to minimize the effects of head motion (Tisdall et al. 2012).
Diffusion-weighted images were acquired using a 2D spin-echo EPI sequence with b-values of 500 (6 directions), 1000 (15 directions), 2000 (15 directions), and 3000 (60 directions) s/mm2, as well as 6 reference acquisitions without diffusion weighting. Diffusion imaging was acquired with 1.7 mm isotropic spatial resolution, TE/TR = 88/4100ms, 81 slices with a multiband factor of 3, axial field of view of 240 × 240 mm2. Corresponding field maps were acquired to enable correction of EPI susceptibility distortions.
MRI data processing
Volumetric data
Imaging data was preprocessed as described previously (Hagler et al. 2019). Segmentation of subcortical structures was performed using FreeSurfer (Fischl et al. 2002), providing measures of the volume of each structure and masks for the aligned diffusion data. Subcortical volumes were normalized to the total intracranial volume.
Restriction spectrum imaging
A restriction spectrum imaging (RSI) (White et al. 2013) model was applied to the diffusion data to generate measures of the isotropic and directional restricted and hindered components of diffusion, as well as a free water component. The isotropic restricted component has been shown to be sensitive to tissue cellularity (White et al. 2013). Deep gray matter structures display low directional anisotropy, so we limited our analysis to the restricted normalized isotropic (RNI) component.
Diffusion tensor imaging
A conventional diffusion tensor imaging (DTI) model was also applied to generate measures of mean diffusivity (MD) to compare its sensitivity to the more sophisticated RSI model.
Zygosity status was genetically determined. All data was residualized to account for the effects of age, sex, and site.
Descriptive statistics
Intraclass correlations (ICCs) between MZ and DZ twin pairs were calculated as descriptive statistics to visualize the relative twin similarity for the phenotypes. Within subject Pearson correlations for phenotypes derived at baseline and year 2 were calculated to provide a proxy measure of the test-retest reliability of the imaging metrics. The relationship between the estimated heritability for each region/metric and the within-subject stability was calculated as a Pearson correlation.
Assumptions of the ACE model
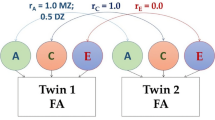
The analysis presented is based on the ACE model. MZ twins share 100% of their genetic information, while DZ twins share approximately 50%. Additive genetic effects imply that the expected genetically-based correlation between DZ twins should be half that of MZ twins. Non-linear effects, such as those caused by genetic dominance, are not included in the model. The ACE model also incorporates the equal environments assumption (EEA), that the effect of the common environment is the same for MZ and DZ twins. Violation of this assumption would be expected to result in greater similarity of MZ twins relative to DZ twins, increasing the apparent heritability. The unique environment term models uncorrelated differences between twin pairs, including differences caused by measurement error.
Structural equation modeling
Structural Equation Modelling was used to assess the additive genetic (A), common environment (C), and unique environmental (E) contributions to tissue microstructure using the OpenMx (Boker et al. 2011) package in R (R version 3.6.1, R Core Team). A direct variance ACE model was used (Verhulst et al. 2019). Correlated factors models were employed to assess the stability of the variance components by decomposing the cross-time covariance of the trait into 3 latent sources of contributions (rA, rC, or rE)(Loehlin 1996). Likelihood-based confidence intervals are provided to account for the non-normal distribution of the variance components (Neale and Miller 1997). If the estimates’ confidence interval does not include 1.00, this is suggestive of novel etiological influences. Model fit was assessed using the criteria of root-mean-square error (RMSEA) < 0.06 and a Comparative Fit Index (CFI) > 0.95 (Hu and Bentler 1998). Chi-square difference tests were used to assess significance (p < 0.05) of the etiological correlations to account for the standard error invariance to model parameterization in genetic models (Neale et al. 1989).
Results
Subjects
The demographics of participants recruited and included in structural and diffusion analyses are shown in Table 1. For the volumetric measures at baseline, 284 MZ and 369 DZ twin pairs had acceptable quality data and were included in the analysis. For diffusion analysis, 148 MZ and 212 DZ twin pairs were included at baseline. At two-year follow-up the corresponding numbers were 207/247 and 134/158 for volume and diffusion respectively. The MZ and DZ groups did not differ significantly in age, sex, household income, parental education, or race/ethnicity (chi-squared test, p > 0.05).
Baseline results
Intraclass correlations (ICCs) for MZ and DZ twin pairs for each region and metric at baseline are shown in Fig. 1. For MZ twins, ICCs ranged from 0.610 (caudate) to 0.867 (pallidum) for RNI, 0.347 (hippocampus) to 0.673 (putamen) for MD, and 0.642 (nucleus accumbens) to 0.910 (brain stem) for volumes. With the exception of MD in the hippocampus and brainstem, the ICCs for DZ twins were significantly less than those from MZ twins (two sample t-test, p < 0.05).
Structural equation modelling estimates of the additive genetic, common, and unique environmental contributions to each region and metric are shown in Table 2; Fig. 2 (top). The model fit statistics can be found in Supplementary Tables 1–3. Heritability estimates (estimate, 95% confidence interval) for RNI at baseline in the pallidum (0.859, 0.818–0.889), putamen (0.859, 0.819–0.889), and thalamus (0.855, 0.814–0.887) were especially high. For MD, heritability was generally lower than for RNI, but highest in the putamen (0.704, 0.622–0.768), pallidum (0.672, 0.596–0.734), and thalamus (0.670, 0.573–0.744). For structure volumes, the putamen (0.906, 0.875–0.914) and caudate (0.831, 0.688–0.992) were found to be highly heritable.
Additive genetic (A), common environmental (C), and unique environmental contributions to restricted normalized isotropic (RNI, left) signal, mean diffusivity (MD, middle), and volumes (right) in subcortical regions at baseline (top) and two-year follow-up (bottom). Negative estimates for contributions to MD in nucleus accumbens have been replaced with zeros for display
Two-year follow-up results
The contributions to each region and metric are shown in Table 2; Fig. 2 (bottom). The model fit statistics can be found in Supplementary Tables 1–3. Heritability estimates for RNI showed a similar pattern at two-year follow-up, with the pallidum (0.835, 0.787–0.871), putamen (0.874, 0.838–0.902) and thalamus (0.819, 0.767–0.857) again demonstrating the highest heritability. For MD, the amygdala, caudate, pallidum, putamen, and thalamus had similar heritabilities, with estimates in the range of 0.627 to 0.701. As at baseline, structure volumes were most highly heritable for the putamen (0.906, 0.885–0.923) and caudate (0.848, 0.701–1.011).
At two-year follow-up, significant (p < 0.05) contributions of the common environment to MD in the nucleus accumbens (0.465, 0.185–0.728) and the volume of the brainstem (0.225, 0.074–0.355) were observed.
Longitudinal model results
The correlated factors models assessed the stability of etiological influences on subcortical regions between measures at baseline and the two-year follow-up (Table 3). For RNI, the genetic correlations ranged from 0.743 (amygdala) to 0.970 (caudate). The hippocampus and caudate appear to be genetically stable across time (rA confidence interval includes 1), but the other phenotypes may have novel genetic influences (rA confidence interval do not include 1). The unique environmental correlations ranged from 0.183 (caudate) to 0.660 (amygdala). For MD, the genetic correlations ranged from 0.772 (ventral diencephalon) to 0.967 (amygdala). The confidence intervals for the brainstem, amygdala, hippocampus, pallidum, ventral diencephalon include 1.00, suggesting genetic stability across time. The rA and rC coefficients could not be estimated for the nucleus accumbens MD measure as there was not both a positive A or positive C variance component to correlate. The unique environmental correlations were − 0.038 (nucleus accumbens) to 0.527 (hippocampus). For volume, the genetic correlations were 0.865 (pallidum) to 1 (brainstem). The confidence intervals for the brainstem, nucleus accumbens, caudate, hippocampus, and ventral diencephalon included 1. The unique environmental correlations for the volumetric measures ranged from 0.292 (pallidum) to 0.610 (hippocampus).
Stability of measures and heritability estimates
The Pearson correlations between measures at baseline and two-year follow-up are shown in Fig. 3. For RNI, the correlations ranged from 0.717 (amygdala) to 0.879 (pallidum). For MD, the correlations were substantially lower, ranging from 0.388 (ventral diencephalon) to 0.751 (pallidum). The volume of the pallidum displayed the lowest correlation between time points (0.647), while the caudate was highest (0.931).
Figure 4 shows that those measures that were stable between time points (r = 0.821–0.922, pallidum, putamen, thalamus for RNI, and brainstem, caudate, hippocampus and putamen for volume) demonstrated high heritability estimates (0.714–0.906 at baseline; 0.659–0.906 at two-year follow-up). Conversely, those measures that were least stable showed relatively low heritabilities (such as MD in the brain stem, nucleus accumbens, and ventral diencephalon). The Pearson correlation between heritability and stability was 0.800 (95% CI 0.604–0.905), suggesting that 64% on the variance in heritability across regions/metrics can be explained by the stability of the measurement.
Discussion
The RNI metric based on an RSI model was more consistent between time points and generally yielded higher heritability estimates compared to MD derived from a diffusion tensor model. The RSI model is biologically based, and substantially more sophisticated than the DTI model, modelling the restricted, hindered, isotropic, anisotropic, and free fluid components separately. By comparison the DTI model combines these components, and the derived MD values are susceptible to contamination, for example from partial volume averaging of free fluid (which has a very high value of MD) with adjacent tissue due to poor tissue segmentation. RNI may be a more reliable microstructure measure to be used in the scope of twin research. A previous study (den Braber et al. 2013) noted poor 5-year retest reliability in the volumes of the nucleus accumbens and pallidum.
The highest microstructure heritability estimates were for the pallidum, putamen, and thalamus. To our knowledge, this is the first study to examine the etiology of subcortical microstructure in adolescents. A prior study (Gillespie et al. 2017) considered heritability of both subcortical MD and volume in middle-aged males. Their results for MD are similar to the present study, with some structures showing higher heritability in adults (caudate, pallidum, putamen), and some lower (amygdala, thalamus). Increasing heritability with age may represent the evolving influence of genetics over development, while decreasing may be due to diverging environmental impacts, especially during adulthood.
For volumes, the highest heritability estimates were for the brainstem, caudate, and putamen. The heritability of subcortical volumes were generally higher or comparable in our cohort compared to prior studies (Swagerman et al. 2014; Yoon et al. 2011). Our results are similar to those of den Braber (den Braber et al. 2013), who found the highest heritability in the left (0.88) and right (0.86) caudate in a cohort of young adults.
The consistency of high heritability estimates across RNI microstructure and volume measures provides evidence that individual variation in these subcortical measures is highly influenced by additive genetic effects. Anomalies in adolescent subcortical structure have been associated with negative outcomes, such as psychopathology (Gurholt et al. 2022; Guyer 2020). The high heritability of these phenotypes suggests they could serve as intermediary phenotypes to identify risk genes for heritable psychopathologies as well as provide insight into possible mechanisms (Glahn et al. 2007; Gurholt et al. 2022).
There was evidence of shared environmental effects influencing brainstem volume and mean diffusivity of the nucleus accumbens at follow-up year 2. However, the nucleus accumbens is often susceptible to low test-retest reliability due to its size and location and this finding should be approached with caution. The brainstem has been associated with neurodevelopmental disorders (Dadalko and Travers 2018). Future research could explore whether shared environmental risk factors, such as nutrition, share variance with the brainstem at this developmentally-sensitive age.
The hippocampus and caudate showed genetic stability (rA confidence intervals included 1) across all three imaging modalities. The putamen and thalamus displayed the possibility of novel genetic influences in all three modalities; however, given the magnitude of the genetic correlations, the genetic effects are mostly stable. All the other phenotypes demonstrated high genetic correlations but were inconsistent across modality on whether there were novel genetic influences. The putamen and thalamus displayed the possibility of novel genetic influences in all three modalities; however, given the magnitude of the genetic correlations, the genetic effects are mostly stable. All the other phenotypes demonstrated high genetic correlations but were inconsistent across modality on whether there were novel genetic influences.
The large genetic correlations across time are suggestive that genetic influences on subcortical development remain relatively stable from ages 10 to 12. Besides the hippocampus and caudate, most of the genetic correlations were less than one in at least one of the imaging modalities, suggesting some novel genetic influences emerging. These findings contradict one previous longitudinal twin study which found no new genetic effects on subcortical brain volumes from ages 9 to 12 (Swagerman et al. 2014); however, the current study had a larger sample size.
In addition to high genetic stability, the subcortical regions displayed unique environmental correlations across time, which were strongest in magnitude for the hippocampus, amygdala, and caudate. Unique environmental correlations can be indicative of a unique risk factor that is not explained by familial influences, such as a traumatic event. Environmental stressors in childhood have been implicated in atypical, sub-nuclei development (McCrory et al. 2012). The literature is especially abundant on structural alterations in the amygdala and hippocampus due to childhood adversity (Calem et al. 2017) and the possible exacerbating impacts these anomalies may have on later psychopathology (McCrory et al. 2012). The unique environmental correlations support that there may be events not captured by familial influence which impact sub-cortical development, and that not all E on subcortical structures is random measurement error. Future research can explore etiological correlations between these subcortical measures in relation to risk factors with psychopathology, such as childhood adversity.
One previous study found that the longitudinal change rate of childhood thalamus, pallidum, and amygdala volumes had a significant heritable component (Brouwer et al. 2017). Future research could explore the heritability of the change rate of subcortical microstructure.
Limitations
The sample size is an important limitation of twin studies that involve imaging, and this study is no exception. The uncertainties in the derived heritability estimates are large, especially for those regions with moderate or low heritability, perhaps limited by measurement uncertainty. Particularly at baseline (ages 9–10), subject motion can also be problematic, although the ABCD study benefits from the use of prospective motion correction for the structural data acquisition, combined with recommended imaging quality control criteria.
Measurement error is expected to reduce the correlations between MZ twins and DZ twins similarly, resulting in lower heritability estimates for metrics that have low test-retest reliability (Fig. 4, r = 0.8). This implies that the estimates of heritabilities presented may represent a lower bound on the true heritability.
While we controlled for the effect of sex, we did not specifically investigate it. Prior studies have postulated that there are no sex differences in subcortical volumes (den Braber et al. 2013; Vink et al. 2012) which may suggest that the same genes influence the same phenotype in both sexes. However, these studies concentrated on adults.
Conclusions
Volume and diffusion MRI metrics demonstrated substantial heritability across all subcortical regions in this sample both at baseline and two-year follow-up. The RNI component of diffusion derived from the RSI model was found to be more heritable than MD, perhaps due to its better test-retest reliability. Particularly high heritability of greater than 0.8 was found using the RSI model in the pallidum, putamen, and thalamus, and in volumes of the caudate and putamen. These findings were consistent across time points. The subcortical microstructures and volumes were largely genetically stable, with some small, novel genetic influences. Almost all of the phenotypes had cross-time unique environmental correlations, suggesting that a non-familial risk factor may impact subcortical development.
Data availability
The ABCD data repository grows and changes over time. The ABCD data used in this report came from Curated Data Release 4.0, https://doi.org/10.15154/1523041.
Code availability (software application or custom code)
The direct variance ACE model used in this work was based on twoACEvc.R, available at. https://hermine-maes.squarespace.com/two.
References
Bis JC et al (2012) Common variants at 12q14 and 12q24 are associated with hippocampal volume. Nat Genet 44(5):545–551
Blokland GAM et al (2012) Genetic and environmental influences on neuroimaging phenotypes: a meta-analytical perspective on twin imaging studies. Twin Res Hum Genet 15(3):351–371
Bohlken MM et al (2014) Genes contributing to subcortical volumes and intellectual ability implicate the Thalamus. Hum Brain Mapp 35(6):2632–2642
Boker S et al (2011) OpenMx: an open source extended structural equation modeling framework. Psychometrika 76(2):306–317
Brouwer RM et al (2017) Genetic influences on individual differences in longitudinal changes in global and subcortical brain volumes: results of the ENIGMA plasticity working group. Hum Brain Mapp 38(9):4444–4458
Calem M et al (2017) Meta-analysis of associations between childhood adversity and hippocampus and amygdala volume in non-clinical and general population samples. NeuroImage Clin 14:471–479
Casey BJ et al (2018) The adolescent brain cognitive development (ABCD) study: imaging acquisition across 21 sites. Dev Cogn Neurosci 32:43–54
Dadalko OI, Travers BG (2018) Evidence for brainstem contributions to autism spectrum disorders. Front Integr Neurosci 12:47
den Braber A et al (2013) Heritability of subcortical brain measures: a perspective for future genome-wide association studies. NeuroImage 83:98–102
Fischl B et al (2002) Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33(3):341–355
Gillespie NA et al (2017) Genetic and environmental influences on mean diffusivity and volume in subcortical brain regions. Hum Brain Mapp 38(5):2589–2598
Glahn DC et al (2007) Imaging genomics: mapping the influence of genetics on brain structure and function. Hum Brain Mapp 28(6):461–463
Gurholt TP et al (2022) Intracranial and subcortical volumes in adolescents with early-onset psychosis: a multisite mega-analysis from the ENIGMA consortium. Hum Brain Mapp 43(1):373–384
Guyer AE (2020) Adolescent psychopathology: the role of brain-based Diatheses, Sensitivities, and susceptibilities. Child Dev Perspect 14(2):104–109
Hagler DJ Jr et al (2019) Image processing and analysis methods for the adolescent brain cognitive development study. NeuroImage 202:116091
Herting MM et al (2018) Development of subcortical volumes across adolescence in males and females: a multisample study of longitudinal changes. NeuroImage 172:194–205
Holzschneider K, Mulert C (2011) Neuroimaging in anxiety disorders. Dialogues Clin Neurosci 13(4):453–461
Hu LT, Bentler PM (1998) Fit indices in covariance structure modeling: sensitivity to underparameterized model misspecification. Psychol Methods 3(4):424–453
Iacono WG et al (2018) The utility of twins in developmental cognitive neuroscience research: how twins strengthen the ABCD research design. Dev Cogn Neurosci 32:30–42
Jansen AG et al (2015) What twin studies tell us about the heritability of brain development, morphology, and function: a review. Neuropsychol Rev 25(1):27–46
Koolschijn PC et al (2009) Brain volume abnormalities in major depressive disorder: a meta-analysis of magnetic resonance imaging studies. Hum Brain Mapp 30(11):3719–3735
Kremen WS et al (2010) Genetic and environmental influences on the size of specific brain regions in midlife: the VETSA MRI study. NeuroImage 49(2):1213–1223
Lenroot RK, Giedd JN (2006) Brain development in children and adolescents: insights from anatomical magnetic resonance imaging. Neurosci Biobehav R 30(6):718–729
Loehlin JC (1996) The Cholesky approach: a cautionary note. Behav Genet 26(1):65–69
McCrory E et al (2012) The link between child abuse and psychopathology: a review of neurobiological and genetic research. J R Soc Med 105(4):151–156
Neale MC, Miller MB (1997) The use of likelihood-based confidence intervals in genetic models. Behav Genet 27(2):113–120
Neale MC et al (1989) Fitting genetic models with lisrel - hypothesis-testing. Behav Genet 19(1):37–49
Peper JS et al (2009) Heritability of Regional and Global Brain structure at the Onset of Puberty: a magnetic resonance imaging study in 9-Year-old twin pairs. Hum Brain Mapp 30(7):2184–2196
Posthuma D et al (2000) Multivariate genetic analysis of brain structure in an extended twin design. Behav Genet 30(4):311–319
Schmitt JE et al (2007) A multivariate analysis of neuroanatomic relationships in a genetically informative pediatric sample. NeuroImage 35(1):70–82
Shepherd AM et al (2012) Systematic meta-review and quality assessment of the structural brain alterations in schizophrenia. Neurosci Biobehav Rev 36(4):1342–1356
Stein JL et al (2012) Identification of common variants associated with human hippocampal and intracranial volumes. Nat Genet 44(5):552–561
Swagerman SC et al (2014) Development and heritability of subcortical brain volumes at ages 9 and 12. Genes Brain Behav 13(8):733–742
Thompson PM et al (2001) Genetic influences on brain structure. Nat Neurosci 4(12):1253–1258
Tisdall MD et al (2012) Volumetric navigators for prospective motion correction and selective reacquisition in neuroanatomical MRI. Magnetic resonance in medicine: official journal of the society of magnetic resonance in medicine / society of magnetic resonance. Medicine 68(2):389–399
Verhulst B et al (2019) Type I error rates and parameter bias in multivariate behavioral genetic models. Behav Genet 49(1):99–111
Vink JM et al (2012) Sex differences in genetic architecture of complex phenotypes? PLoS ONE 7(12):e47371
Watts R et al (2003) Fiber tracking using magnetic resonance diffusion tensor imaging and its applications to human brain development. Ment Retard Dev Disabil Res Rev 9(3):168–177
White NS et al (2013) Probing tissue microstructure with restriction spectrum imaging: histological and theoretical validation. Hum Brain Mapp 34(2):327–346
Yoon U et al (2011) Genetic and environmental influences on structural variability of the brain in pediatric twin: deformation based morphometry. Neurosci Lett 493(1–2):8–13
Funding
Data used in the preparation of this article were obtained from the Adolescent Brain Cognitive DevelopmentSM (ABCD) Study (https://abcdstudy.org), held in the NIMH Data Archive (NDA). This is a multisite, longitudinal study designed to recruit more than 10,000 children age 9–10 and follow them over 10 years into early adulthood. The ABCD Study® is supported by the National Institutes of Health and additional federal partners under award numbers U01DA041048, U01DA050989, U01DA051016, U01DA041022, U01DA051018, U01DA051037, U01DA050987, U01DA041174, U01DA041106, U01DA041117, U01DA041028, U01DA041134, U01DA050988, U01DA051039, U01DA041156, U01DA041025, U01DA041120, U01DA051038, U01DA041148, U01DA041093, U01DA041089, U24DA041123, U24DA041147. A full list of supporters is available at https://abcdstudy.org/federal-partners.html. A listing of participating sites and a complete listing of the study investigators can be found at https://abcdstudy.org/consortium_members/. ABCD consortium investigators designed and implemented the study and/or provided data but did not necessarily participate in the analysis or writing of this report. This manuscript reflects the views of the authors and may not reflect the opinions or views of the NIH or ABCD consortium investigators.
Author information
Authors and Affiliations
Contributions
RW: conception, data analysis, and writing; LR: data analysis, and writing; JG: writing, review and editing; CGF: conception, writing, review, and editing.
Corresponding author
Ethics declarations
Competing interest (include appropriate disclosures)
Richard Watts, Lydia Rader, Justin Grant and Christopher G. Filippi these authors report no conflicts of interest.
Ethics approval (include appropriate approvals or waivers)
The anonymized data used in the preparation of this article (secondary data analysis) were obtained from the Adolescent Brain Cognitive DevelopmentSM (ABCD) Study (https://abcdstudy.org), held in the NIMH Data Archive (NDA). Procedures of the ABCD study were approved by a central Institutional Review Board at the University of California, San Diego, and in some cases by local IRB.
Consent to participate (include appropriate consent statements)
Parents or guardians provided written informed consent, and children assented to their participation.
Consent for publication (consent statement regarding publishing an individual’s data or image)
Not Applicable.
Additional information
Handling Editor: Sylia Wilson.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Watts, R., Rader, L., Grant, J. et al. Genetic and Environmental Contributions to Subcortical Gray Matter Microstructure and Volume in the Developing Brain. Behav Genet 53, 208–218 (2023). https://doi.org/10.1007/s10519-023-10142-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10519-023-10142-1