Abstract
Breeding of ornamental marine fishes was initiated in Makassar, South Sulawesi, Indonesia, to protect reef ecosystems from destructive fishing and to supplement income of local communities. However, stress associated with captive breeding conditions could enhance vulnerability of fish to disease. Wild- and captive-bred clownfish Amphiprion percula showed clinical signs such as white spots and warts typically associated with lymphocystis disease. Sequencing a portion of the DNA polymerase gene confirmed the identity of a lymphocystis disease virus (LCDV) in clownfish tissues that closely resembled LCDV from paradisefish; however, both strains were only 90% similar at the nucleotide level. Phylogenetic analysis suggests the LCDV is a new strain representing the first sequence of LCDV in A. percula. To determine the virus presence in clownfish and other ornamental marine species (OMS) in the hatchery, we developed conventional PCR and loop-mediated isothermal amplification (LAMP) assays that were comparatively sensitive at detecting the new LCDV strain. Among the PCR-tested clownfish, 19.4% (n = 14/72) were positive for the LCDV DNA that were either symptomatic or apparently healthy. Other OMS (n = 29) bred in the hatchery including seahorse Hippocampus spp., mandarin fish Synchiropus spp., and pajama cardinal fish Sphaeremia nepatoptera were negative by the LCDV PCR. The PCR and LAMP assays developed in this study will facilitate disease management by early detection of the LCDV and removal of potential reservoirs of the virus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The capture of wild ornamental marine species (OMS) provides short-term but an important source of livelihood in Indonesia (Reksodihardjo-Lilley and Lilley 2007). Recurring destructive fishing practices, however, such as blast (dynamite) and cyanide fishing, have severely damaged coral reefs (Williams et al. 2019) and have endangered other wildlife in the region (Halim 2002). Thus, prospects for environmentally sustainable fishery resources are vital such as aquaculture although OMS breeding is poorly understood compared with ornamental freshwater fishes (Tlusty 2002). Among the 100 OMS species bred in captivity (Dawes 1998), only 21 species are deemed commercially profitable (Schiemer 2001).
Stress factors associated with intensive culture conditions such as water quality, handling, stocking density, and nutrition promote the vulnerability of fish to disease-causing pathogens (Hedrick 1998). Fish collected from highly polluted coastal waters harbor excessive pathogen loads that may lead to diseases in captivity (Lafferty et al. 2015). Parasitic, bacterial, viral, and fungal pathogens have been reported as agents of diseases in ornamental fish production (Fioravanti and Florio 2017). Lymphocystis disease virus (LCDV) can threaten ornamental fish through disease and loss of profit due to the warty appearance of fish (Yanong 2013). Our study is focused on LCDV as there are no cost-effective treatments against viral infections (e.g., Assefa and Abunna 2018). Lymphocystis is an iridovirus (DNA virus) broadly documented to occur in many freshwater and marine fish around the world in both wild and farmed fish (e.g., Sahoo and Goodwin 2012; Yanong 2013; Borrego et al. 2017). Typical signs of lymphocystis-infected fish are mostly external nodular, wart-like growths on the fins, skin, or gills that can advance from chronic to systemic infections (Yanong 2013). Lymphocystis disease has only been identified by histopathology and electron microscopy as reported in common clownfish, Amphiprion ocellaris (Pirarat et al. 2011). Thus far, there has been no genetic information on LCDV in clownfish. Studies to address this knowledge gap are relevant as the two clownfish species A. percula and A. ocellaris are among the top five species in the aquarium industry (Wabnitz et al. 2003; Rhyne et al. 2012).
In high-density culture environments, LCDV can be horizontally transmitted from fish surviving the viral infection and remain carriers of the disease (McDermott and Palmeiro 2013; Fioravanti and Florio 2017). Early detection of LCDV is therefore important to minimize disease transmission and spread through identification and removal of carriers as reservoirs of infection. One method of pathogen detection is loop-mediated isothermal amplification (LAMP) assay, which employs a DNA amplification technique with DNA polymerase, 4–6 primers, autocycling strand displacement, and isothermal incubation temperature (Notomi et al. 2000). Unlike conventional and real-time PCR, the LAMP assay with colorimetric detection eliminates the need for expensive laboratory equipment such as a thermocycler, gel electrophoresis, and gel documentation system. The LAMP assay is comparatively sensitive as conventional PCR with shorter reaction time (Notomi et al. 2000; Ranjbar and Afshar 2015), providing rapid and sensitive disease diagnosis. Various LAMP assays have been developed for the detection of several iridoviruses and other fish disease agents (Biswas and Sakai 2014).
Mars Symbioscience Indonesia conducted feasibility projects from 2009 to 2011 to engage coastal community residents in Badi Island, South Sulawesi, on captive breeding of the seahorse Hippocampus barbouri (Williams et al. 2014). Breeding efforts subsequently included another species of seahorse (H. kuda), the orange clownfish Amphiprion percula, two species of mandarin fish Synchiropus picturatus (spotted mandarin) and S. splendidus (mandarin dragonet), and pajama cardinal fish Sphaeremia nepatoptera. During captive breeding from 2015 to 2017 at Takalar hatchery, some of these species showed various infections as reported here. The purpose of our study is to describe a new LCDV strain in wild- and captive-bred clownfish A. percula in South Sulawesi, Indonesia, and to assess the virus prevalence among captive-bred OMS by developing conventional PCR and LAMP assays. We identified the LCDV by sequencing a portion of the DNA polymerase gene and determined its phylogenetic relationship with other LCDV strains. Early and specific detection of untreatable viral agent is relevant to ensure fish health in culture conditions where infectious diseases are highly likely to occur.
Materials and methods
Disease and mortalities
New clownfish broodstock (F0) were introduced to the breeding facility in February 2015 from Balai Laut Ambon (Ambon Marine Aquaculture Center) while F1 generations were bred in the hatchery from mixed broodstock sourced from various locations in Indonesia including Balai Laut Ambon and Balai Laut Takalar. Tumors, warts, and white spots were observed in adult clownfish including chronic fin rot among larvae and broodstock (Fig. 1, Table S1). Quarantine procedures were initiated in September 2015; however, marine ich, fin rot, rapid breathing, and flared operculum ensued in clownfish throughout 2015 until 2017 (Table S1) with increased mortalities (Fig. S1). Fish were also screened for presence of external parasites or bacteria from tissue scrapes (i.e., skin, gill/operculum, fins) using light microscopy. Kidney/spleen of clinically infected clownfish were aseptically inoculated onto Brain Heart Infusion Agar (BHIA) prepared with filtered sterile seawater; dominant representative colonies that grew on BHIA were processed for genomic DNA extraction using Qiagen Blood and Tissue Kit, PCR amplified with EUBA and EUBB primers for bacterial 16S rRNA genes (Buller 2004; Austin 2011), and submitted for sequencing at UC Davis (http://dnaseq.ucdavis.edu/).
Clownfish Amphiprion percula showing symptoms and presence of the lymphocystis disease virus (LCDV) as confirmed by the LCDV PCR developed in this study. a F1 fish (case 070316-1) with whitish wart (red circle) on dorsal fin from which the LCDV was originally detected, from Balai Laut Takalar (Takalar Marine Aquaculture Center, Indonesia). b F0 fish (case 170215, LCDV-positive wart) showing severe warts (red circle) on the body surface, from Balai Laut Ambon (Ambon Marine Aquaculture Center). c F1 fish (case 070316-2, LCDV-positive fin) with severe fin rot and lesions on the caudal fin, from Balai Laut Takalar. The grid indicates the estimated size of the fish image: 1 grid = 1 cm
Water quality in breeding facility
A continuous water flow-through system was in place at the Takalar hatchery during the study. Incoming ocean water was pumped through an extended intake pipe; filtered through a series of mechanical filters; stored in a 50-ton reservoir; and passed to sand filtration, microfiltration, and finally to a UV sterilizer system (Pro Pond Advantage UV 110 W) prior to water inflow into breeding tanks. Water quality parameters were monitored daily including water temperature using minimum-maximum thermometer (Sper Scientific Direct), salinity (ppt) as measured with a hand refractometer (Atago Co., Ltd), and pH with HANNA pHep Tester (Hanna Instruments, Inc.). Dissolved oxygen was measured using HACH LDo101 (Hach), while ammonia and nitrate levels were measured with Sera test kits (Sera). Water temperature in larval tanks was maintained at 28–30 °C using water heaters (Resun). Supplemental figures show the profile of average water quality from 2015 to 2017 (Fig. S2).
Fish samples and genomic DNA extraction
Fish were sampled across tanks and life stages; anesthetized with 50 ppm tricaine methane sulfonate (Argent); examined for external signs, bacteria, and parasites as above; and collected tissues for DNA extraction. General health assessment showed clownfish exhibiting flared operculum, fin rot, white spots, and warts (Fig. 1). For genomic DNA extraction, various OMS were sampled: clownfish (n = 72) that were apparently healthy or symptomatic (Table S1), seahorses (n = 24), mandarin fish (n = 4), and pajama cardinal fish (n = 1) (Table S2). The tissues that were collected from these fish include pieces of fins, gills, operculum, skin with warts or white spots, and other affected tissues (chin, snout) as well as internal organs (spleen, kidney, liver) (Table S1, Table S2). For larval samples, the whole fish was processed for genomic DNA extraction. Eggs (n = 10/broodstock pair) from three pairs of clownfish broodstock (male and female/pair) were processed as one sample per pair. Mysids (n = 3 at 5 mysids/pool) were also sampled and extracted for genomic DNA (Table S2).
If possible, fresh tissue was used for genomic DNA extraction; alternatively, whole fish samples were stored in 75% ethanol for DNA extraction of tissue using the Qiagen Blood and Tissue Kit. Genomic DNA were analyzed for the presence or absence of lymphocystis disease virus (LCDV) using the LCDV PCR as described below (see “Assessing conventional PCR and LAMP for sensitivity of LCDV detection”).
Sequencing a portion of the DNA polymerase gene from LCDV
Clownfish fin with white spots (Fig. 1a) was initially PCR-tested using a set of degenerate primers HV and Cons lower (Table 1) that amplify a portion of DNA polymerase gene from iridoviruses and herpes viruses (Hanson et al. 2006). The PCR reaction mixture contained 1× High Fidelity PCR buffer, 2.0 mM MgSO4, 0.2 mM dNTP mixture, 0.2 μM forward primer and reverse primer, and 1 U/rxn Platinum Taq DNA Polymerase High Fidelity (Invitrogen Life Technologies). Cycling conditions included 95 °C for 5 min, 35 cycles at 95 °C for 30 s, 45 °C for 30 s and 72 °C for 1 min and 30 s, and 72 °C for 5 min.
The PCR products were electrophoresed in 1.5% agarose gel and visualized (Wide Mini-Sub Cell GT Cell, Bio-Rad). Electrophoresis used 100 V for 30 min in 1× TAE buffer followed by staining in 1× GelRed dye (Phoenix Research) for 30 min. GelDoc XR (Bio-Rad) was used for gel imaging; PCR products with strong bands were identified and purified using DNA Clean and Concentrator-5 Kit (Zymo Research) for cloning reactions. The PCR-amplified DNA fragment (646 bp) was cloned into a vector (TOPO TA cloning kit pCR 4 vector, Thermo Fisher Scientific), followed by sequencing with M13 forward and reverse primers (Thermo Fisher Scientific). Sequencing reaction was performed by UC Davis (http://dnaseq.ucdavis.edu/).
Loop-mediated isothermal amplification primer design and conditions
The LCDV DNA polymerase gene from the clownfish fin tissue (Fig. 1a) was used to design the inner and outer loop primers for the LAMP assay (Table 1) using PrimerExplorerV5 (https://primerexplorer.jp/lampv5/index.html) (Eiken Chemical Co., Ltd.). The LAMP reaction mixture contained 1× isothermal amplification buffer, 8 mM MgSO4, 1.4 mM dNTP mix, 1.6 μM FIP/BIP primers, 0.2 μM F3/B3 primers, 0.4 μM LF/LB primers, and 320 U/mL Bst 2.0 DNA Polymerase Warmstart in 25 μL reaction volume (Bst 2.0 WarmStart DNA Polymerase, New England BioLabs Inc.). Reaction tubes were incubated at 65 °C for 1 h then 80 °C for 5 min in SimpliAMP thermal cycler (Thermo Fisher Scientific).
LAMP products were visualized after electrophoresis (0.8% gel) as described above with positive and negative controls. Positive controls were genomic DNA samples from which the LCDV was originally sequenced (case no. 120415-1 and 070316-1) while negative controls were nuclease-free water samples treated similarly as fish tissue from collection through extraction (method control).
Assessing conventional PCR and LAMP for sensitivity of LCDV detection
Plasmid DNA harboring a portion of clownfish LCDV DNA polymerase gene (199 bp; GeneArt, Thermo Fisher) was used for 1:10 serial dilutions of DNA standards to compare PCR and LAMP on detection limit of the LCDV. We designed a total of 9 LAMP primer sets (data not shown) and ran LAMP reactions with the serially diluted DNA standards and compared the results. The primer sets 13 and 14 produced DNA band at lower copy numbers of the target gene with higher signal intensity compared with the other primer sets; hence, we selected the primer set (Table 1) for amplifying the LCDV DNA in our study. Conventional PCR used the LCDV outer LAMP primers: primer 13_F3 as forward and primer 14_B3 as reverse primer (Table 1). The PCR reaction mixture contained 1× PCR buffer, 1.5 mM MgCl2, 0.2 mM dNTP mixture, 0.8 μM forward primer and reverse primer, and 1 U/rxn Platinum Taq DNA polymerase (Invitrogen Life Technologies). Cycling conditions included 95 °C for 5 min, 30 cycles at 95 °C for 30 s, 58 °C for 30 s and 72 °C for 30 s, and 72 °C for 5 min. The LAMP assay followed the reaction steps as described above, products were gel-electrophoresed, GelRed-stained, and imaged. Colorimetric LAMP used a colorimetric kit (New England Biolabs) following the manufacturer’s instructions and cycling conditions like the non-colorimetric LAMP assay.
LCDV detection in fish tissue
The genomic DNA of tissues from clownfish and other OMS (Table S1, Table S2) were screened for the LCDV DNA using the conventional PCR assay established in this study. Furthermore, the detection of the LCDV DNA in fish tissues was compared using conventional PCR and LAMP assay using the primers (Table 1) and conditions described above. Samples with good DNA quality (> 1 ng/μL DNA) were selected for the comparison (Table S3).
Sequence similarity searches and phylogenetic analysis
A portion of the LCDV DNA polymerase sequence from the Indonesian clownfish was used for similarity searches by the BLAST programs (Altschul et al. 1990). In addition, the sequence was used to generate a phylogenetic tree with other LCDV strains (Table 2) as follows: deduced amino acid sequences of DNA polymerase sequences from various LCDVs were aligned with MAFFT software ver. 7.427 (algorithm: L-INS-i) (Katoh et al. 2002). Based on the alignment of DNA polymerase protein sequences, gaps were inserted in the corresponding original nucleotide sequences using a custom perl script. The obtained nucleotide alignment was then used to generate a phylogenetic tree using MrBayes program ver. 3.2.6 with the following settings: substitution model: Hasegawa-Kishono-Yano (HKY); rate variation: gamma; outgroup: MG791866.1; gamma categories: 4; number of chains: 4; number of generations: 25,000,000; temperature: 0.5; subsampling freq: 500; burn-in: 25% (Huelsenbeck and Ronquist 2001). The jModelTest ver. 2.1.10 selected the HKY85 model and gamma rate variation as the best model for the dataset (Guindon and Gascuel 2003; Darriba et al. 2012).
Results
Disease and mortalities
Infected clownfish showed clinical symptoms such as white spots or small wart on dorsal fin from which the LCDV was originally detected (Fig. 1a). Fully developed warts on the body surface were also present (Fig. 1b) including severe fin rot and lesions on the caudal fin (Fig. 1c).
Bacteria and parasites were observed on light microscopy of scrapes from fin rots and gill tissues of clownfish across life stages (Table S1) in addition to LCDV data (Fig. 1, Tables 3 and 4, Table S1). Marine “ich,” Cryptocaryon irritans, was prevalent in clownfish and in seahorses (Table S2). Gene sequencing of DNA from dominant bacterial colonies isolated from diseased clownfish showed Gram-negative bacteria such as Pseudomonas, Pseudoalteremonas, Vibrio, Enterovibrio, and Photobacterium and Gram-positive bacteria such as Staphylococcus and Arthrobacter. Numerous clownfish succumbed to infections in November 2016; rearing to marketable size in 2017 showed even greater mortalities with increased disease monitoring (Fig. S1).
Water quality in breeding facility
The mean water temperature in all 3 years ranged from 28 to 31 °C throughout November to June, decreasing slightly from June to October (Fig. S2). While clownfish prefer 24–28 °C (Moorhead and Zeng 2010), water temperatures were maintained at 28–31 °C for optimum growth of OMS bred at the Takalar hatchery. The pH and salinity were at 8.1–8.2 and 33.5–34.2, respectively; pH spiked to 8.4 and nitrate levels increased to 1.25 ppm in September 2016; ammonia levels were generally above 0.02 mg/L (Fig. S2) deemed sub-optimal for fish health (Moorhead and Zeng 2010).
Amplification and sequencing of LCDV DNA polymerase gene
The Hanson primers (Hanson et al. 2006) amplified a DNA band in two clownfish samples at the expected size for iridovirus (ca. 700 bp). The samples were F0 fish with fin rot and F1 fish with fin tumor (Fig. 1a), each produced a 636-bp sequence (GenBank accession no. MN123250, Table 2) with high sequence similarity to DNA polymerase gene from Lymphocystis sp. The BLASTN search showed a query cover of 99% and identity match of 90% to LCDV-PF from paradise fish Macropodus opercularis (Table 5). This sequence was used to design LAMP primers that amplified the LCDV gene fragment by the LAMP and PCR, using the two LCDV-positive fish as positive controls. Nuclease-free water was used for negative control as fish with no history of viral infection was not available in the facility. Based on results of the LCDV PCR (Table 3, Table S1, Table S2), clownfish and other OMS were negative for the virus suggesting that amplification of the host gene(s) is likely very low.
Sensitivity of conventional PCR and LAMP for LCDV detection
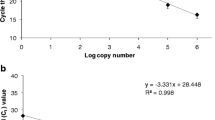
Using the 10-fold dilutions of LCDV plasmid DNA standard as template, the conventional PCR amplified a 200-bp product (Fig. 2a) while the LAMP assay showed intense streaking (Fig. 2b). The conventional PCR and LAMP assay detected the LCDV at 100 and 1000 gene copy numbers per reaction, respectively (Fig. 2a, b). The colorimetric LAMP detected the LCDV at 100 gene copy numbers per reaction (Fig. 2c).
Sensitivity of the PCR and loop-mediated isothermal amplification (LAMP) assays for detection of the lymphocystis disease virus (LCDV) from clownfish Amphiprion percula in Indonesia using primers designed in this study. Limit of detection is indicated by a white triangle. a Conventional PCR detects the 200-bp LCDV-amplified product at 100 gene copy numbers. b LAMP detects the LCDV at 1000 gene copy numbers as shown in gel streaking of products. c Colorimetric LAMP detects the LCDV at 100 gene copy numbers. 1 kb: 1 kb plus DNA ladder. −: negative control (DNase-free water). +(1): positive control of DNA from fin tumor (case no. 120415-1). +(2): positive control of the LCDV PCR product. Numbers 0 to 8 indicate the 1:10 serial dilution of LCDV plasmid DNA to create standard curves that correspond to 10× gene copy numbers of LCDV per microliter of sample
Gel electrophoresis of LAMP products from strongly positive samples and standards with 1 × 106 or higher gene copy numbers showed bright smears that may have overexposed the gel, making the smears from the 1000 and 100 gene copy number standards dim (Fig. 2b). The colorimetric detection of LAMP products produced distinct results for positive (yellow-orange PCR solution) and negative (pink PCR solution) samples (Fig. 2c).
LCDV detection by conventional PCR
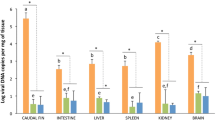
Among the 72 clownfish that were screened for the LCDV DNA, both symptomatic and apparently healthy fish were LCDV PCR-positive (Table 3, Table S1). Out of the 45 unhealthy fish, 20.0% were LCDV-positive while among the 24 healthy fish, five fish or 20.8% were LCDV-positive (Table 3). These five fish were in the same tank holding ca. 100 juvenile fish from different broodstock. Not all symptomatic fish tested positive for the LCDV. Of fish exhibiting fin rot (n = 22), 9.1% tested positive for the LCDV. Among fish exhibiting warts (n = 6) and white spots (n = 3), 50.0% and 33.3% were positive for the LCDV, respectively (Table 3). From fish exhibiting various symptoms without skin warts (n = 12), 25% tested positive for the LCDV. Disparity on LCDV presence was observed among the clownfish generations. Among the F0 broodstock sampled (n = 8), 37.5% were positive for the LCDV while for F1 fish (n = 61), 18.0% were positive for the LCDV (Table 4).
Although most of the LCDV PCR-positive clownfish had warts and tumors (Table 3), the LCDV DNA was also detected in the spleen of fish whose fin tumor also tested positive for the LCDV (case # 070316-1, Table S1). From the three broodstock pairs sampled, the 10 eggs from each pair and processed as a single sample/pair were all negative for LCDV DNA by conventional PCR (Table 3, Table S1). All other OMS that were PCR-tested were negative for the LCDV DNA (Table S2). Mysids (n = 3, 5 mysids/sample) were also negative for the LCDV DNA (Table S2).
The LAMP assay showed comparative sensitivity as the conventional PCR assay for LCDV detection in clownfish. Fish positive and negative for the LCDV DNA matched the results of the LAMP assay and conventional PCR (Table S3).
Phylogenetic analyses
The clownfish LCDV in our study was placed in clade I with other lymphocystis viruses found in marine fish species as shown in the phylogenetic tree generated from DNA polymerase genes (Fig. 3). The clownfish LCDV in Indonesia is most closely related to LCDV from paradise fish in China with high sequence similarities at the nucleotide and deduced amino acid levels (90.0 and 91.2% similarity, respectively) (Fig. 3, Table 5). The partial DNA polymerase gene distinctly separated the clade for LCDV-Clownfish-Indonesia and LCDV-ParadiseFish-China from the clade of LCDV-SeaBream with a high posterior probability (1.00) (Fig. 3).
Phylogenetic relationship of the lymphocystis disease virus (LCDV) from clownfish Amphiprion percula in Indonesia with other LCDV strains. The phylogenetic tree was generated by MrBayes program ver. 3.2.6 using DNA polymerase nucleotide sequences of LCDV strains. Posterior probabilities are shown at each node. The scale bar indicates the estimated substitutions per site. Ranavirus from black-spotted pond frog (Rana nigromaculata) was used as an outgroup
Discussion
Pathogens causing diseases are commonly found in natural environments and can become problematic once introduced into stressful aquaculture conditions (Lafferty et al. 2015). In our study, various infections observed in captive-bred clownfish and other OMS are mainly associated with bacteria and protozoans. While disease associated with these organisms can be managed with various treatment options, combating viral infections is challenging due to the lack of effective chemical treatments against viral agents (e.g., Assefa and Abunna 2018). Viral infections can reduce the production of healthy fish for sustainable aquaculture of OMS (Sivasankar 2017). For this reason, the focus of our study was to assess virus infections encountered in the captive breeding of OMS and to develop the tools for specific detection of the viral agent(s).
Clownfish exhibited white spots, warts, and tumors that led to the presumptive identification of lymphocystis disease virus (LCDV). Gene sequencing using a portion of the DNA polymerase gene identified the virus as LCDV at 91.2% sequence similarity (amino acid level) with the DNA polymerase of paradisefish LCDV (Table 5). However, further investigation is needed to confirm whether the LCDV strain from clownfish is different from the LCDV in paradisefish. Specific guidelines on cutoff value for species differentiation in iridoviruses that include the LCDV, based on DNA polymerase and/or other gene markers, are currently lacking. As such, although the LCDV identified from the Indonesian clownfish likely belongs to a new species, whole-genome sequencing and electron microscopy are needed for accurate taxonomic classification of the virus. Using a portion of the DNA polymerase gene addressed our core objective of describing the identity and prevalence of the new LCDV strain in clownfish and other OMS in the breeding facility. Furthermore, the partial DNA polymerase gene adequately established the genetic relationship of the clownfish LCDV with other known LCDV strains (i.e., phylogenetic tree). Several studies have used a portion of DNA polymerase gene sequence for screening and initial identification of LCDV (Hanson et al. 2006; Xu et al. 2014).
To the best of our knowledge, the LCDV strain that we found in the Indonesian clownfish represents the first sequence of LCDV in A. percula. Previous LCDV strains in clownfish were identified using other methods of virus characterization (Pirarat et al. 2011; Hick et al. 2016; Borrego et al. 2017). The complete genome of LCDV-1 was first reported from European flounder Platichthys flesus (Tidona and Darai 1997) then in Japanese flounder Paralichthys olivaceus (Zhang et al. 2004). LCDV has also been found in Chinese flounder, paradise fish, rockfish, and gilthead seabream (Zhang et al. 2003; Kitamura et al. 2006; Xu et al. 2014; Valverde et al. 2017). Our phylogenetic analysis using various LCDV strains suggests that the LCDV from the Indonesian clownfish is a new strain of LCDV but very closely related to paradise fish LCDV.
The other OMS in the breeding facility were negative for the LCDV suggesting that the host range of the clownfish LCDV strain is limited. Further investigation is needed to confirm whether the clownfish LCDV strain is host-specific to clownfish. Previous studies indicate that LCDV is host-specific although geographically cosmopolitan in distribution (Kitamura et al. 2006; Hossain et al. 2008; Yan et al. 2011).
We demonstrated that even apparently healthy clownfish harbored the LCDV. The five generally healthy juvenile fish that were LCDV-positive were from a group of ca. 100 fish suggesting that the LCDV was horizontally transmitted (fish to fish) among fish sharing the same tank. Iridovirus can spread among fish in the holding container (Kurobe et al. 2011). Furthermore, clownfish that succumbed to disease showed the presence or absence of LCDV, bacteria, and marine “ich” Cryptocaryon irritans suggesting that these pathogens, together, contributed to disease and mortalities.
Although we did not detect the LCDV DNA from broodstock eggs, LCDV could be vertically transmitted via egg surface of carrier parents (Cano et al. 2013). Our study design did not include assessment of the LCDV transmission. The clownfish breeding protocol at the Takalar hatchery did not separate progenies from the parent broodstock for certain periods, and in this setting, the LCDV was likely transmitted through direct contact of LCDV-positive fish with healthy fish or via direct ingestion of feces or food contaminated with the LCDV. Horizontal fish to fish transmission of disease typically occurs through direct ingestion of contaminated food particles and feces found in the water (e.g., Baxa-Antonio et al. 1992). The mysids that were fed to OMS were negative for the LCDV DNA suggesting they are not potential reservoir of the virus at least from the mysid population examined for the LCDV PCR.
Due to the detection of the LCDV in clownfish across life stages and health status, assessing the virus presence among sick as well as in apparently healthy fish in the hatchery was deemed relevant. We demonstrated that non-lethal sampling and the LCDV PCR testing of fin clips and skin scrapes of fin rots and lymphocytis lesions successfully detected the presence or absence of the LCDV. The LCDV PCR screening informed the prevalence of fish harboring the virus within suspect populations. Removal of LCDV-positive fish may help to diminish likely reservoirs of the virus in rearing tanks.
Following disease outbreaks in the hatchery, decreased egg production (data not shown) and sudden death among clownfish juveniles and adults ensued in the absence or presence of the LCDV. Bacterial infections and parasites that occurred at the time of the LCDV outbreak combined with elevated water temperatures, and high concentrations of ammonia and nitrates in 2015 and 2016, potentially contributed to clownfish stress that led to mortalities. The first disease outbreak occurred after the arrival of new broodstock, which likely introduced the virus and other pathogens from polluted marine environments (Lafferty et al. 2015). Interestingly, the F0 wild broodstock had a higher percentage of LCDV-positive (37.5%) compared with F1 apparently healthy adults (14%) suggesting that F0 wild broodstock are potential reservoirs of the LCDV in the culture facility over time (Valverde et al. 2017).
The LAMP assay showed comparable sensitivity as the conventional PCR on the LCDV detection in symptomatic and asymptomatic fish indicating that both methods could sufficiently detect the LCDV even in carrier state. The LAMP method can be adapted for on-site field-testing at fish farms or culture facilities along coastal Sulawesi without access to pathogen identification. The PCR and LAMP assays that we developed will aid in the detection of the LCDV at critical phases of OMS breeding: (1) LCDV screening of new broodstock and their progeny; (2) quarantine of infected fish from other healthy fish; and (3) screening of various OMS, live diets, and other potential reservoirs of the LCDV.
Although quarantine and disease screening likely helped in minimizing the LCDV spread among stock fish, total elimination of the virus in the OMS populations is not feasible. Good husbandry can help to reduce disease outbreaks that cause loss of expensive fish stocks and progeny. More importantly, the new feeding regimes containing enriched algae and rotifers improved fish nutrition. These changes in fish care significantly reduced the development of warts and white spots as well as decreased the incidence of mortalities associated with bacterial infections. Mediating risk factors that enhance the transmission and progress of viral infections is critically important for sustaining fish health in captivity (Stevens et al. 2017).
As only a few OMS are bred using aquaculture technology (Dawes 1998; Schiemer 2001), studies that focus on species-specific biology can help to improve broodstock, larval production, and nutrition (Tlusty 2002). Discovering breeding technologies for OMS that are less intricate to breed can be collaborated with academic research (Pouil et al. 2019). To date, clownfish are deemed the most abundant species traded in the industry (Maison and Graham 2016). Although captive breeding is costly and untenable in the long term for fish farmers (Williams et al. 2014), sustainably sourced OMS support environmentally friendly practices and supplement livelihoods of coastal communities (Williams et al. 2014; Militz et al. 2017; Marchio 2018; King 2019). The benefits of OMS breeding compared with wild harvesting deemed detrimental to the environment, due to destructive fishing practices and overfishing, must be balanced in a broader context (Tlusty 2002).
Our study underscores the importance of partnership between academic research (University of California, Davis) and business enterprise (Mars Symbioscience) to inform the relevance of disease screening in OMS breeding. Developing the tools for detection of causative agents is vital for health assessment of OMS that are vulnerable to disease under stressful breeding conditions. Early detection of pathogens prior to disease onset is essential for the viability of OMS that undergo a supply chain from breeding facilities to business retailers to consumers.
Conclusion
We discovered a unique LCDV strain in captive-bred clownfish in Indonesia representing the first gene sequence of LCDV in clownfish. The LCDV strain in our study was detected by PCR from 14 of 69 clownfish tested (20%) on the skin and fin tissues of symptomatic and asymptomatic fish including wild broodstock (F0) and F1 generation. Eggs from clownfish broodstock sourced from various locations in Indonesia including other OMS bred in the facility were all negative for LCDV. The conventional PCR and LAMP assays developed in this study will facilitate LCDV disease management via early detection of the virus and removal of LCDV-positive fish as reservoirs of contamination and transmission. Although intensive breeding conditions may enhance fish vulnerability to stress and disease, good husbandry, early detection of pathogens causing diseases, and improved nutrition are useful measures to address fish health management aimed at attaining sustainable OMS production.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Assefa A, Abunna F (2018) Maintenance of fish health in aquaculture: review of epidemiological approaches for prevention and control of infectious disease of fish. Vet Med Int 2018:1–10. https://doi.org/10.1155/2018/5432497
Austin B (2011) Taxonomy of bacterial fish pathogens. Vet Res 42(1):20. https://doi.org/10.1186/1297-9716-42-20
Baxa-Antonio D, Groff JM, Hedrick RP (1992) Experimental horizontal transmission of Enterocytozoon salmonis to Chinook salmon, Oncorhynchus tshawytscha. J Protozool 39:699–702
Biswas G, Sakai M (2014) Loop-mediated isothermal amplification (LAMP) assays for detection and identification of aquaculture pathogens: current state and perspectives. Appl Microbiol Biotechnol 98:2881–2895. https://doi.org/10.1007/s00253-014-5531-z
Borrego JJ, Valverde EJ, Labella AM, Castro D (2017) Lymphocystis disease virus: its importance in aquaculture. Rev Aquac 9:179–193. https://doi.org/10.1111/raq.12131
Buller NB (2004) Bacteria from fish and other aquatic animals: a practical identification manual. https://doi.org/10.1079/9780851997384.0000
Cano I, Valverde EJ, Garcia-Rosado E, Alonso MC, Lopez-Jimena B, Ortiz-Delgado JB, Borrego JJ, Sarasquete C, Castro D (2013) Transmission of lymphocystis disease virus to cultured gilthead seabream, Sparus aurata L., larvae. J Fish Dis 36:569–576. https://doi.org/10.1111/jfd.12011
Darriba D, Taboada GL, Doallo R, Posada D (2012) JModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
Dawes J (1998) International experience in ornamental marine species management. Part 1: perspectives. Ornamental Fish Int J 26:1–3
Fioravanti ML, Florio D (2017) Common diseases in marine ornamental fishes. In: Calado R, Olivoto I, Oliver MP, Holt GJ (eds) Marine ornamental species aquaculture. Wiley, Hoboken, pp 347–380
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704. https://doi.org/10.1080/10635150390235520
Halim A (2002) Adoption of cyanide fishing practice in Indonesia. Ocean Coast Manag 45:313–323
Hanson LA, Rudis MR, Vasquez-Lee M, Montgomery RD (2006) A broadly applicable method to characterize large DNA viruses and adenoviruses based on the DNA polymerase gene. Virol J 3:1–10. https://doi.org/10.1186/1743-422X-3-28
Hedrick RP (1998) Relationships of the host, pathogen, and environment: implications for diseases of cultured and wild fish populations. J Aquat Anim Health 10:107–111. https://doi.org/10.1577/1548-8667(1998)010%3C0107:ROTHPA%3E2.0.CO;2
Hick PM, Becker JA, Whittington RJ (2016) Iridoviruses of fish (chapter 8). In: Kibenge FSB, Godoy M (eds) Aquaculture virology. Elsevier Inc, Amsterdam, pp 127–152
Hossain M, Song J-Y, Kitamura S-I, Jung S-J, Oh M-J (2008) Phylogenetic analysis of lymphocystis disease virus from tropical ornamental fish species based on a major capsid protein gene. J Fish Dis 31:473–479. https://doi.org/10.1111/j.1365-2761.2008.00917.x
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinform 17:754–755. https://doi.org/10.1093/bioinformatics/17.8.754
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
King TA (2019) Wild caught ornamental fish: a perspective from the UK ornamental aquatic industry on the sustainability of aquatic organisms and livelihoods. J Fish Biol 94:1–12. https://doi.org/10.1111/jfb.13900
Kitamura S-I, Jung S-J, Kim W-S, Nishizawa T, Yoshimizu M, Oh M-J (2006) A new genotype of lymphocystivirus, LCDV-RF, from lymphocystis diseased rockfish. Arch Virol 151:607–615. https://doi.org/10.1007/s00705-005-0661-3
Kurobe T, MacConnell E, Hudson C, McDowell TS, Mardones FO, Hedrick RP (2011) Iridovirus infections among Missouri River sturgeon: initial characterization, transmission, and evidence for establishment of a carrier state. J Aquat Anim Health 23:9–18. https://doi.org/10.1080/08997659.2011.545697
Kvitt H, Heinisch G, Diamant A (2008) Detection and phylogeny of Lymphocystivirus in sea bream Sparus aurata based on the DNA polymerase gene and major capsid protein sequences. Aquac 275:58–63
Lafferty KD, Harvell CD, Conrad JM, Friedman CS, Kent ML, Kuris AM, Powell EN, Rondeau D, Saksida SM (2015) Infectious diseases affect marine fisheries and aquaculture economics. Annu Rev Mar Sci 7:471–496
Lopez-Bueno A, Mavian C, Labella AM, Castro D, Borrego JJ, Alcami A, Alejo A (2016) Concurrence of iridovirus, polyomavirus, and a unique member of a new group of fish papillomaviruses in lymphocystis disease-affected gilthead sea bream. J Virol 90:8768–8779
Maison KA, Graham KS (2016) Status review report: orange clownfish (Amphiprion percula). NOAA Technical Memorandum NMFS-PIFSC-52. https://doi.org/10.7289/V5J10152
Marchio EA (2018) The art of aquarium keeping communicates science and conservation. Front Commun 3. https://doi.org/10.3389/fcomm.2018.00017
McDermott C, Palmeiro B (2013) Selected emerging infectious diseases of ornamental fish. Vet Clin North Am Exot Anim Pract 16:261–282. https://doi.org/10.1016/j.cvex.2013.01.006
Militz TA, Foale S, Kinch J, Southgate PC (2017) Consumer perspectives on theoretical certification schemes for the marine aquarium trade. Fish Res 193:33–42. https://doi.org/10.1016/j.fishres.2017.03.022
Moorhead JA, Zeng C (2010) Development of captive breeding techniques for marine ornamental fish: a review. Rev Fish Sci 18:315–343. https://doi.org/10.1080/10641262.2010.516035
Mou W, Geng Y, Ouyang P, Chen D, Huang X, Wang K, Zhao R, Bai M (2019) Isolation, identification and phylogenetic analysis of a ranavirus isolated from Rana nigromaculata. J South China Agric Univ 40:40–46. https://doi.org/10.7671/j.issn.1001-411X.201805012
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, Hase T (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28:63e–663e. https://doi.org/10.1093/nar/28.12.e63
Pirarat N, Pratakpiriya W, Jongnimitpaiboon K, Sajjawiriyakul K, Rodkhum C, Chansue N (2011) Lymphocystis disease in cultured false clown anemonefish (Amphiprion ocellaris). Aquac 315:414–416. https://doi.org/10.1016/j.aquaculture.2011.01.014
Pouil S, Tlusty MF, Rhyne AL, Metian M (2019) Aquaculture of marine ornamental fish: overview of the production trends and the role of academia in research progress. Rev Aquac 12:1217–1230. https://doi.org/10.1111/raq.12381
Ranjbar R, Afshar D (2015) Development of a loop-mediated isothermal amplification assay for rapid detection of Yersinia enterocolitica via targeting a conserved locus. Iran J Microbiol 7:185–190
Reksodihardjo-Lilley G, Lilley R (2007) Towards a sustainable marine aquarium trade: an Indonesian perspective. SPC Live Reef Fish Inf Bull 17:11–19
Rhyne AL, Tlusty MF, Schofield PJ, Kaufman L, Morris JA, Bruckner AW (2012) Revealing the appetite of the marine aquarium fish trade: the volume and biodiversity of fish imported into the United States. PLoS One 7:e35808. https://doi.org/10.1371/journal.pone.0035808
Sahoo PK, Goodwin AE (2012) Viruses of freshwater finfish in the Asian-Pacific region. Indian J Virol 23:99–105. https://doi.org/10.1007/s13337-012-0102-3
Schiemer G (2001) Captive-bred marine fish. Aquar Fish Mag, March, 41–45
Sivasankar P (2017) A review on DNA viral diseases of fish. Int J Fish Aquat Study 5:443–450
Stevens CH, Croft DP, Paull GC, Tyler CR (2017) Stress and welfare in ornamental fishes: what can be learned from aquaculture? J Fish Biol 91:409–428. https://doi.org/10.1111/jfb.13377
Tidona CA, Darai G (1997) The complete DNA sequence of lymphocystis disease virus. Virol 230:207–216. https://doi.org/10.1006/viro.1997.8456
Tlusty MF (2002) The benefits and risks of aquacultural production for the aquarium trade. Aquac 205:203–219
Valverde EJ, Borrego JJ, Sarasquete MC, Ortiz-Delgado JB, Castro D (2017) Target organs for lymphocystis disease virus replication in gilthead seabream (Sparus aurata). Vet Res 48:1–11. https://doi.org/10.1186/s13567-017-0428-3
Wabnitz C, Taylor M, Green E, Razak T (2003) From ocean to aquarium: the global trade in marine ornamental species. UNEP/WCMC, Cambridge https://www.unenvironment.org/resources/report/ocean-aquarium-global-trade-marine-ornamental-species. Accessed 12 Jul 2020
Williams SL, Janetski N, Abbott J, Blankenhorn S, Cheng B, Crafton RE, Hameed SO, Rapi S, Trockel D (2014) Ornamental marine species culture in the coral triangle: seahorse demonstration project in the Spermonde Islands, Sulawesi, Indonesia. Environ Manag 54:1342–1355. https://doi.org/10.1007/s00267-014-0343-6
Williams SL, Sur C, Janetski N, Hollarsmith JA, Rapi S, Barron L, Heatwole SJ, Yusuf AM, Yusuf S, Jompa J, Mars F (2019) Large-scale coral reef rehabilitation after blast fishing in Indonesia. Restor Ecol 27:447–456. https://doi.org/10.1111/rec.12866
Xu L, Feng J, Huang Y (2014) Identification of lymphocystis disease virus from paradise fish Macropodus opercularis (LCDV-PF). Arch Virol 159:2445–2449. https://doi.org/10.1007/s00705-014-2060-0
Yan XY, Wu ZH, Jian JC, Lu YS, Sun XQ (2011) Analysis of the genetic diversity of the lymphocystis virus and its evolutionary relationship with its hosts. Virus Genes 43:358–366. https://doi.org/10.1007/s11262-011-0646-0
Yanong RPE (2013) Lymphocystis disease in fish 1. IFAS Ext University Florida. 2013;FA181(November):1-4. https://edis.ifas.ufl.edu/pdffiles/FA/FA18100.pdf. Accessed 12 Jul 2020
Zhang QY, Ruan HM, Li ZQ, Yuan XP, Gui JF (2003) Infection and propagation of lymphocystis virus isolated from the cultured flounder Paralichthys olivaceus in grass carp cell lines. Dis Aquat Org 57:27–34. https://doi.org/10.3354/dao057027
Zhang QY, Xiao F, Xie J, Li ZQ, Gui J-F (2004) Complete genome sequence of lymphocystis disease virus isolated from China. J Virol 78:6982–6994. https://doi.org/10.1128/jvi.78.13.6982-6994.2004
Acknowledgments
We extend our gratitude to Mars Symbioscience staff including Indar Dewa, Muhammad Agam Alpharesy, Rosdiana, and Abdul Rahman for their support on hatchery phases of the project. Khiet Huynh at UC Davis Aquatic Health Program (AHP) is appreciated for his assistance on cloning procedures. We dedicate this work in honor of Professor Susan L. Williams for her invaluable contributions on conservation of ornamental marine species and coral reef rehabilitation in Indonesia. Professor Williams established the MOU on research collaboration for UC Davis; Mars Symbioscience; and Hasanuddin University in South Sulawesi, Indonesia, in 2014.
Funding
Agreement no. 201602101 and Marine Resources Grant from Mars Symbioscience and Mars Inc. provided funds to DVB with SVM–AHP team to investigate infectious diseases of marine resources, especially fish, development of diagnostic tools, and travels to MSS Indonesia. DVB received supplemental travel grants from UC Davis Academic Federation Committee. AHP funded the online publication of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed by the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lam, C., Khairunissa, I., Damayanti, L. et al. Detection of a new strain of lymphocystis disease virus (LCDV) in captive-bred clownfish Amphiprion percula in South Sulawesi, Indonesia. Aquacult Int 28, 2121–2137 (2020). https://doi.org/10.1007/s10499-020-00581-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10499-020-00581-6