Abstract
Outcomes of melanoma patient treatment remain unsatisfactory despite accessibility of oncoprotein-targeting drugs and immunotherapy. Here, we reported that 17-aminogeldanamycin more potently activated caspase-3/7 in BRAFV600E melanoma cells than geldanamycin, another inhibitor of heat shock protein 90 (HSP90). 17-aminogeldanamycin alleviated self-triggered compensatory increase in HSP70 mRNA level and induced endoplasmic reticulum (ER) stress, which was followed by selective diminution of cytoprotective IRE1α-XBP1s pathway activity of unfolded protein response (UPR), inhibition of ERK1/2 activity and induction of apoptosis. Concomitantly, ATF6/p50 level and expression of PERK-dependent genes, CHOP and BIM, remained unaltered. This might result from an inframe deletion in EIF2AK3 leading to a PERKL21del variant revealed by whole-exome sequencing in melanoma cell lines. 17-aminogeldanamycin exhibited similar activity in NRASQ61R melanoma cells that harbored a heterozygous inactivating variant of NAD(P)H:quinone oxidoreductase 1 (NQO1P187S). In addition, 17-aminogeldanamycin acted cooperatively with trametinib (an inhibitor of MEK1/2) and vemurafenib (an inhibitor of BRAFV600E) in induction of apoptosis in melanoma cell lines as evidenced by in-cell caspase-3/7 activation and PARP cleavage that occurred earlier compared with either drug used alone. As trametinib and vemurafenib did not significantly affect HSP70 and GRP78 transcript levels, cooperation of MEK/BRAFV600E inhibitors and 17-aminogeldanamycin might result from a concurrent inhibition of the RAS/RAF/MEK/ERK cascade and IRE1α-dependent signaling, and cell-intrinsic ER homeostasis can determine the extent of the drug cooperation. Our study indicates that 17-aminogeldanamycin takes several advantages compared with other HSP90-targeting compounds, and can complement activity of BRAF/MEK inhibitors in melanoma cells of different genetic subtypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genomic classification has been a gauge for clinical management of melanoma patients by using immunotherapy or targeted inhibitors of BRAFV600 and MEK1/2 [1]. However, lack of hot-spot BRAF, RAS, or NF1 driver mutations in the triple wild-type subtype accounting for 6–20% of melanomas [2, 3], and variability of phenotype of patient-derived melanoma cell lines representing the same genetic subtype [4] enforce combining both genetic and phenotypic traits to achieve more accurately stratification of melanoma patients. In addition, phenotype-based approaches can limit the number of potential therapeutic targets by pointing to master regulators of cell identity as demonstrated by selection of either MEK or HSP90, whose inhibition substantially affected 75% of melanoma cell lines [5]. Heat shock protein 90 (HSP90) is a molecular chaperone involved in a proper folding and multiprotein complex assembly of a myriad of client proteins including several oncoproteins [6, 7], whereas a membrane-bound HSP90 in dying cells facilitates activation of the immune clearance [8]. HSP90 is frequently overexpressed in cancer [6]. Accordingly, expression of HSP90 substantially increases from nevi to melanoma resulting in high HSP90 level in more than 50% of melanoma tumors, and augments with advanced melanoma stage [9, 10]. In addition, also serum levels of HSP90 are higher in melanoma patients than in healthy controls, with median values 49.76 ng/ml versus 27.07 ng/ml, respectively [11]. More interestingly, it has been demonstrated that HSP90 isoform present in melanoma-derived exosomes contributes to creation of a pre-metastatic niche by ‘educating’ bone marrow progenitors [12].
HSP90 predominantly exerts its function via N-terminal ATPase domain, thus preventing from ATP binding largely interferes with HSP90 activity [13]. Regarding a pleiotropic role of this chaperone, inhibition of HSP90 is associated with an accumulation of improperly folded client proteins, which is followed by induction of endoplasmic reticulum (ER) stress and unfolded protein response (UPR) governed by glucose-regulated protein 78/binding immunoglobulin protein (GRP78/BiP). UPR engages three pathways initiated by the GRP78/BiP release of inositol-requiring enzyme 1 alpha (IRE1α), protein kinase R-like endoplasmic reticulum kinase (PERK) and activating transcription factor 6 (ATF6). These pathways either restore cell homeostasis or promote cell death in case of an excessive proteotoxic stress [14]. In preclinical melanoma studies, structurally different inhibitors of HSP90 produced ER stress [15], induced apoptosis and reduced tumorigenicity of vemurafenib-resistant cells [16, 17], circumvented mitochondria biogenesis [18] and mitigated immunosuppressing activity of melanoma cells [19]. Combining XL888 (Exelixis), a non-benzoquinone ATP-competitive inhibitor of HSP90, with targeted inhibitors of the RAS/RAF/MEK/ERK (MAPK) signaling pathway (XL888 + vemurafenib, and XL888 + vemurafenib + cobimetinib) is currently evaluated in phase I clinical trials in patients with unresectable melanoma (clinicaltrials.gov). In a dose escalation trial of XL888 and vemurafenib combination, 15 out of 20 patients (75%) responded to the treatment with a median overall survival of 34.6 months [20]. Resistance to a combination of XL888 and BRAFV600 inhibitor has been recently linked to a CDK2high/MITFhigh phenotype of melanoma cells [21]. Concerning high protein levels of both MITF and CDK2 reported in five out of 12 melanoma cell lines [22] and the most significant correlation between MITF and CDK2 mRNA levels in melanoma tumor samples compared with other types of cancer [21], XL888 and BRAFV600 inhibitor combination is likely ineffective in a subset of patients. In the study by Azimi et al., it has been also demonstrated that the same melanoma cell line can exhibit a variable sensitivity to different HSP90 inhibitors [21]. It might result from dissimilar chemical structures of these compounds underlying execution of specific molecular effects as exemplified by BRAFV600E degradation exhibited by benzoquinone inhibitors of HSP90 [23]. Therefore, further research on inhibitors structurally unrelated to XL888 is of interest.
Geldanamycin, a natural benzoquinone inhibitor of the N-terminal ATPase activity of HSP90, was first purified from Streptomyces hydroscopicus, and has been a prototype of a class of anti-cancer agents [24]. Geldanamycin-induced toxicity and low solubility have limited its clinical use [24]. Its derivatives, 17-substituted geldanamycin analogues are less hepatotoxic [25, 26]. 17-aminogeldanamycin is a metabolic product of cytochrome P450 3A4 (CYP3A4)-dependent conversion of 17-N-allylamino-17-demethoxygeldanamycin (tanespimycin) [27], and exhibits higher water solubility than a parental compound, and higher affinity to HSP90 than a number of other geldanamycin derivatives probably due to additional hydrogen bonds engaging an amine group [28]. It has been demonstrated that 17-aminogeldanamycin is a bioavailable compound upon oral administration [29, 30], and can reduce tumor growth and vessel density in xenografts of gastrointestinal stromal tumors (GIST) [31]. In our previous study on 120 natural agents, we have found that 17-aminogeldanamycin is more cytotoxic than geldanamycin, and both compounds are more potent against melanoma than leukemic cells [32]. In addition, 17-aminogeldanamycin significantly reduces c-MYC transcript level while not affecting the frequency of cells positive for ATP-binding cassette, sub-family B, member 5 (ABCB5) [32], which is a drug efflux transporter that mediates chemoresistance and marks melanoma-initiating cells [33]. These preliminary results prompted us to evaluate the activity of 17-aminogeldanamycin in melanoma cells more extensively.
Materials and methods
Cell culture
Melanoma cell lines were derived from tumors obtained during surgical interventions. The study was approved by Ethical Commission of Medical University of Lodz, and informed consent was obtained from all patients. Tumor fragments were washed, minced with scissors and incubated in HBSS (Sigma-Aldrich, St Louis, MO, USA) supplemented with 3 mM CaCl2 and 1 mg/ml collagenase IV for few hours at 37 °C. 10 μg/ml DNase I was added and cells were filtered through a 70 μm pore size filter. Cells were cultured in a complete medium (RPMI-1640 supplemented with 10% FBS) for 1 day to remove dead and non-adherent cells. Then, they were transferred to serum-free stem cell medium (SCM) consisting of DMEM/F12 low osmolality medium (Gibco, Paisley, UK), B-27 supplement (Gibco), 10 μg/ml insulin, 1 ng/ml heparin, 10 ng/ml bFGF, 20 ng/ml EGF (BD Biosciences, San Jose, CA, USA), 100 IU/ml penicillin and 100 μg/ml streptomycin [34,35,36]. Cell lines were named DMBC12, DMBC21, DMBC28, DMBC29 and DMBC22 (Department of Molecular Biology of Cancer, DMBC). DMBC12, DMBC21, DMBC28 and DMBC29 cells were assigned to the BRAF subtype as they harbored either a homozygous (DMBC12) or heterozygous (DMBC21, DMBC28 and DMBC29) BRAFV600E variant, whereas DMBC22 cells harbored a homozygous Q61R substitution in NRAS [4]. For experiments, melanoma cells were seeded at final density, and drugs were added after 2.5 h.
Drugs
Vemurafenib and trametinib were purchased from Selleck Chemicals LLC (Houston, TX, USA), geldanamycin from Sigma-Aldrich and 17-aminogeldanamycin from BOC Sciences (Shirley, NY, USA). Chemical formulas of geldanamycin and 17-aminogeldanamycin were prepared using ISIS/Draw (version 2.3). Drug stocks were prepared in DMSO. For experiments, drugs were dissolved in the culture medium to final concentrations as following: 5 μM vemurafenib (PLX), 50 nM trametinib (TRA), 0,1 and 0,4 μM geldanamycin (GEL) and 17-aminogeldanamycin (AG).
Whole-exome sequencing
Whole-exome sequencing was performed as described previously [4]. Raw data are available at ArrayExpress and European Nucleotide Archive (ENA) under the numbers E-MTAB-6978 and ERP109743, respectively. Functional effects of amino acid substitutions were predicted in silico by the Polyphen-2 software (genetics.bwh.harvard.edu/pph2/index.shtml). The Polyphen-2-based predictions were classified as benign (scores 0.000–0.449), possibly damaging (scores 0.450–0.959) or probably damaging (scores 0.960–1.000).
A time-lapse fluorescent microscopy
Melanoma cells were grown in 96-well plates (8 × 103 cells/well) and treated with drugs at indicated concentrations and IncuCyte Caspase-3/7 Apoptosis Assay Reagent at 4 μM for 3 days. Activation of caspase-3/7 was monitored every 3 h by using a time-lapse fluorescence microscope system IncuCyte ZOOM (IncuCyte, Essen Bioscience). Data were analyzed using the IncuCyte Zoom original software. Percent of cells with active caspase-3/7 was calculated by dividing the percentages of confluence of apoptotic cells by the percentages of confluence of all cells at particular time points.
Acid phosphatase activity assay
Acid phosphatase activity was assessed to determine a number of viable melanoma cells. Melanoma cells were grown for 0, 24, 48 and 72 h, then the plates were centrifuged and medium was replaced with assay buffer as described previously [34]. The absorbance values were measured using a microplate reader Infinite M200Pro (Tecan, Salzburg, Austria).
Flow cytometry
Cells were incubated with drugs for 24 and 48 h, then collected, trypsinized and stained with Annexin V-FLUOS Staining Kit (Roche, Manheim, Germany) for 15 min. Flow cytometric data were acquired with FACSVerse (BD Biosciences), and analyzed using BD FACSuite.
Cell lysate preparation and Western blotting
Melanoma cells were lysed in RIPA buffer containing 50 mmol/l Tris–HCl pH 8.0, 150 mmol/l NaCl, 1% TritonX-100, 0.5% sodium deoxycholate, 0.1% SDS supplemented with freshly added protease and phosphatase inhibitors (Sigma-Aldrich). Cell lysates were diluted in 2 × Laemmli buffer and protein samples (15 μg) were loaded on standard 7% SDS–polyacrylamide gel. After electrophoresis, the proteins were transferred onto Immobilon-P PVDF membrane (Millipore, Billerica, MA, USA) followed by incubation in a blocking solution: 5% nonfat milk in PBS-Tween 0.05% or 5% phospho-BLOCKER (Cell Biolabs, San Diego, CA, USA) in PBS-Tween 0.05%. Primary antibodies detecting PARP, ATF6, IRE1α, GRP78 and p53 were from Santa Cruz Biotechnology (Santa Cruz, CA, USA), p-IRE1α from Abcam (Cambridge, UK), β-actin from Sigma-Aldrich, p-ERK1/2 (Thr202/Tyr204) and ERK1/2 from Cell Signaling Technology (Danvers, MA, USA). Secondary HRP-conjugated anti-mouse or anti-rabbit antibodies (Santa Cruz Biotechnology) and Pierce ECL Western Blotting Substrate (Pierce, Rockford, IL, USA) were used to visualize proteins on the X-ray film (Foton-Bis, Bydgoszcz, Poland) or by using ChemiDoc Imaging System (Biorad). The quantification of the Western blotting data was performed by using ImageJ software.
RNA isolation, cDNA synthesis and quantitative RT-PCR (qRT-PCR)
Total RNA isolation, cDNA synthesis and amplification procedures were extensively described elsewhere [32]. Primer sequences are shown in the Online Resource 1. To calculate the relative expression of target genes versus a reference gene RPS17, a mathematical model including an efficiency correction was used.
Statistical analysis
Graphs are presented as mean ± SD. To calculate statistical significance of differences, Statistica v.13 software was used. Normality of a sample distribution was assessed by the Shapiro–Wilk test. The Levene’s test was used to determine equality of variances. The unpaired t test was used to compare two samples with normal distribution and equal variances. In case of different sample size or when n = 2, Mann–Whitney U test was used. To compare three samples with normal distribution and equal variances, ANOVA was used. The differences were considered significant if p ≤ 0.05.
Results
17-Aminogeldanamycin (AG) is more effective than geldanamycin (GEL) in caspase-3/7 activation in melanoma cells
17-aminogeldanamycin (AG) is a geldanamycin (GEL) derivative in which a methoxy substituent attached to the C17 of the benzoquinone moiety is replaced by an amine group (Fig. 1a). First, we compared efficiency of GEL and AG in induction of apoptosis in two BRAFV600E patient-derived melanoma cell lines, DMBC21 and DMBC28. Using a time-lapse fluorescence microscopy, we showed that 0.4 μM AG caused in-cell caspase-3/7 activation already after 24 h. This was followed by further increase in the percentages of cells with active caspase-3/7 up to 30–35% and cell detachment (Fig. 1b; Online Resource 3), whereas GEL at this concentration was ineffective.
17-Aminogeldanamycin (AG) is more potent than geldanamycin (GEL) against BRAFV600E melanoma cell lines a Structural formulas of GEL and AG. b Percentages of cells with active caspase-3/7 were assessed by time-lapse imaging system IncuCyte ZOOM in DMBC21 and DMBC28 cell lines incubated with either AG or GEL at indicated concentrations for 72 h
AG at 0.4 μM reduces viability and inhibits ERK1/2 activity in BRAFV600E melanoma cells
We used additional BRAFV600E patient-derived cell lines to assess AG activity more extensively. AG reduced viable cell numbers after 72 h by 26, 62, 34 and 54% in DMBC12, DMBC21, DMBC28 and DMBC29 cell lines, respectively, as assessed by acid phosphatase activity (Fig. 2a). Double Annexin V/propidium iodide staining followed by flow cytometry revealed that 0.4 μM AG increased the frequency of Annexin V-positive cells in a time-dependent manner (Fig. 2b), whereas AG at 0.1 μM did not consistently induce apoptosis (Fig. 2b) indicating that AG at this concentration could rather exhibit cytostatic effect in melanoma cell lines (Fig. 2a). Apoptosis induced by 0.4 μM AG was also confirmed by detection of a caspase-mediated cleavage product of PARP already after 24 h (Fig. 2c), consistently with caspase-3/7 activation shown in DMBC21 and DMBC28 cell lines (Fig. 1b). In addition, AG-dependent diminution of the MAPK signaling pathway activity was observed (Fig. 2d). While this effect was almost undetectable after 4 h of exposure to drug, longer incubation markedly reduced ERK1/2 activity in both ERK1/2high (DMBC12) and ERK1/2low (DMBC21, DMBC28 and DMBC29) melanoma cell lines (Fig. 2d).
17-Aminogeldanamycin (AG) reduces viable cell number and inhibits activity of the MAPK signaling pathway a Melanoma cells were incubated with AG at 0.1 μM or 0.4 μM. Changes in viable cell number were assessed by acid phosphatase activity assay over the course of 72 h. Representative results are shown. *p ≤ 0.05 vs. control b–c Induction of apoptosis by AG is shown as percentages of Annexin V-positive cells after 24 and 48 h (*p ≤ 0.05 vs. control), and the level of cleaved PARP (cPARP) after 4 and 24 h. An equal loading was confirmed by β-actin. d Level of phosphorylated ERK1/2 (p-ERK1/2) was determined by Western Blotting after 4 and 24 h of cell incubation with 0.4 μM AG. β-actin was used as a loading control. p-ERK1/2 level was normalized to β-actin level, and shown below the blots
AG transiently increases mRNA levels of HSP70 and GRP78
Upregulation of HSP70 expression compensates for attenuation of HSP90 activity by N-terminal inhibitors including geldanamycin and geldanamycin analogues [37]. AG at 0.4 μM significantly increased the transcript level of HSP70 after 6 h, which was a transient effect as HSP70 mRNA level decreased (p < 0.05) after additional 16 h of incubation with a drug (Fig. 3a). In addition, the transcript level of GRP78, a marker of ER stress induction, was significantly increased in all melanoma cell lines after 6 h, and reduced (p < 0.05) to the level of control after 22 h (Fig. 3b). Changes in GRP78 expression were not associated with alterations in the level of corresponding protein assessed at two time intervals (Fig. 3c).
17-Aminogeldanamycin (AG) at 0.4 μM transiently increases mRNA levels of HSP70 and GRP78, and inhibits IRE1α-dependent pathway of UPR a, b HSP70 and GRP78 transcript levels were assessed by qRT-PCR after 6 and 22 h, and expressed relatively to the control. *p ≤ 0.05 c GRP78 protein level was determined by Western blotting after 4 and 24 h. An equal loading was confirmed by β-actin. Quantification of the protein level is shown below the blots. d Levels of active (phosphorylated IRE1α; p-IRE1α) and total IRE1α were determined after 4 and 24 h. β-actin was used as a loading control. Quantifications of p-IRE1α and IRE1α levels are shown below the blots. e XBP1 s transcript level after 22 h is shown relatively to the control. *p ≤ 0.05 f Activity of ATF6 was determined as the level of ATF6 cleavage product (p50) after 4 and 24 h. An equal loading was confirmed by β-actin. Quantification of the protein level is shown below the blots. g The transcript levels of CHOP and BIM were assessed by qRT-PCR after 22 h, and expressed relatively to the control. *p ≤ 0.05
AG selectively diminishes activity of IRE1α-dependent pathway of UPR
AG-mediated upregulation of GRP78 expression suggested induction of ER stress. AG slightly increased IRE1α activity after 4 h (Fig. 3d), but levels of both total and phosphorylated IRE1α (p-IRE1α) were markedly reduced after additional 20 h of incubation with a drug (Fig. 3d). As a consequence, level of spliced XBP1 (XBP1s) mRNA was significantly diminished in all melanoma cell lines (Fig. 3e). Importantly, GEL at the same concentration did not affect the transcript level of XBP1s in DMBC21 and DMBC29 cells (Online Resource 4) that exhibited the largest decrease in XBP1s mRNA level in response to AG (Fig. 3e). Apart from its evident effect on the IRE1α-XBP1s pathway activity, 0.4 μM AG did not markedly affect level of a nuclear form of ATF6 (p50) (Fig. 3f), and did not significantly induce CHOP and BIM expression (Fig. 3g) encoding for executioners of PERK-dependent apoptosis. For that reason, we assessed p53/BIK-dependent route as an alternative pathway of apoptosis induced in response to prolonged ER stress [38]. AG at 0.4 μM did not substantially alter the protein level of p53, except for an increase in DMBC12 cells already after 4 h (Online Resource 5a). This was associated with an insignificant effect of AG on BIK transcript level in all melanoma cell lines (Online Resource 5b). To elucidate lack of apparent mechanism of ER stress-triggered apoptosis, we used whole-exome sequencing data [4] to determine the mutation status of genes encoding components of the UPR cascades. We found that PERK-dependent pathway might be affected by an inframe deletion in EIF2AK3 leading to a PERKL21del variant that was harbored in all melanoma cell lines (Online Resource 2), although genes encoding downstream components of the PERK signaling, ATF4 and eIF2α, either harbored alteration leading to a variant predicted as benign or were unaltered (Online Resource 2).
AG cooperates with inhibitors of BRAFV600E and MEK1/2 in induction of apoptosis
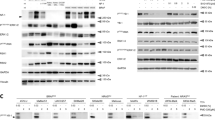
HSP90 is a chaperone protein for several oncoproteins that contribute to melanoma cell response to inhibitors of the MAPK signaling pathway, vemurafenib (PLX; an inhibitor of BRAFV600E) and trametinib (TRA; an inhibitor of MEK1/2). Therefore, we investigated whether a combination of AG and PLX or TRA could cooperatively induce apoptosis. In the following experiments, we used also 0.1 μM AG. AG already at this low concentration significantly increased HSP70 transcript level after 6 h, which was followed by a reduction (p < 0.05) after additional 16 h of incubation in all melanoma cell lines (Online Resource 6a). It also significantly augmented GRP78 mRNA level after 6 h in DMBC29 cells (Online Resource 6b) while not triggering apoptosis when used alone (Figs. 1b and 2b). PLX at 5 μM and TRA at 50 nM did not induce significant changes in the transcript levels of HSP70 (Fig. 4a) and GRP78 (Fig. 4b) in all BRAFV600E cell lines. As expected, PLX and TRA inhibited ERK1/2 activity already after 4 h (Fig. 4c). Similar effect was exerted by 0.4 μM AG after 24 h (Figs. 2d, 4c), whereas AG at 0.1 μM poorly affected level of phospho-ERK1/2 (Fig. 4c). In drug combinations, the effect of PLX and TRA was dominant, and AG did not interfere with PLX- and TRA-mediated attenuation of ERK1/2 activity (Fig. 4c). Real-time monitoring of cells incubated with a combination of 50 nM TRA and 0.4 μM AG revealed that activation of caspase-3/7 occurred earlier and in a larger number of cells compared with cells incubated with either drug used alone (Fig. 4d). For example, the percentage of cells with active caspase-3/7 was around or < 10% upon incubation with either 50 nM TRA or 0.4 μM AG, but reached almost 20% and 30% when a drug combination was used in DMBC21 and DMBC28 cells, respectively (Fig. 4d). This cooperatively induced apoptosis was observed in DMBC21, DMBC28 and DMBC29 cell lines, but not in DMBC12 cells (Fig. 4d), which was consistently reflected in the level of cleaved PARP after 24 h (Fig. 4e; Online Resource 7a). Combined effect of 5 μM PLX and 0.4 μM AG was also observed as induction of PARP cleavage, however, it was less pronounced than effect of the TRA + AG combination in DMBC21 and DMBC29 cell lines, and not observed in DMBC12 cells (Fig. 4e; Online Resource 7a). We also determined the level of IRE1α and XBP1s in melanoma cells exposed to PLX and TRA, and their combinations with AG. We found that PLX and TRA inhibited IRE1α activity in DMBC12 cells after 24 h, but it was associated with insignificant alteration in the transcript level of XBP1s (Online Resource 7b). PLX and TRA also slightly diminished levels of phospho-IRE1α and XBP1s in DMBC21 cells. In combination with AG, PLX/TRA did not cooperatively reduce the XBP1s mRNA level in any melanoma cell line (Online Resource 7b).
17-Aminogeldanamycin (AG) inhibits ERK1/2 activity and cooperates with trametinib (TRA) and vemurafenib (PLX) in induction of apoptosis in BRAFV600E melanoma cell lines a, b HSP70 and GRP78 transcript levels were assessed by qRT-PCR after 6 and 22 h of cell incubation with either 5 μM PLX or 50 nM TRA, and expressed relatively to the control. ns not significant. c–e Melanoma cell lines were incubated with AG, PLX and TRA used either alone or in combinations at indicated concentrations. c Level of phosphorylated ERK1/2 (p-ERK1/2) was assessed by Western blotting after 4 and 24 h. An equal loading was confirmed by β-actin. d Percentages of cells with active caspase-3/7 were assessed by time-lapse imaging system IncuCyte ZOOM. e Level of cleaved PARP (cPARP) was determined after 4 and 24 h. An equal loading was confirmed by β-actin. Quantification of cPARP level after 24 h of cell incubation with drugs is shown in the Online Resource 7a
AG is also effective against NRASQ61R melanoma cells
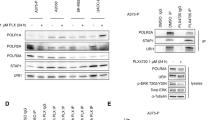
We also employed DMBC22 cell line that was previously assigned to the NRAS subtype of melanoma as it harbored a homozygous NRASQ61R variant and a wild-type BRAF [4]. AG at 0.4 μM markedly reduced viability in DMBC22 cell line by 52% after 72 h as evidenced by changes in acid phosphatase activity (Fig. 5a), and significantly increased transcript levels of HSP70 and GRP78 after 6 h, which was followed by a decrease (p < 0.05) to the level of control after 22 h (Fig. 5b). In addition, 0.4 μM AG reduced the levels of phospho-IRE1α and XBP1s transcript (Fig. 5c). To elucidate whether drug cooperation reported in most of BRAFV600E cell lines was also executed in NRASQ61R melanoma cells, DMBC22 cells were incubated with 50 nM TRA and 0.4 μM AG, used either alone or in combination. While TRA inhibited ERK1/2 activity already after 4 h, AG reduced level of phospho-ERK1/2 level after 24 h of incubation (Fig. 5d). Combination of TRA and AG increased the frequency of Annexin V-positive cells in a time-dependent manner (Fig. 5e) and augmented the level of cleaved PARP after 24 h (Fig. 5f) to larger extents than those observed for either drug used alone. Importantly, the occurrence of AG-induced apoptosis in DMBC22 cell line was slightly delayed compared to BRAFV600E cells as consistently reported at the cellular and molecular levels (Fig. 5e, f vs. Fig. 2b, c). Notably, this was clearly visible in real-time measurement of the percentages of cells with active caspase-3/7 (Fig. 5g vs. Fig. 4d). Moreover, AG and TRA used in combination induced caspase-3/7 much earlier than either drug alone (Fig. 5g). For example, the percentage of cells with active caspase-3/7 was ~ 5% when cells were incubated with either 50 nM TRA or 0.4 μM AG, but reached ~ 25% when DMBC22 cells were exposed to a combination of drugs (Fig. 5g). In addition, p53-BIK axis was unlikely involved in the induction of apoptosis (Online Resource 5a, b), whereas PERK pathway-dependent upregulation of CHOP and BIM expression was not detected (data not shown) possibly due to a homozygous PERKL21del variant harbored in DMBC22 cells similarly to BRAFV600E cell lines (Online Resource 2).
Cellular and molecular effects of 17-aminogeldanamycin (AG) activity used either alone or in combination with trametinib (TRA) in NRASQ61R melanoma cells a Changes in a viable cell number were assessed by acid phosphatase assay. Representative results are shown. *p ≤ 0.05 versus control b HSP70 and GRP78 transcript levels were assessed by qRT-PCR in cells incubated with 0.4 μM AG for 6 and 22 h, and expressed relatively to the control. *p ≤ 0.05 c Level of phospho-IRE1α (p-IRE1α) was determined after 4 and 24 h. β-actin was used as a loading control. Quantifications of p-IRE1α and IRE1α levels are shown below the blots. XBP1s transcript level was assessed by qRT-PCR after 22 h, and expressed relatively to control. d–g DMBC22 cells were incubated with 0.4 μM AG and 50 nM TRA used either alone or in combination. d Level of phosphorylated ERK1/2 (p-ERK1/2) was assessed by Western blotting, and β-actin was used as a loading control. e Percentages of Annexin V-positive cells after 24 and 48 h were determined by flow cytometry. Representative contour plots are shown. *p ≤ 0.05 drug combination vs. either drug used alone f Cleaved PARP (cPARP) was immunoblotted after 24 h. An equal loading was confirmed by β-actin. Quantification of cPARP level is shown below the blots. g Percentages of cells with active caspase-3/7 were assessed by time-lapse microscopy
Discussion
In the present study, we have demonstrated that 17-aminogeldanamycin perturbates ER homeostasis, predominantly by interfering with activity of IRE1α-dependent pathway, and induces apoptosis in melanoma cells of the BRAFV600E and NRASQ61R subtypes. 17-aminogeldanamycin takes several advantages over geldanamycin and other geldanamycin derivatives as supported by previously published and present results showing that 17-aminogeldanamycin (i) is more potent than geldanamycin against melanoma cells; (ii) exerts anti-melanoma activity at lower concentrations compared with IC50 values for other geldanamycin analogues assessed in different human cancer cell lines [26], and (iii) attenuates self-triggered increase in HSP70 transcript level.
17-Aminogeldanamycin alleviates self-triggered upregulation of HSP70 expression
It has been demonstrated that N-terminal inhibitors of HSP90 activate heat shock factor-1 (HSF-1) to upregulate expression of other chaperones including HSP70 and HSP27 [37, 39, 40]. This cell response contributes to development of resistance to HSP90 inhibitors [41, 42] as compensatory chaperones can remain at increased levels in cells incubated with geldanamycin, its analogues [43,44,45] and XL888, a geldanamycin-unrelated inhibitor of HSP90 [46]. Here, we have shown that 17-aminogeldanamycin upregulates HSP70 expression, which returns to baseline mRNA level in both BRAFV600E and NRASQ61R melanoma cell lines. It is consistent with other results showing insignificant alterations in the protein level of HSP70 in GIST cell lysates after 17-aminogeldanamycin administration [31]. This suggests that attenuation of self-induced upregulation of HSP70 expression is rather drug-specific than cell type-dependent, and might be crucial for a high activity of 17-aminogeldanamycin in melanoma cells.
17-Aminogeldanamycin selectively inhibits cytoprotective IRE1α-XBP1s signaling pathway of UPR
Upregulation of HSP70 expression following inhibition of HSP90 compensates for HSP90 function and deters from activation of prolonged ER stress and UPR, and induction of apoptosis emerging from an increased burden of misfolded proteins [47]. Disturbances in the execution of UPR support adaptation of cancer cells to proteotoxic stress [48]. In melanoma, UPR and other stress-attenuating signaling pathways are hyperactivated during tumor progression and cell response to therapy [49, 50]. They engage different anti-apoptotic proteins [51] and microphthalmia-associated transcription factor (MITF) [52]. Increasing evidence suggests that targeting the ER stress and UPR programs can be a promising anti-melanoma strategy [53, 54], also against resistant cells due to the contribution of GRP78/BiP to ERK1/2 reactivation [55]. In the present study, we have found that 17-aminogeldanamycin induces ER stress as evidenced by a transient increase in GRP78 transcript level and a slight activation of IRE1α already after 4 h. IRE1α autophosphorylation followed by its dimerization leads to the excision of a 26-nt intron from XBP1 transcript and a generation of XBP1s mRNA translated into XBP1, a transcription factor that regulates expression of ER stress-attenuating genes [56, 57]. XBP1 has been implicated in the regulation of melanoma cell proliferation by activating inteleukin-6/STAT3 pathway [58] in addition to its critical role in melanoma cell survival during ER stress [59]. As both IRE1α and ATF6 can upregulate expression of GRP78 in melanoma [60], the contribution of IRE1α to the increase in GRP78 mRNA level is more likely because ATF6 activity remained unaltered following 17-aminogeldanamycin treatment. More importantly, while IRE1α activation could be an acute cytoprotective response of melanoma cells to 17-aminogeldanamycin, prolonged incubation with this compound led to a substantial diminution of both activity and protein level of IRE1α, probably due to a chaperoning role of HSP90 on IRE1α [61]. This effect can be specific for particular HSP90 inhibitors because tanespimycin inhibited XBP1s generation without abrogating IRE1α protein level even when used in higher concentration (1 µM) [62] than 0.4 µM 17-aminogeldanamycin used in our study. Therefore, 17-aminogeldanamycin is capable to reduce IRE1α activity and also, by leading to IRE1α degradation, to prevent from undesired activation of cytoprotective IRE1α signaling, which might be valuable considering 17-aminogeldanamycin as a part of combined treatment.
17-Aminogeldanamycin induces apoptosis in melanoma cells harboring PERKL21del and NQO1P187S variants
ER stress-induced apoptosis has been commonly attributed to PERK signaling-dependent upregulation of CHOP and BIM expression [14], although CHOP-deficient cells also undergo apoptosis in response to ER stress inducers [63]. Alternatively, p53-dependent suppression of GRP78 mRNA translation and induction of BIK expression can induce apoptosis [38]. In the present study, we have demonstrated that apoptosis induced by 17-aminogeldanamycin unlikely depends on any of these pathways. PERK-dependent signaling might be affected due to an inframe deletion in EIF2AK3 leading to a PERKL21del variant that was found in all melanoma cell lines. A PERKL21del variant has been already reported in one tumor sample excised from a patient before treatment with vemurafenib [64]. It needs to be elucidated how deletion of Leu21 affects PERK structure and activity. On the other hand, an increase in p53 level was found exclusively in one cell line and this was not associated with either decrease in GRP78 protein level (Fig. 3c) or upregulation of BIK expression (Online Resource 5), despite transcriptionally active p53 in all melanoma cell lines [4]. Thus, while 17-aminogeldanamycin apparently diminishes activity of cytoprotective IRE1α-XBP1s axis and induces apoptosis in melanoma cells, specific mediators of cell death need to be determined.
Cell sensitivity to geldanamycin derivatives correlates with expression of NAD(P)H:quinone oxidoreductase 1 (NQO1) [65], and NQO1-mediated quinone-to-hydroquinone conversion of geldanamycin and its analogues enhances activity of these compounds because of increased hydrogen bonding of hydroquinone derivatives [66]. It has also been shown that a P187S variant of NQO1 has a reduced activity compared with wild-type NQO1 [67], but genetic alterations affecting a His80 residue in NQO1 can compensate for P187S substitution [68]. In our study, a heterozygous NQO1P187S variant (rs1800566) was harbored exclusively in DMBC22 cells. As DMBC22 cells lack additional alterations in NQO1, NQO1P187S variant might contribute to a delayed pro-apoptotic response of DMBC22 cells to 17-aminogeldanamycin compared with the response of cell lines harboring wild-type NQO1. However, it has been demonstrated that the affinity of 17-aminogeldanamycin to purified HSP90 is not considerably enhanced upon reduction to the hydroquinone [29], and cell response to 17-aminogeldanamycin is not associated with NQO1 protein level [31]. It suggests that activity of 17-aminogeldanamycin is at least partially independent of NQO1 that can be advantageous because a loss of NQO1 expression and acquisition of an inactive NQO1P187S variant have been demonstrated as causative factors in development of resistance to tanespimycin in melanoma and glioblastoma cells [69].
17-Aminogeldanamycin and inhibitors of the RAS/RAF/MEK signaling cascade cooperatively induce apoptosis
A tolerable side effect profile has been recently shown for a combination of HSP90 and BRAFV600E inhibitors [20]. Clinical trials NCT01657591 and NCT02721459 on co-treatment of melanoma patients with XL888 and inhibitors of the MAPK signaling pathway are ongoing (clinicaltrials.gov). The rationale for combining these drugs is that several proteins, which are linked to melanoma cell response and resistance to BRAFV600/MEK-targeting agents, are HSP90 clients including ARAF, CRAF, BRAFV600E, CDK4, COT, IGF1R, PDGFR-β and AKT [16, 70, 71]. In addition, it has been demonstrated that oncogenic MAPK signaling increases intracellular protein load and maintains cytoprotective autophagy at elevated level [59, 72], and sustains IRE1α and ATF6 at activated states to adapt melanoma cells to a chronic ER stress [60]. Indeed, attenuation of the MAPK pathway activity sensitizes melanoma cells to ER stress-inducing drugs [73]. Moreover, it even directly causes the release of Ca2+ from endoplasmic reticulum [74] and enhances interaction between mutant BRAF and GRP78 to induce PERK-dependent apoptosis [75]. In our study, vemurafenib and trametinib did not upregulate GRP78 expression, but affected IRE1α-XBP1s signaling in certain melanoma cell lines. Therefore, we propose a putative model of cooperation between 17-aminogeldanamycin and trametinib or vemurafenib (Fig. 6a). According to this model, 17-aminogeldanamycin exerts dual time-dependent activity. By inhibiting HSP90, 17-aminogeldanamycin triggers accumulation of unfolded proteins thereby rapidly inducing ER stress as evidenced by an increase in GRP78 and HSP70 transcript levels and a slight activation of IRE1α. It is followed, however, by a decay of IRE1α protein, which results in the attenuation of cytoprotective IRE1α-XBP1s axis, in addition to a diminution of the transcript levels of HSP70 and GRP78 as well as activity of ERK1/2. ERK1/2 maintains basal ER homeostasis in melanoma cells. In comparison to 17-aminogeldanamycin, trametinib and vemurafenib rapidly attenuate ERK1/2 activity, which might be crucial for potentiating effects triggered by 17-aminogeldanamycin. Consequently, apoptosis is induced earlier and in a larger number of cells compared with either drug used alone, but not in all cell lines. We have found that the transcript levels of XBP1s, GRP78, HSP70 and CHOP were the lowest in drug-naïve DMBC12 cells compared with other BRAFV600E and NRASQ61R melanoma cell lines (Fig. 6b). Already low adaptation to ER disturbances might determine lack of cooperation between 17-aminogeldanamycin and targeted therapeutics in DMBC12 cells. Therefore, our study suggests that cooperatively induced apoptosis might result from a concurrent inhibition of both, the MAPK signaling and cytoprotective IRE1α-XBP1s axis, and cell-intrinsic ER homeostasis can narrow the extent of drug cooperation.
Cooperation of 17-aminogeldanamycin (AG) and trametinib (TRA) or vemurafenib (PLX) a A schematic summary of molecular effects of TRA, and PLX activity, and 17-aminogeldanamycin-mediated HSP90 inhibition in melanoma cells. Alterations accompanying acute (4–6 h) and prolonged (22–24 h) response are shown. See discussion for additional comments. ERAD, endoplasmic reticulum-associated protein degradation b Transcript levels of XBP1 s, GRP78, HSP70 and CHOP were determined by qRT-PCR in drug-naïve BRAFV600E and NRASQ61R melanoma cell lines, and expressed relatively to the median value in all five cell lines. *p ≤ 0.05 versus median value
Conclusion
17-Aminogeldanamycin is a more potent HSP90 inhibitor than geldanamycin against melanoma cells, and exerts previously unidentified activities including diminution of self-triggered upregulation of HSP70 expression and selective inhibition of cytoprotective IRE1α-XBP1s axis of unfolded protein response. In addition, 17-aminogeldanamycin cooperates with inhibitors of the MAPK signaling pathway in apoptosis induction that might be exploited in BRAFV600E and NRASQ61R melanomas.
Abbreviations
- AG:
-
17-Aminogeldanamycin
- ATF6:
-
Activating transcription factor 6
- BIK:
-
BCL2-interacting killer
- BIM:
-
BCL-2-interacting mediator of cell death
- BRAF:
-
B-Raf proto-oncogene serine/threonine kinase
- CHOP:
-
C/EBP homologous protein
- DMBC:
-
Department of Molecular Biology of Cancer
- ER:
-
Endoplasmic reticulum
- ERAD:
-
Endoplasmic reticulum-associated protein degradation
- ERK1/2:
-
Extracellular signal-regulated kinase 1/2
- GEL:
-
Geldanamycin
- GRP78/BiP:
-
Glucose-regulated protein 78/binding immunoglobulin protein
- HSP70:
-
Heat shock protein 70
- HSP90:
-
Heat shock protein 90
- MEK1/2:
-
Mitogen-activated protein kinase 1/2
- IRE1α:
-
Inositol-requiring enzyme 1 alpha
- NRAS:
-
Neuroblastoma RAS viral oncogene homolog
- NQO1:
-
NAD(P)H:quinone oxidoreductase 1
- PARP1:
-
Poly(ADP-ribose) polymerase 1
- PERK:
-
Protein kinase R-like endoplasmic reticulum kinase
- PLX:
-
Vemurafenib an inhibitor of BRAFV600E
- TRA:
-
Trametinib an inhibitor of MEK1/2
- UPR:
-
Unfolded protein response
- XBP1 (XBP1s):
-
X-box binding protein 1 (spliced XBP-1)
References
Kaufman HL, Margolin K, Sullivan R (2018) Management of metastatic melanoma in 2018. JAMA Oncol 4(6):857–858. https://doi.org/10.1001/jamaoncol.2018.0170
Cancer Genome Atlas Network (2015) Genomic classification of cutaneous melanoma. Cell 161(7):1681–1696. https://doi.org/10.1016/j.cell.2015.05.044
Garman B, Anastopoulos IN, Krepler C, Brafford P, Sproesser K, Jiang Y, Wubbenhorst B, Amaravadi R, Bennett J, Begiri M, Elder D, Flaherty KT, Frederick DT, Gangadhar TC, Guarino M, Hoon D, Karakousis G, Liu Q, Mitra N, Petrelli NJ, Schuchter L, Shannan B, Shields CL, Wargo J, Wenz B, Wilson MA, Xiao M, Xu W, Xu X, Yin X, Zhang NR, Davies MA, Herlyn M, Nathanson KL (2017) Genetic and genomic characterization of 462 melanoma patient-derived xenografts, tumor biopsies, and cell lines. Cell Rep 21(7):1936–1952. https://doi.org/10.1016/j.celrep.2017.10.052
Hartman ML, Sztiller-Sikorska M, Czyz M (2019) Whole-exome sequencing reveals novel genetic variants associated with diverse phenotypes of melanoma cells. Mol Carcinog 58(4):588–602. https://doi.org/10.1002/mc.22953
Senses KM, Ghasemi M, Akbar MW, Isbilen M, Fallacara AL, Frankenburg S, Schenone S, Lotem M, Botta M, Gure AO (2017) Phenotype-based variation as a biomarker of sensitivity to molecularly targeted therapy in melanoma. Medchemcomm 8(1):88–95. https://doi.org/10.1039/C6MD00466K
Schopf FH, Biebl MM, Buchner J (2017) The HSP90 chaperone machinery. Nat Rev Mol Cell Biol 18(6):345–360. https://doi.org/10.1038/nrm.2017.20
Morán Luengo T, Mayer MP, Rüdiger SGD (2019) The Hsp70-Hsp90 chaperone cascade in protein folding. Trends Cell Biol 29(2):164–177. https://doi.org/10.1016/j.tcb.2018.10.004
Zhu H, Fang X, Zhang D, Wu W, Shao M, Wang L, Gu J (2016) Membrane-bound heat shock proteins facilitate the uptake of dying cells and cross-presentation of cellular antigen. Apoptosis 21(1):96–109. https://doi.org/10.1007/s10495-015-1187-0
Shipp C, Weide B, Derhovanessian E, Pawelec G (2013) Hsps are up-regulated in melanoma tissue and correlate with patient clinical parameters. Cell Stress Chaperones 18(2):145–154. https://doi.org/10.1007/s12192-012-0363-1
Strickler AG, Vasquez JG, Yates N, Ho J (2014) Potential diagnostic significance of HSP90, ACS/TMS1, and L-plastin in the identification of melanoma. Melanoma Res 24(6):535–544. https://doi.org/10.1097/CMR.0000000000000115
Tas F, Bilgin E, Erturk K, Duranyildiz D (2017) Clinical significance of circulating serum cellular Heat Shock Protein 90 (HSP90) level in patients with cutaneous malignant melanoma. Asian Pac J Cancer Prev 18(3):599–601. https://doi.org/10.22034/APJCP.2017.18.3.599
Peinado H, Alečković M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M, Williams C, García-Santos G, Ghajar C, Nitadori-Hoshino A, Hoffman C, Badal K, Garcia BA, Callahan MK, Yuan J, Martins VR, Skog J, Kaplan RN, Brady MS, Wolchok JD, Chapman PB, Kang Y, Bromberg J, Lyden D (2012) Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 18(6):883–891. https://doi.org/10.1038/nm.2753
Hoter A, El-Sabban ME, Naim HY (2018) The HSP90 family: structure, regulation, function, and implications in health and disease. Int J Mol Sci 19(9):E2560. https://doi.org/10.3390/ijms19092560
Almanza A, Carlesso A, Chintha C, Creedican S, Doultsinos D, Leuzzi B, Luís A, McCarthy N, Montibeller L, More S, Papaioannou A, Püschel F, Sassano ML, Skoko J, Agostinis P, deBelleroche J, Eriksson LA, Fulda S, Gorman AM, Healy S, Kozlov A, Muñoz-Pinedo C, Rehm M, Chevet E, Samali A (2018) Endoplasmic reticulum stress signalling—from basic mechanisms to clinical applications. FEBS J 286(2):241–278. https://doi.org/10.1111/febs.14608
Barbagallo I, Parenti R, Zappalà A, Vanella L, Tibullo D, Pepe F, Onni T, Li Volti G (2015) Combined inhibition of Hsp90 and heme oxygenase-1 induces apoptosis and endoplasmic reticulum stress in melanoma. Acta Histochem 117(8):705–711. https://doi.org/10.1016/j.acthis.2015.09.005
Paraiso KH, Haarberg HE, Wood E, Rebecca VW, Chen YA, Xiang Y, Ribas A, Lo RS, Weber JS, Sondak VK, John JK, Sarnaik AA, Koomen JM, Smalley KS (2012) The HSP90 inhibitor XL888 overcomes BRAF inhibitor resistance mediated through diverse mechanisms. Clin Cancer Res 18(9):2502–2514. https://doi.org/10.1158/1078-0432.CCR-11-2612
Smyth T, Paraiso KHT, Hearn K, Rodriguez-Lopez AM, Munck JM, Haarberg HE, Sondak VK, Thompson NT, Azab M, Lyons JF, Smalley KSM, Wallis NG (2014) Inhibition of HSP90 by AT13387 delays the emergence of resistance to BRAF inhibitors and overcomes resistance to dual BRAF and MEK inhibition in melanoma models. Mol Cancer Ther 13(12):2793–2804. https://doi.org/10.1158/1535-7163.MCT-14-0452
Zhang G, Frederick DT, Wu L, Wei Z, Krepler C, Srinivasan S, Chae YC, Xu X, Choi H, Dimwamwa E, Ope O, Shannan B, Basu D, Zhang D, Guha M, Xiao M, Randell S, Sproesser K, Xu W, Liu J, Karakousis GC, Schuchter LM, Gangadhar TC, Amaravadi RK, Gu M, Xu C, Ghosh A, Xu W, Tian T, Zhang J, Zha S, Liu Q, Brafford P, Weeraratna A, Davies MA, Wargo JA, Avadhani NG, Lu Y, Mills GB, Altieri DC, Flaherty KT, Herlyn M (2016) Targeting mitochondrial biogenesis to overcome drug resistance to MAPK inhibitors. J Clin Invest 126(5):1834–1856. https://doi.org/10.1172/JCI82661
Janssen N, Speigl L, Pawelec G, Niessner H, Shipp C (2018) Inhibiting HSP90 prevents the induction of myeloid-derived suppressor cells by melanoma cells. Cell Immunol 327:68–76. https://doi.org/10.1016/j.cellimm.2018.02.012
Eroglu Z, Chen YA, Gibney GT, Weber JS, Kudchadkar RR, Khushalani NI, Markowitz J, Brohl AS, Tetteh LF, Ramadan H, Arnone G, Li J, Zhao X, Sharma R, Darville LNF, Fang B, Smalley I, Messina JL, Koomen JM, Sondak VK, Smalley KSM (2018) Combined BRAF and HSP90 inhibition in patients with unresectable BRAF V600E-mutant melanoma. Clin Cancer Res 24(22):5516–5524. https://doi.org/10.1158/1078-0432.CCR-18-0565
Azimi A, Caramuta S, Seashore-Ludlow B, Boström J, Robinson JL, Edfors F, Tuominen R, Kemper K, Krijgsman O, Peeper DS, Nielsen J, Hansson J, Brage ES, Altun M, Uhlen M, Maddalo G (2018) Targeting CDK2 overcomes melanoma resistance against BRAF and Hsp90 inhibitors. Mol Syst Biol 14(3):e7858. https://doi.org/10.15252/msb.20177858
Desai BM, Villanueva J, Nguyen TT, Lioni M, Xiao M, Kong J, Krepler C, Vultur A, Flaherty KT, Nathanson KL, Smalley KS, Herlyn M (2013) The anti-melanoma activity of dinaciclib, a cyclin-dependent kinase inhibitor, is dependent on p53 signaling. PLoS ONE 8(3):e59588. https://doi.org/10.1371/journal.pone.0059588
Fukuyo Y, Inoue M, Nakajima T, Higashikubo R, Horikoshi NT, Hunt C, Usheva A, Freeman ML, Horikoshi N (2008) Oxidative stress plays a critical role in inactivating mutant BRAF by geldanamycin derivatives. Cancer Res 68(15):6324–6330. https://doi.org/10.1158/0008-5472.CAN-07-6602
Fukuyo Y, Hunt CR, Horikoshi N (2010) Geldanamycin and its anti-cancer activities. Cancer Lett 290(1):24–35. https://doi.org/10.1016/j.canlet.2009.07.010
Samuni Y, Ishii H, Hyodo F, Samuni U, Krishna MC, Goldstein S, Mitchell JB (2010) Reactive oxygen species mediate hepatotoxicity induced by the Hsp90 inhibitor geldanamycin and its analogs. Free Radic Biol Med 48(11):1559–1563. https://doi.org/10.1016/j.freeradbiomed.2010.03.001
Li YP, Chen JJ, Shen JJ, Cui J, Wu LZ, Wang Z, Li ZR (2015) Synthesis and biological evaluation of geldanamycin analogs against human cancer cells. Cancer Chemother Pharmacol 75(4):773–782. https://doi.org/10.1007/s00280-015-2696-9
Goetz MP, Toft D, Reid J, Ames M, Stensgard B, Safgren S, Adjei AA, Sloan J, Atherton P, Vasile V, Salazaar S, Adjei A, Croghan G, Erlichman CJ (2005) Phase I trial of 17-allylamino-17-demethoxygeldanamycin in patients with advanced cancer. Clin Oncol 23(6):1078–1087. https://doi.org/10.1200/JCO.2005.09.119
Tian ZQ, Liu Y, Zhang D, Wang Z, Dong SD, Carreras CW, Zhou Y, Rastelli G, Santi DV, Myles DC (2004) Synthesis and biological activities of novel 17-aminogeldanamycin derivatives. Bioorg Med Chem 12(20):5317–5329. https://doi.org/10.1016/j.bmc.2004.07.053
Lee J, Grenier L, Holson E, Slocum K, Coco J, Ge J, Normant E, Hoyt J, Lim A, Cushing J, Sydor J, Wright J (2008) IPI-493, a potent, orally bioavailable HSP90 inhibitor of the ansamycin class. Abstract, EORTC-NCI-AACR Symposium on “Molecular Targets and Cancer Therapeutics
Senzer NN, Mendelson D, Weekes C, Rosen L, LoRusso P, Just R, Smith D, Ritchie E, DeLucia D, Quigley T, Dunbar J, Schmalbach T, Cortes J (2011) Safety, tolerability, and pharmacokinetics of IPI-493, an oral Hsp90 inhibitor, in patients with advanced cancers. Mol Cancer Ther 11(11 Suppl):Abstract nr B101
Floris G, Sciot R, Wozniak A, Van Looy T, Wellens J, Faa G, Normant E, Debiec-Rychter M, Schöffski P (2011) The Novel HSP90 inhibitor, IPI-493, is highly effective in human gastrostrointestinal stromal tumor xenografts carrying heterogeneous KIT mutations. Clin Cancer Res 17(17):5604–5614. https://doi.org/10.1158/1078-0432.CCR-11-0562
Sztiller-Sikorska M, Koprowska K, Majchrzak K, Hartman M, Czyz M (2014) Natural compounds’ activity against cancer stem-like or fast-cycling melanoma cells. PLoS ONE 9(3):e90783. https://doi.org/10.1371/journal.pone.0090783
Wilson BJ, Saab KR, Ma J, Schatton T, Putz P, Zhan Q, Murohy GF, Gasser M, Waaga-Gasser AM, Frank NY, Frank MH (2014) ABCB5 maintains melanoma-initiating cells through a proinflammatory cytokine signaling circuit. Cancer Res 74(15):4196–4207. https://doi.org/10.1158/0008-5472.CAN-14-0582
Sztiller-Sikorska M, Koprowska K, Jakubowska J, Zalesna I, Stasiak M, Duechler M, Czyz ME (2012) Sphere formation and self-renewal capacity of melanoma cells is affected by the microenvironment. Melanoma Res 22(3):215–224. https://doi.org/10.1097/CMR.0b013e3283531317
Sztiller-Sikorska M, Hartman ML, Talar B, Jakubowska J, Zalesna I, Czyz M (2015) Phenotypic diversity of patient-derived melanoma populations in stem cell medium. Lab Invest 95(6):672–683. https://doi.org/10.1038/labinvest.2015.48
Talar B, Gajos-Michniewicz A, Talar M, Chouaib S, Czyz M (2016) Pentoxifylline inhibits WNT signaling in β-cateninhigh patient-derived melanoma cell populations. PLoS ONE 11(6):e0158275. https://doi.org/10.1371/journal.pone.0158275
Wang Y, McAlpine SR (2015) Heat-shock protein 90 inhibitors: will they ever succeed as chemotherapeutics? Future Med Chem 7(2):87–90. https://doi.org/10.4155/fmc.14.154
López I, Tournillon AS, Prado Martins R, Karakostis K, Malbert-Colas L, Nylander K, Fåhraeus R (2017) p53-mediated suppression of BiP triggers BIK-induced apoptosis during prolonged endoplasmic reticulum stress. Cell Death Differ 24(10):1717–1729. https://doi.org/10.1038/cdd.2017.96
Neckers L, Blagg B, Haystead T, Trepel JB, Whitesell L, Picard D (2018) Methods to validate Hsp90 inhibitor specificity, to identify off-target effects, and to rethink approaches for further clinical development. Cell Stress Chaperones 23(4):467–482. https://doi.org/10.1007/s12192-018-0877-2
Kijima T, Prince TL, Tigue ML, Yim KH, Schwartz H, Beebe K, Lee S, Budzynski MA, Williams H, Trepel JB, Sistonen L, Calderwood S, Neckers L (2018) HSP90 inhibitors disrupt a transient HSP90-HSF1 interaction and identify a noncanonical model of HSP90-mediated HSF1 regulation. Sci Rep 8(1):6976. https://doi.org/10.1038/s41598-018-25404-w
Zorzi E, Bonvini P (2011) Inducible hsp70 in the regulation of cancer cell survival: analysis of chaperone induction, expression and activity. Cancers (Basel) 3(4):3921–3956. https://doi.org/10.3390/cancers3043921
Davis AL, Cabello CM, Qiao S, Azimian S, Wondrak GT (2013) Phenotypic identification of the redox dye methylene blue as an antagonist of heat shock response gene expression in metastatic melanoma cells. Int J Mol Sci 14(2):4185–4202. https://doi.org/10.3390/ijms14024185
Lee CH, Hong HM, Chang YY, Chang WW (2012) Inhibition of heat shock protein (Hsp) 27 potentiates the suppressive effect of Hsp90 inhibitors in targeting breast cancer stem-like cells. Biochimie 94(6):1382–1389. https://doi.org/10.1016/j.biochi.2012.02.034
Ghadban T, Dibbern JL, Reeh M, Miro JT, Tsui TY, Wellner U, Izbicki JR, Güngör C, Vashist YK (2017) HSP90 is a promising target in gemcitabine and 5-fluorouracil resistant pancreatic cancer. Apoptosis 22(3):369–380. https://doi.org/10.1007/s10495-016-1332-4
Schaefer S, Svenstrup TH, Guerra B (2017) The small-molecule kinase inhibitor D11 counteracts 17-AAG-mediated up-regulation of HSP70 in brain cancer cells. PLoS ONE 12(5):e0177706. https://doi.org/10.1371/journal.pone.0177706
Rebecca VW, Wood E, Fedorenko IV, Paraiso KH, Haarberg HE, Chen Y, Xiang Y, Sarnaik A, Gibney GT, Sondak VK, Koomen JM, Smalley KS (2014) Evaluating melanoma drug response and therapeutic escape with quantitative proteomics. Mol Cell Proteomics 13(7):1844–1854. https://doi.org/10.1074/mcp.M113.037424
Murphy ME (2013) The HSP70 family and cancer. Carcinogenesis 34(6):1181–1188. https://doi.org/10.1093/carcin/bgt111
Madden E, Logue SE, Healy SJ, Manie S, Samali A (2019) The role of the unfolded protein response in cancer progression: from oncogenesis to chemoresistance. Biol Cell 111(1):1–17. https://doi.org/10.1111/boc.201800050
Hill DS, Lovat PE, Haass NK (2014) Induction of endoplasmic reticulum stress as a strategy for melanoma therapy: is there a future? Melanoma Manag 1(2):127–137. https://doi.org/10.2217/mmt.14.16
Sykes EK, Mactier S, Christopherson RI (2016) Melanoma and the unfolded protein response. Cancers (Basel) 8(3):E30. https://doi.org/10.3390/cancers8030030
Hartman ML, Czyz M (2013) Anti-apoptotic proteins on guard of melanoma cell survival. Cancer Lett 331(1):24–34. https://doi.org/10.1016/j.canlet.2013.01.010
Hartman ML, Czyz M (2015) Pro-survival role of MITF in melanoma. J Invest Dermatol 135(2):352–358. https://doi.org/10.1038/jid.2014.319
Cerezo M, Lehraiki A, Millet A, Rouaud F, Plaisant M, Jaune E, Botton T, Ronco C, Abbe P, Amdouni H, Passeron T, Hofman V, Mograbi B, Dabert-Gay AS, Debayle D, Alcor D, Rabhi N, Annicotte JS, Héliot L, Gonzalez-Pisfil M, Robert C, Moréra S, Vigouroux A, Gual P, Ali MMU, Bertolotto C, Hofman P, Ballotti R, Benhida R, Rocchi S (2016) Compounds triggering ER stress exert anti-melanoma effects and overcome BRAF inhibitor resistance. Cancer Cell 29(6):805–819. https://doi.org/10.1016/j.ccell.2016.04.013
Eigner K, Filik Y, Mark F, Schütz B, Klambauer G, Moriggl R, Hengstschläger M, Stangl H, Mikula M, Röhrl C (2017) The unfolded protein response impacts melanoma progression by enhancing FGF expression and can be antagonized by a chemical chaperone. Sci Rep 7(1):17498. https://doi.org/10.1038/s41598-017-17888-9
Ojha R, Leli NM, Onorati A, Piao S, Verginadis II, Tameire F, Rebecca VW, Chude CI, Murugan S, Fennelly C, Noguera-Ortega E, Liu S, Xu X, Krepler C, Xiao M, Xu W, Wei Z, Frederick DT, Boland G, Mitchell TC, Karakousis GC, Schuchter LM, Flaherty KT, Zhang G, Herlyn M, Koumenis C, Amaravadi RK (2018) ER translocation of the MAPK pathway drives therapy resistance in BRAF mutant melanoma. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-18-0348
Matos L, Gouveia AM, Almeida H (2015) ER stress response in human cellular models of senescence. J Gerontol A Biol Sci Med Sci 70(8):924–935. https://doi.org/10.1093/gerona/glu129
Xiang C, Wang Y, Zhang H, Han F (2017) The role of endoplasmic reticulum stress in neurodegenerative disease. Apoptosis 22(1):1–26. https://doi.org/10.1007/s10495-016-1296-4
Chen C, Zhang X (2017) IRE1α-XBP1 pathway promotes melanoma progression by regulating IL-6/STAT3 signaling. J Transl Med 15(1):42. https://doi.org/10.1186/s12967-017-1147-2
Croft A, Tay KH, Boyd SC, Guo ST, Jiang CC, Lai F, Tseng HY, Jin L, Rizos H, Hersey P, Zhang XD (2014) Oncogenic activation of MEK/ERK primes melanoma cells for adaptation to endoplasmic reticulum stress. J Invest Dermatol 134(2):488–497. https://doi.org/10.1038/jid.2013.325
Tay KH, Luan Q, Croft A, Jiang CC, Jin L, Zhang XD, Tseng HY (2014) Sustained IRE1 and ATF6 signaling is important for survival of melanoma cells undergoing ER stress. Cell Signal 26(2):287–294. https://doi.org/10.1016/j.cellsig.2013.11.008
Marcu MG, Doyle M, Bertolotti A, Ron D, Hendershot L, Neckers L (2002) Heat shock protein 90 modulates the unfolded protein response by stabilizing IRE1alpha. Mol Cell Biol 22(24):8506–8513. https://doi.org/10.1128/MCB.22.24.8506-8513.2002
Mimura N, Fulciniti M, Gorgun G, Tai YT, Cirstea D, Santo L, Hu Y, Fabre C, Minami J, Ohguchi H, Kiziltepe T, Ikeda H, Kawano Y, French M, Blumenthal M, Tam V, Kertesz NL, Malyankar UM, Hokenson M, Pham T, Zeng Q, Patterson JB, Richardson PG, Munshi NC, Anderson KC (2012) Blockade of XBP1 splicing by inhibition of IRE1α is a promising therapeutic option in multiple myeloma. Blood 119(24):5772–5781. https://doi.org/10.1182/blood-2011-07-366633
Urra H, Dufey E, Lisbona F, Rojas-Rivera D (1833) Hetz C (2013) When ER stress reaches a dead end. Biochim Biophys Acta 12:3507–3517. https://doi.org/10.1016/j.bbamcr.2013.07.024
Van Allen EM, Wagle N, Sucker A, Treacy DJ, Johannessen CM, Goetz EM, Place CS, Taylor-Weiner A, Whittaker S, Kryukov GV, Hodis E, Rosenberg M, McKenna A, Cibulskis K, Farlow D, Zimmer L, Hillen U, Gutzmer R, Goldinger SM, Ugurel S, Gogas HJ, Egberts F, Berking C, Trefzer U, Loquai C, Weide B, Hassel JC, Gabriel SB, Carter SL, Getz G, Garraway LA, Schadendorf D, Dermatologic Cooperative Oncology Group of Germany (DeCOG) (2014) The genetic landscape of clinical resistance to RAF inhibition in metastatic melanoma. Cancer Discov 1:94–109
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW, Greninger P, Thompson IR, Luo X, Soares J, Liu Q, Iorio F, Surdez D, Chen L, Milano RJ, Bignell GR, Tam AT, Davies H, Stevenson JA, Barthorpe S, Lutz SR, Kogera F, Lawrence K, McLaren-Douglas A, Mitropoulos X, Mironenko T, Thi H, Richardson L, Zhou W, Jewitt F, Zhang T, O’Brien P, Boisvert JL, Price S, Hur W, Yang W, Deng X, Butler A, Choi HG, Chang JW, Baselga J, Stamenkovic I, Engelman JA, Sharma SV, Delattre O, Saez-Rodriguez J, Gray NS, Settleman J, Futreal PA, Haber DA, Stratton MR, Ramaswamy S, McDermott U, Benes CH (2012) Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483(7391):570–575. https://doi.org/10.1038/nature11005
Chang CH, Drechsel DA, Kitson RR, Siegel D, You Q, Backos DS, Ju C, Moody CJ, Ross D (2014) 19-substituted benzoquinone ansamycin heat shock protein-90 inhibitors: biological activity and decreased off-target toxicity. Mol Pharmacol 85(6):849–857. https://doi.org/10.1124/mol.113.090654
Siegel D, Yan C, Ross D (2012) NAD(P)H:quinone oxidoreductase 1 (NQO1) in the sensitivity and resistance to antitumor quinones. Biochem Pharmacol 83(8):1033–1040. https://doi.org/10.1016/j.bcp.2011.12.017
Muñoz IG, Morel B, Medina-Carmona E, Pey AL (2017) A mechanism for cancer-associated inactivation of NQO1 due to P187S and its reactivation by the consensus mutation H80R. FEBS Lett 591(18):2826–2835. https://doi.org/10.1002/1873-3468.12772
Gaspar N, Sharp SY, Pacey S, Jones C, Walton M, Vassal G, Eccles S, Pearson A, Workman P (2009) Acquired resistance to 17-allylamino-17-demethoxygeldanamycin (17-AAG, tanespimycin) in glioblastoma cells. Cancer Res 69(5):1966–1975. https://doi.org/10.1158/0008-5472.CAN-08-3131
Grbovic OM, Basso AD, Sawai A, Ye Q, Friedlander P, Solit D, Rosen N (2006) V600E B-Raf requires the Hsp90 chaperone for stability and is degraded in response to Hsp90 inhibitors. Proc Natl Acad Sci USA 103(1):57–62. https://doi.org/10.1073/pnas.0609973103
Haarberg HE, Paraiso KH, Wood E, Rebecca VW, Sondak VK, Koomen JM, Smalley KS (2013) Inhibition of Wee1, AKT, and CDK4 underlies the efficacy of the HSP90 inhibitor XL888 in an in vivo model of NRAS-mutant melanoma. Mol Cancer Ther 12(6):901–912. https://doi.org/10.1158/1535-7163.MCT-12-1003
Corazzari M, Rapino F, Ciccosanti F, Giglio P, Antonioli M, Conti B, Fimia GM, Lovat PE, Piacentini M (2015) Oncogenic BRAF induces chronic ER stress condition resulting in increased basal autophagy and apoptotic resistance of cutaneous melanoma. Cell Death Differ 22(6):946–958. https://doi.org/10.1038/cdd.2014.183
Jiang CC, Chen LH, Gillespie S, Wang YF, Kiejda KA, Zhang XD, Hersey P (2007) Inhibition of mek sensitizes human melanoma cells to endoplasmic reticulum stress-induced apoptosis. Cancer Res 67:9750–9761. https://doi.org/10.1158/0008-5472.CAN-07-2047
Beck D, Niessner H, Smalley KS, Flaherty K, Paraiso KH, Busch C, Sinnberg T, Vasseur S, Iovanna JL, Drießen S, Stork B, Wesselborg S, Schaller M, Biedermann T, Bauer J, Lasithiotakis K, Weide B, Eberle J, Schittek B, Schadendorf D, Garbe C, Kulms D, Meier F (2013) Vemurafenib potently induces endoplasmic reticulum stress-mediated apoptosis in BRAFV600E melanoma cells. Sci Signal 6(260):ra7. https://doi.org/10.1126/scisignal.2003057
Ma XH, Piao SF, Dey S, McAfee Q, Karakousis G, Villanueva J, Hart LS, Levi S, Hu J, Zhang G, Lazova R, Klump V, Pawelek JM, Xu X, Xu W, Schuchter LM, Davies MA, Herlyn M, Winkler J, Koumenis C, Amaravadi RK (2014) Targeting ER stress-induced autophagy overcomes BRAF inhibitor resistance in melanoma. J Clin Invest 124(3):1406–1417. https://doi.org/10.1172/JCI70454
Acknowledgements
This study was funded by National Science Centre (Poland), Grant No. 2014/15/B/NZ7/00947. We thank Dr. Anna Gajos-Michniewicz for cell culture propagation and helpful discussion, and Ewa Lewandowska for excellent administrative and technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Mielczarek-Lewandowska, A., Sztiller-Sikorska, M., Osrodek, M. et al. 17-Aminogeldanamycin selectively diminishes IRE1α-XBP1s pathway activity and cooperatively induces apoptosis with MEK1/2 and BRAFV600E inhibitors in melanoma cells of different genetic subtypes. Apoptosis 24, 596–611 (2019). https://doi.org/10.1007/s10495-019-01542-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-019-01542-y