Abstract
Prosopis laevigata (mesquite; Fabaceae) forms fertility islands in soils of semi-arid lands where microbial diversity concentrates in response to the accumulation of resources in the soil beneath individual plants, promoting organic matter decomposition and nutrient cycling. This phenomenon provides suitable conditions for the proliferation of key edaphic elements such as fungi and mites. Mite-fungal interactions are central for our understanding of nutrient cycling processes in resource-limited arid food webs; yet, no information is available about fertility islands in semi-arid lands. Thus, we aimed to determine in vitro fungal-based feeding preferences and molecular gut content of the oribatid mite species Zygoribatula cf. floridana and Scheloribates cf. laevigatus, which are abundant under the canopy of P. laevigata in an intertropical semi-arid zone in Central Mexico. Our results on the gut content analysis of these oribatid species resulted in the ITS-based identification of the following fungi: Aspergillus homomorphus, Beauveria bassiana, Filobasidium sp., Mortierella sp., Roussoella sp., Saccharomyces cerevisiae, Sclerotiniaceae sp. and Triparticalcar sp. Furthermore, under laboratory conditions both oribatid mite species exhibited feeding preferences on melanized fungi, such as Cladosporium spp., whereas A. homomorphus and Fusarium penzigi were avoided. Our findings indicated that the analyzed oribatid mite species have similar feeding preferences for melanized fungi, which might suggest resource partitioning and a degree of preference, explaining the coexistence of both oribatid species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Desert soils are highly susceptible to degradation due to the reduced biological productivity, demanding an efficient nutrient mobilization in a short period of time (Whitford and Duval 2019). In these stressed environments, the canopy and the rhizosphere of trees enhance higher microbial and plant activities under them (fertility island effect), sustaining complex soil food webs during dryness, in contrast to sun-exposed or degraded areas (Ochoa-Hueso et al. 2018). Under these circumstances, complex biotic interactions proliferate in response to environmental stress or degradation pressures.
Plant health and nutrition depend on microbial communities, recycling nutrients via organic matter degradation (Klironomos and Kendrick 1995). In addition, the invertebrate community (mesofauna) facilitates microbial recycling of lignin, tannins, phenols, waxy cuticles, and other polymers (Stott and Martin 1989; Bailey et al. 2002; Schneider et al. 2004a, b; Walter and Proctor 2013). Among this invertebrate community, oribatid mites are acknowledged as important nutrient mobilizers (Hartenstein 1962; Siepel and Maaskamp 1994; Maraun et al. 1998; Schneider et al. 2004a, b).
Oribatid mites are the most abundant and active microarthropods in fertility islands under the arid-active trees Prosopis laevigata (Humb. & Bonpl. ex Willd.) M.C. Johnst. (mesquite) even during the driest part of the year in a semi-arid zone from southern Mexico (Rodríguez-Zaragoza et al. unpubl. data). Their activity has a strong impact on the biological productivity in arid systems (Santos and Whitford 1981; Cepeda-Pizarro and Whitford 1989; Klironomos and Kendrick 1995; Todria et al. 2021) where fungal-based food webs are characteristic due to fungal metabolic ability to exploit high molecular weight compounds such as litter and woody structures (Wardle et al. 2004). Oribatid mites, by grazing on fungal hyphae and spores, mobilize nutrients, control fungal dispersion and community structure (Behan-Pelletier and Hill 1978; Renker et al. 2005; Vašutová et al. 2019). Hence, oribatid mites enhance the decomposition rates, nutrient cycling, microbial respiration, and plant growth (Siepel and Maaskamp 1994; Zak and Whitford 1988; Crowther et al. 2012).
Zygoribatula floridana and Scheloribates laevigatus are cosmopolitan oribatid mite species (Murvanidze et al. 2018; Subias 2022). These taxa have been reported from resource-limited and stressful systems, such as arid and semi-arid areas (Neher et al. 2009; Krantz and Walter 2009) feeding potentially on litter (Hubert et al. 2000; Wickings and Grandy 2011; Gergócs and Hufnagel 2016), lichen, algae (Erdmann et al. 2007), and nematodes (Rockett and Woodring 1966; Muraoka and Ishibashi 1976; Rockett 1980; Ramakrishnan and Neravathu 2019). However, several reports suggest they prefer fungal hyphae and spores over the rest of potential items (Behan-Pelletier and Hill 1978; Siepel and de Ruiter-Dijkman 1993; Hubert et al. 2001, 2004; Maraun et al. 2023), placing them on the fungal-based food web. As fungal consumers, these oribatid mite species might compete for similar resources, posing the possibility of resource partitioning through selective feeding to avoid strong competition (Schneider and Maraun 2005).
Direct in situ observations have been attempted to elucidate mite-fungal interactions and their impact on the soil system (Smrž 1991, 2002; Hubert et al. 2001). However, these approaches face several methodological limitations such as: (1) the cryptic nature of both oribatid mite species in field, and (2) difficulties for distinguishing between parasitic fungi and those serving as food, since fungal and mite morphological identification of both fungi and mite taxa is difficult, time consuming, and the analysis of mites’ gut content is extremely challenging to perform in the field (Remén et al. 2010). At present, these limitations may be overcome by in vitro feeding experiments coupled with the analysis of gut contents through amplification of fungal ITS region (Velez et al. 2018). Moreover, knowledge of mites' fungal diet would provide more insights on the ‘fungal path’ of soil nutrients in semi-arid environments. However, the potential fungal resources on which abundant oribatid taxa feed in the surroundings of P. laevigata in a semiarid land remain unknown. Hence, we aimed to determine fungal taxa associated with oribatid mites Z. cf. floridana and S. cf. laevigatus, collected under the canopy of P. laevigata in a preserved and a degraded terrace during rainy season from an intertropical semiarid soil, as well as to determine their feeding preferences on the isolated fungi under laboratory conditions. We hypothesized that each of the terraces harbors a characteristic acarofauna that establishes close trophic interactions with the local mycobiota. This work provides major insights into how the oribatid mite species coexist within their specific assemblage.
Materials and methods
Zapotitlán belongs to the Tehuacán-Cuicatlán Valley Reserve, located in the southeast of the State of Puebla and northeast of the State of Oaxaca, Mexico (Fig. 1). Semi-aridity of this zone is due to the rain shadow effect, producing a 4-month rainy season each year. Climate is dry-hot with summer rains, corresponding to Köppen’s Bs class (Peel et al. 2007). The valley has an average temperature of 21 °C and 400–450 mm average rainfall (García 2004). Vegetation is xerophytic shrubland, dominated by legume trees like mesquite (P. laevigata), Palo Verde (Parkinsonia praecox), Mimosa luisiana and several species of cacti (Dávila et al. 2002).
The Zapotitlán area displays alluvial terraces deposited by ‘El Salado’ river during the Pleistocene (López-Galindo et al. 2003) with patches of well-developed herbaceous, shrub and scattered tree species (conserved patches) intermixed with patches having scarce or no vegetation (degraded patches). Several of these patches are transitioning into badlands (López-Galindo et al. 2003), where the fertility island effect of P. laevigata is well observed (González-Ruiz et al. 2008).
Samplings were conducted during the season of heavy rains (late May-late July 2018). Individual litter-soil samples were collected under the canopy of six P. laevigata trees at 10 cm depth from the conserved terrace (18°19′30.0″N, 97°27′12.3″W) and six more, with the same conditions, from the degraded one (18°19′34.8″N, 97°27′17.1″W). Samples were deposited in 2-kg Ziploc bags and transferred to the Laboratory of Microbial Ecology, FES Iztacala, UNAM, for extraction and sorting of oribatid mites.
Microarthropods were extracted from the samples using the Berlese funnel extraction method for 4 days and collected into plastic vials. Half of the vials contained 90% alcohol for mite preservation and identification (Crossley and Blair 1991). The other half of the vials contained moistened filter paper to collect mites alive for culturing and feeding preferences tests.
For morphological identification, adult specimens were, first, macerated with lactic acid for increasing transparency and a better microscopic observation. Second, mites were transferred to permanent slide mounts with Hoyer’s medium for their identification (Krantz and Walter 2009). The keys of Franklin et al. (2008) and Balogh and Balogh (1990) were used to identify the specimens by morphology achieving identification to genus or species level.
From the mites collected in vials with alcohol, 12 mites from each of the two most abundant taxa were separated and individually stored in 2-ml Eppendorf tubes with 100% alcohol, for gut dissection and molecular analysis. Cuticles of these mites were sanitized prior to gut dissection to ensure the absence of external contamination for DNA analysis of gut content. Thus, individual mites were transferred to 2-ml Eppendorf tubes and vortexed 30 s in a solution of 1 ml sterile water and 0.01% Tween 80, for a period of 30 s twice in 1 ml sterile water (Velez et al. 2018). Subsequently, under sterile conditions and using a stereomicroscope (Nikon), the exoskeleton was opened by inserting an insect needle (no. 00) in the base of the notogaster (separating the ventral and dorsal plates of the body), revealing the gut. Next, the gut was extracted with another sterile insect needle and placed individually in Eppendorf tubes for DNA extraction.
Additionally, another 12 specimens corresponding to the most abundant oribatid taxa were extracted and their cuticles were processed for fungal isolation, due to its potential to carry fungi (Remén et al. 2010). Each individual mite was placed in a sterile Eppendorf tube containing a solution of 1 ml distilled water and 0.01% Tween 80, and then vortexed for 1 min. Subsequently, 500 µl was plated into Petri dishes with Potato Dextrose Agar (PDA; supplemented with 0.5 g/ml of benzathine benzylpenicillin to prevent bacterial growth) and incubated at room temperature (22 ± 3 °C) for 2 weeks. Developing fungi were isolated and grouped based on the macroscopic characteristics. We selected and sub-cultured the top-20 most abundant isolates for subsequent feeding preference tests. All of the isolates were identified and deposited in the Laboratory C-121, Institute of Biology, UNAM, and are available for research upon request. For identification, the genomic DNA of each isolate was obtained using the protocol described by Doyle and Doyle (1987). The ITS1-5.8S-ITS4 region of rDNA was amplified using the primer set ITS 1 and ITS 4 (5′-TCCGTAGGTGAACCTGCGG-3′ and 5′-TCCTCCGCTTATTGATATGC-3′) (White 1990). PCR products were sequenced in both directions by the Laboratory of Biodiversity and Health Genomic Sequencing, Institute of Biology, UNAM.
The fungal ITS region of DNA mixture obtained from the dissected guts was directly amplified using the Thermo Scientific Phire Animal Tissue kit following the manufacturers’ instructions, with the primer set ITS 1 and ITS 4 (White et al. 1990). In this experiment, we included positive (Cladosporium cladosporioides) and negative (no mite’s gut in the tube mix) controls. The PCR products were visualized on a 1% agarose gel stained with GelRed to verify amplification products. Electrophoresis revealed amplification in nearly all samples, but only strongly stained and well-defined bands of amplified DNA were considered for sequencing in the Laboratory of Biodiversity and Health Genomic Sequencing.
Fungal sequences from both experiments (fungal isolation and gut content analysis) were manually edited and trimmed using BioEdit software (v.7.2; Hall 1999), and were compared to the GenBank database, using Basic Local Alignment Search Tool (BLASTn) (Altschul et al. 1990). For each OTU, several sequences, preferably from published studies, with a minimum cut-off value of 98–100% for presumed species and 94–97% for genus level, were considered (Millberg et al. 2015).
Adult specimens of the abundant taxa collected in vials with moistened filter paper were sorted and subsequently cultured in 100-ml glass vials with Paris plaster and charcoal (9:1), soil litter from the sampling site, and Baker’s yeast as diet. Temperature and humidity in cultures were kept at 20 ± 1 °C and 60 ± 10%, before feeding preference tests (Krantz and Walter 2009). Prior to tests, the oribatid mite specimens in culture were starved for 5 days. Tests were conducted in 100-ml glass containers, where 15 individuals of Z. cf. floridana and S. cf. laevigatus mites were added in separated containers. Thus, four tests were conducted according to the sampling site where fungi and oribatid mites were extracted: (A) individuals of Z. cf. floridana collected from a conserved area, (B) individuals of Z. cf. floridana from a degraded area, (C) individuals of S. cf. laevigatus from a conserved area, and (D) individuals of S. cf. laevigatus collected from a degraded area (Fig. 2). Five replicates were used per test. Each test contained five fungal species collected from their respective oribatid cuticle and baker’s yeast as control (Maraun et al. 1998; Hubert et al. 2001; Schneider et al. 2004a, b). Observations under a stereomicroscope (Nikon) were made every 30 min, until nearly all food items were depleted, so the test duration depended on the intensity of consumption. Furthermore, three criteria were considered to determine feeding preferences: (1) preferred, where > 6 mites in the glass vial stayed consuming the food choice; (2) transitory, where < 6 mites consumed the food choice; and (3) avoided, no mites approached or consumed the food choice.
Feeding preference tests including five fungal species and a control (baker’s yeast). Zygoribatula cf. floridana collected in a conserved area (A) and a degraded area (B). Scheloribates cf. laevigatus collected in a conserved area (C) and in a degraded area (D). Aspergillus homomorphus (Aho), Chaetomiaceae sp. (Cha), Cladosporium cladosporioides (Ccl), Cladosporium sp. (Csp), control (Ctrl), Didmellaceae sp. (Didi), Endomycetaceae sp. (End), Fusarium equisetum (Feq), Fusarium penzigii (Fpe), Leptosphaerulina australis (La), Mortierella schmuckeri (Msc), Mortierella sp. (Msp), Penicillium olsonii (Pol), Sarcopodium sp. (Ssp)
For this study, sampling sites were not considered variables which might have influenced the feeding preference test, as mites and fungi were cultured afterwards under laboratory conditions and fed with the same diet before tests. Hence, the sampling site effect was disregarded. To approach diversity differences in terms of acarofauna, a principal component analysis (PCA) was performed using R software (R core team 2021) with the vegan package (v.2.6–4) (Oksanen et al. 2017) using the oribatid taxa abundance against both terraces (conserved terrace and degraded terrace). Furthermore, a Kruskal–Wallis test was performed to determine significant differences in feeding preferences. Subsequently, a Dunn post-hoc test was used to contrast differences among treatments. Statistical analysis was carried out using R (R Core Team 2021).
Results
The oribatid mites Z. cf. floridana and S. cf. laevigatus were the most abundant oribatid taxa collected under the canopies of P. laevigata from both terraces. These species belong to families Oribatulidae and Scheloribatidae, respectively, both with a worldwide distribution (Subias 2022), and accounting for 97 (Z. cf. floridana) and 75 (S. cf. laevigatus) individuals (Table S1), being Z. cf. floridana the most abundant in both sites.
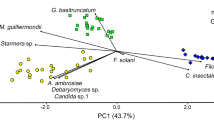
The PCA biplot (Fig. 3) displayed the differences in oribatid mite taxa assemblage between both terraces. PC1 and PC2 explained 52.4 and 15.1% of the variance, respectively. Nearly half (12 specimens) of the oribatid individuals dissected displayed traces of fungal DNA from gut content DNA analysis. However, only a fraction of DNA amplified from mite gut extractions yielded to fungal DNA suitable material for sequencing and identification, which indicates that fungal DNA might degrade between sample collection and amplification, reducing its detectability. This might also be due to individuals feeding at different times and rates on a variety of sources as they are heterogeneously distributed in soil (Rohde 1959; Anderson and Healey 1972).
The amplified ITS sequences from mites’ gut produced fragments of different sizes, ranging from 500 bp (Mortierella sp.) to 200 bp (Aspergillus sp.). Based on these sequences, eight fungal species from the gut of oribatid mites were identified: Beauveria bassiana, Filobasidium sp., Saccharomyces cerevisiae and Triparticalcar sp. in the gut of Z. cf. floridana, and Aspergillus homomorphus, Mortierella sp., Roussoella sp. and Sclerotiniaceae sp. in the gut of S. cf. laevigatus (Table 1).
Among the 20 fungi isolated from the cuticle of several individuals of both oribatid mites, C. cladosporioides was the most representative fungus species, followed by Mortierella schmuckeri. Further isolates obtained from the cuticle of mites included: A. homomorphus and two unidentified species of Cladosporium (sp. 1 and sp. 2), Fusarium equiseti, Fusarium penzigii, Leptosphaerulina australis, Mortierella sp., Penicillium olsoni and Sarcopodium sp. (Table 2).
Differences in the feeding preferences by the two oribatid taxa were observed (Fig. 4). Zigoribatula cf. floridana showed a marked preference for Cladosporium sp. 1 (test A; H5, 0.5 = 25.28, P < 0.05) which was consumed until depletion (Table 3). However, Z. cf. floridana from test B displayed a strong preference on M. schmuckeri (H5, 0.5 = 28.78, P < 0.05) whereas A. homomorphus was avoided (Table 3).
Boxplot depicting feeding preferences of Zygoribatula cf. floridana (Zyg1 in test 1 and Zyg2 in test 2) and Scheloribates cf. laevigatus (Sch1 in test 3 and Sch2 in test 4). The boxes display the interquartile range, and the bold line displays the median; whiskers of the boxplots represent the upper and lower quartiles (25th and 75th percentiles). Fungal choices: Aho—Aspergillus homomorphus, Chae—Chaetomiaceae sp., Ccl—Cladosporium cladosporioides, Csp—Cladosporium sp., Ctrl—control, Didi—Didmellaceae sp., Endo—Endomycetaceae sp., Feq—Fusarium equisetum, Fpe—Fusarium penzigii, Lau—Leptosphaerulina australis, Msc—Mortierella schmuckeri, Msp—Mortierella sp., Pol—Penicillium olsonii, Ssp—Sarcopodium sp. (n = 4)
Scheloribates cf. laevigatus from test C preferred to feed on C. cladosporioides and Sarcopodium sp. (H5, 0.5 = 26.99, P < 0.05) although neither of these fungi were its first choice. Scheloribates cf. laevigatus fed from them towards the end of the tests. Fusarium penzigii was always avoided in all treatments (Table 4). Scheloribates cf. laevigatus from test D preferred C. cladosporioides and frequently avoided Mortierella sp. and Endomicetacea sp. (H5, 0.5 = 28.21, P < 0.05; Table 4). Baker’s yeast was consumed in nearly all tests, although mites looked first for other suitable food.
Discussion
In this study, both Z. cf. floridana and S. cf. laevigatus were abundant on both terraces. We hypothesized to find a particular oribatid mites’ community assemblage on both terraces due to different soil degradation processes affecting them. This suggests that P. laevigata provides the minimum conditions necessary to harbor a diversity of soil fungi, which is reflected in the abundance of both oribatid taxa on the terraces, since they might feed on a great variety of fungi (Behan-Pelletier and Hill 1978).
Under laboratory conditions, Z. cf. floridana and S. cf. laevigatus shared feeding preferences on several fungal species, which might suggest either competition or resource partitioning between these two taxa in their environment. However, resource partitioning is a plausible explanation for their co-occurrence in soil, i.e., high availability and diversity of food resources in a heterogeneous environment, or consumption of fungal spores by one oribatid taxon and consumption of hyphae by the other (Koukol et al. 2009).
Both oribatid mites fed on C. cladosporioides until depletion, accompanied by a massive production of feces, which according to Smrž and Norton (2004), might be due to intensive feeding for nutrient assimilation from sources with very low nutrient content which is reflected in a massive production of feces with fragments of partially digested fungal structures. It is likely that C. cladosporioides in the field benefits from this mechanism for spore attachment to the mite cuticle while feeding on hyphae, which offers a colonizing advantage for this fungus being dispersed via cuticle transport or fecal deposition, favoring resource colonization and exploitation (Visser et al. 1981; Williams et al. 1998). Preference for melanized taxa such as Cladosporium was also reported by Schneider and Maraun (2005).
The fungus A. homomorphus was avoided all the time by Z. cf. floridana, causing mites to seek for other available fungi during the food choices. However, traces of A. homomorphus were found in the gut of both oribatid taxa collected in the field. Probably, A. homomorphus faces a trade-off between growing or producing toxic compounds which may repel mites during hyphal development (Koukol et al. 2009; Chatterjee et al. 2016). Thus, this fungus might become a more palatable food to mites if hyphae are produced fast, reducing the quantity and quality of toxic molecules (Behan-Pelletier and Hill 1978; Hubert et al. 2003; Velez et al. 2018).
Mortierella spp. hyphae have been reported to be undesirable for microarthropods (Dix and Webster 1995; Maraun et al. 1998; Werner et al. 2016; Yoder et al. 2019). However, in this study Z. cf. floridana preferred to feed on Mortierella sp., and although neither M. schmuckeri nor Mortierella sp. were their first choice, individual mites fed on them. This degree of preference could explain the occurrence of this fungal species as the second most reported fungal species on the mites’ cuticle.
Few individuals of Z. cf. floridana fed on P. olsonii. However, the interaction among the two taxa is not clear as this fungus has been acknowledged to interact differently with oribatid mites. For example, Hartenstein (1962) and Seniczak et al. (2017) reported that Penicillium is repellent to oribatids, whereas Seniczak and Stefaniak (1981) and Hubert et al. (1999) observed that Penicillium was attractive to a different set of mite species, which suggests a complex species-specific relationship among the taxa. Thus, further investigations of this isolate are needed to better understand its interaction with soil mites.
The direct amplification of the fungal barcode region from mites’ gut content is a reliable tool to analyze feeding specificity in the mite-fungus interaction. However, certain factors hamper a good DNA identification. i.e., time after the last food intake. Nonetheless, this minor deviation may be corrected by increasing sample size, as well as by feeding preference tests under laboratory conditions. In this sense, our results showed the presence of different fungal species such as A. homomorphus, Mortierella sp. and S. cerevisiae, in the gut of Z. cf. floridana and S. cf. laevigatus, which may suggest that the analyzed oribatids fulfill their different nutritional needs based on a mixed-fungal diet that provides an assortment of nutrients to successfully fulfill their life cycle (Gan et al. 2014; Maraun et al. 2020).
As a marginal finding, we report the entomopathogenic fungus B. bassiana from the gut of S. cf. laevigatus. This fungus is used as biological pest-control agent against several species of phytoparasitic mites (Jiang et al. 2019). However, the lack of evident disease symptoms and the presence of B. bassiana in the gut contents of the herein examined individuals indicates that hyphae of this fungus may serve as a food source for S. cf. laevigatus, implying some degree of tolerance to toxic molecules, such as beauvericin and beauverolid (Pedrini 2022) and the active role of mites in fungal dispersion. This agrees with former observations reporting the asymptomatic association of B. bassiana with Paradamaeus clavipes (Renker et al. 2005).
The occurrence of yeasts in the gut of oribatid mites is not rare, although these microfungi are scarcely reported from soil (Princz et al. 2010; Molva et al. 2019). Our results suggest that Z. cf. floridana and S. cf. laevigatus were feeding on S. cerevisiae in soil, as it has been previously observed in various feeding preferences tests (van Bronswijk 1971; Andersen 1991; Princz et al. 2010; Molva et al. 2019). Moreover, its occurrence as an endosymbiotic fungus is plausible since several yeast species have been reported as beneficial to mites establishing mutualistic relationships, that provide the host with vitamins or sterols (Ganter 2006; Douglas 2009; Chandler et al. 2012).
The presence of Filobasidium sp., Roussoella sp. and Sclerotiniaceae sp. in the gut content of S. cf. laevigatus resembles previous work describing the high abundance of these fungi in the soil and in association with plants, particularly in the cases of saprophagous Roussoella sp. (Liu et al. 2014) or the putative facultative phytopathogens Sclerotiniaceae sp. (Adams and Ayers 1979). However, the chemical interactions between host plants, phytopathogens, and mites (e.g., S. cf. laevigatus) are still open for research.
In contrast to our hypothesis, our observations indicate that Z. cf. floridana and S. cf. laevigatus have similar fungal-based dietary inclinations, as both oribatid mites share preference for Cladosporium spp. and Mortierella spp. Nevertheless, we observed distinct degrees of preference between both oribatid taxa, which may facilitate an intensive appropriation of fungal resources based on resource partitioning. This feeding behavior might represent a suitable way to avoid competition in a stressful environment such as arid soils. Finally, our findings confirmed that the molecular approach is very useful for the establishment of in situ (in the environment) feeding preferences. However, it is necessary to conduct complementary studies, including a deeper microbiome analysis to discard food sources from endosymbionts. This is needed for understanding how oribatid mites affect fungal-based food-webs in arid and semiarid lands. Microbiome plays an important role in digesting preferred fungal prey due to the molecular specificities of the fungal cell wall (Smrž et al. 2016), it is likely that Z. cf. floridana and S. cf. laevigatus share mutualistic microbiota capable of degrading similar hyphae components.
References
Adams P, Ayers W (1979) Ecology of Sclerotinia species. Phytopathology 69(8):896–898
Altschul S, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Andersen A (1991) Nutritional value of yeast for Dermatophagoides pteronyssinus (Acari: Epidermoptidae) and the antigenic and allergenic composition of extracts during extended culturing. J Med Entomol 28:487–491
Anderson JM, Healey IN (1972) Seasonal and inter-specific variation in major components of the gut contents of some Woodland Collembola. J Anim Ecol 41:359. https://doi.org/10.2307/3473
Bailey VL, Smith JL, Bolton H (2002) Fungal-to-bacterial ratios in soils investigated for enhanced C sequestration. Soil Biol Biochem 34:997–1007. https://doi.org/10.1016/S0038-0717(02)00033-0
Balogh J, Balogh P (1990) Oribatid mites of the Neotropical Region II. Akademiai Kiado, Budapest
Behan-Pelletier VM, Hill SB (1978) Feeding habits and spore dispersal of oribatid mites (Acari, Oribatida) in the North American Arctic. Rev Ecol (terre Vie) 15:497–516
Cepeda-Pizarro JG, Whiford WG (1989) Species abundance distribution patterns of microarthropods in surface decomposing leaf-litter and mineral soil on a desert watershed. Pedobiologia 33:254–268
Chandler JA, Eisen JA, Kopp A (2012) Yeast communities of diverse Drosophila species: comparison of two symbiont groups in the same hosts. Appl Environ Microbiol 78:7327–7336. https://doi.org/10.1128/AEM.01741-12
Chatterjee S, Kuang Y, Splivallo R, Chatterjee P, Karlovsky P (2016) Interactions among filamentous fungi Aspergillus niger, Fusarium verticillioides and Clonostachys rosea: fungal biomass, diversity of secreted metabolites and fumonisin production. BMC Microb 16(1):83. https://doi.org/10.1186/s12866-016-0698-3
Crossley DA, Blair JM (1991) A high-efficiency, “low-technology” Tullgren-type extractor for soil microarthropods. Agric Ecosyst Environ 34:187–192. https://doi.org/10.1016/0167-8809(91)90104-6
Crowther TW, Boddy L, Hefin-Jones T (2012) Functional and ecological consequences of saprotrophic fungus-grazer interactions. ISME J 6:1992–2001. https://doi.org/10.1038/ismej.2012.53
Dávila P, Arizmendi MDC, Valiente-Banuet A et al (2002) Biological diversity in the Tehuacán-Cuicatlán Valley, Mexico. Biodivers Conserv 11:421–442. https://doi.org/10.1023/A:1014888822920
Dix NJ, Webster J (1995) The Mycelium and Substrates for Growth. In: Dix NJ (ed) Fungal Ecology. Springer, Netherlands, pp 12–38. https://doi.org/10.1007/978-94-011-0693-1_2
Douglas AE (2009) The microbial dimension in insect nutritional ecology. Funct Ecol 23:38–47
Doyle J, Doyle J (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Erdmann G, Otte V, Langel R, Scheu S, Maraun M (2007) The trophic structure of bark-living oribatid mite communities analysed with stable isotopes (15N, 13C) indicates strong niche differentiation. Exp Appl Acarol 41(1–2):1–10. https://doi.org/10.1007/s10493-007-9060-7
Franklin E, Norton RA, Crossley DA (2008) Zygoribatula colemani sp. nov (Acari, Oribatida, Oribatulidae) from granite outcrops in Georgia USA, with a highly variable translamella. Zootaxa. https://doi.org/10.11646/zootaxa.1847.1.3
Gan H, Zak DR, Hunter MD (2014) Trophic stability of soil oribatid mites in the face of environmental change. Soil Biol Biochem 68:71–77. https://doi.org/10.1016/j.soilbio.2013.09.019
Ganter PF (2006) Yeast and invertebrate associations. In: Rosa CA, Péter G (eds) Biodiversity and Ecophysiology of Yeasts, 1st edn. Springer-Verlag, Berlin, pp 303–370
García E (2004) Modificaciones al Sistema de Clasificación Climática de Köppen. Instituto de Geografía, UNAM, México
Gergócs V, Hufnagel L (2016) The effect of microarthropods on litter decomposition depends on litter quality. Eur J Soil Biol 75:24–30
González-Ruiz T, Rodríguez-Zaragoza S, Ferrera-Cerrato R (2008) Fertility islands around Prosopis laevigata and Pachycereus hollianus in the drylands of Zapotitlán Salinas, México. J Arid Environ 72:1202–1212. https://doi.org/10.1016/j.jaridenv.2007.12.008
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hartenstein R (1962) Soil Oribatei. I. feeding specificity among forest soil Oribatei (Acarina). Ann Entomol Soc Am 55(2):202–206. https://doi.org/10.1093/aesa/55.2.202
Hubert J, Šustr V, Smrž J (1999) Feeding of the oribatid mite Scheloribates laevigatus (Acari: Oribatida) in laboratory experiments. Pedobiologia 43:328–339
Hubert J, Kubátová A, Šárová J (2000) Feeding of Scheloribates laevigatus (Acari: Oribatida) on different stadia of decomposing grass litter (Holcus lanatus). Pedobiologia 44(5):627–639. https://doi.org/10.1078/S0031-4056(04)70077-3
Hubert J, Žilová M, Pekár S (2001) Feeding preferences and gut contents of three panphytophagous oribatid mites (Acari: Oribatida). Eur J Soil Biol 37(3):197–208. https://doi.org/10.1016/S1164-5563(01)01083-4
Hubert J, Stejskal V, Kubátová A et al (2003) Mites as selective fungal carriers in stored grain habitats. Exp Appl Acarol 29:69–87. https://doi.org/10.1023/A:1024271107703
Hubert J, Jarošík V, Mourek J, Kubátová A, Ždárková E (2004) Astigmatid mite growth and fungi preference (Acari: Acaridida): comparisons in laboratory experiments. Pedobiologia 48(3):205–214. https://doi.org/10.1016/j.pedobi.2003.12.005
Jiang A, Yuan Y, Yang R et al (2019) Beauveria bassiana is a potential effective biological agent against Psoroptes ovis var. cuniculi mites. Biol Control 131:43–48. https://doi.org/10.1016/j.biocontrol.2019.01.010
Klironomos JN, Kendrick WB (1995) Stimulative effects of Arthropods on Endomycorrhizas of Sugar Maple in the presence of Decaying Litter. Funct Ecol 9:528. https://doi.org/10.2307/2390019
Koukol O, Mourek J, Janovský Z, Černá K (2009) Do oribatid mites (Acari: Oribatida) show a higher preference for ubiquitous vs. specialized saprotrophic fungi from pine litter? Soil Biol Biochem 41(6):1124–1131. https://doi.org/10.1016/j.soilbio.2009.02.018
Krantz G, Walter D (2009) A Manual of Acarology (Chapter 7), 3rd edn. Texas Tech University Press, Texas, pp 83–96
Liu JK, Phookamsak R, Dai DQ et al (2014) Roussoellaceae, a new pleosporalean family to accommodate the genera Neoroussoella gen. nov. Roussoella and Roussoellopsis. Phytotaxa 181:1–33. https://doi.org/10.11646/phytotaxa.181.1.1
López-Galindo F, Muñoz-Iniestra D, Hernández-Moreno M et al (2003) Análisis integral de la toposecuencia y su influencia en la distribución de la vegetación y la degradación del suelo en la Subcuenca de Zapotitlán Salinas. Puebla, Bol Soc Geol Mex 56:19–41. https://doi.org/10.18268/bsgm2003v56n1a3
Maraun M, Visser S, Scheu S (1998) Oribatid mites enhance the recovery of the microbial community after a strong disturbance. Appl Soil Ecol 9(1–3):175–181. https://doi.org/10.1016/S0929-1393(98)00072-9
Maraun M, Augustin D, Pollierer M, Scheu S (2020) Variation in trophic niches of oribatid mites in temperate forest ecosystems as indicated by neutral lipid fatty acid patterns. Exp Appl Acarol 81:103–115. https://doi.org/10.1007/s10493-020-00494-2
Maraun M, Thomas T, Fast E et al (2023) New perspectives on soil animal trophic ecology through the lens of C and N stable isotope ratios of oribatid mites. Soil Biol & Biochem. https://doi.org/10.1016/j.soilbio.2022.108890
Millberg H, Boberg J, Stenlid J (2015) Changes in fungal community of Scots pine (Pinus sylvestris) needles along a latitudinal gradient in Sweden. Fungal Ecol 17:126–139. https://doi.org/10.1016/j.funeco.2015.05.012
Molva V, Nesvorna M, Hubert J (2019) Feeding interactions between microorganisms and the house dust mites Dermatophagoides pteronyssinus and Dermatophagoides farinae (Astigmata: Pyroglyphidae). J Med Entomol 56:1669–1677. https://doi.org/10.1093/jme/tjz089
Muraoka M, Ishibashi N (1976) Nematode-feeding mites and their feeding behaviour. Appl Entomol Zool 11(1):1–7
Murvanidze M, Todria N, Mumladze L, Kalatozishvili L (2018) Diversity of soil mite communities in different habitats of Saskhori quarries Georgia. Persian J Acarol 7:297–305. https://doi.org/10.22073/pja.v7i3.37647
Neher DA, Lewins SA, Weicht TR, Darby BJ (2009) Microarthropod communities associated with biological soil crusts in the Colorado Plateau and Chihuahuan deserts. J Arid Environ 73(6–7):672–677. https://doi.org/10.1016/j.jaridenv.2009.01.013
Ochoa-Hueso R, Eldridge DJ, Delgado-Baquerizo M et al (2018) Soil fungal abundance and plant functional traits drive fertile island formation in global drylands. J Ecol 106:242–253. https://doi.org/10.1111/1365-2745.12871
Oksanen FJ et al (2017) Vegan: Community Ecology Package. R package Version 2.4–3. https://CRAN.R-project.org/package=vegan
Pedrini N (2022) The entomopathogenic fungus Beauveria bassiana shows its toxic side within insects: expression of genes encoding secondary metabolites during pathogenesis. J Fungi 8(5):488. https://doi.org/10.3390/jof8050488
Peel MC, Finlayson BL, McMahon TA (2007) Updated world map of the Köppen-Geiger climate classification. Hydrol Earth Syst Sci 11:1633–1644. https://doi.org/10.5194/hess-11-1633-2007
Princz JI, Behan-Pelletier VM, Scroggins RP, Siciliano SD (2010) Oribatid mites in soil toxicity testing-the use of Oppia nitens (C.L. Koch) as a new test species. Environ Toxicol Chem 29:971–979. https://doi.org/10.1002/etc.98
R Core Team (2021) R: A language and environment for statistical computing. R foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Ramakrishnan N, Neravathu R (2019) Oribatid mites as potential predators of the root knot nematode. Meloidogyne Incognita Acarological Studies 1(2):123–128
Remén C, Krüger M, Cassel-Lundhagen A (2010) Successful analysis of gut contents in fungal-feeding oribatid mites by combining body-surface washing and PCR. Soil Biol Biochem 42(11):1952–1957. https://doi.org/10.1016/j.soilbio.2010.07.007
Renker C, Otto P, Schneider K, Zimdars B, Maraun M, Buscot F (2005) Oribatid mites as potential vectors for soil microfungi: study of mite-associated fungal species. Microb Ecol 50(4):518–528. https://doi.org/10.1007/s00248-005-5017-8
Rockett C (1980) Nematode predation by oribatid mites (Acari: Oribatida). Int J Acarol 6(3):219–224. https://doi.org/10.1080/01647958008683222
Rockett CL, Woodring JP (1966) Oribatid mites as predators of soil nematodes. Ann Entomol Soc Am 59(4):669–671. https://doi.org/10.1093/aesa/59.4.669
Rohde CJ (1959) Studies on the biologies of two mite species, predator and prey, including some effects of gamma radiation on selected developmental stages. Ecology 40:572–579. https://doi.org/10.2307/1929810
Santos PF, Whitford WG (1981) The effect of microarthropods on litter decomposition in a chihuahuan desert ecosystem. Ecology 62:654–663
Schneider K, Maraun M (2005) Feeding preferences among dark pigmented fungal taxa (“Dematiacea”) indicate limited trophic niche differentiation of oribatid mites (Oribatida, Acari). Pedobiologia 49(1):61–67. https://doi.org/10.1016/j.pedobi.2004.07.010
Schneider K, Renker C, Scheu S, Maraun M (2004a) Feeding biology of oribatid mites: a minireview. Phytophaga 14:247–256
Schneider K, Migge S, Norton RA et al (2004b) Trophic niche differentiation in soil microarthropods (Oribatida, Acari): Evidence from stable isotope ratios (15N/14N). Soil Biol Biochem 36:1769–1774. https://doi.org/10.1016/j.soilbio.2004.04.033
Secniczak A, Seniczak S, Slowikowska M, Paluszak Z (2017) The effect of different diet on life history parameters and growth of Oppia denticulata (Acari: Oribatida: Oppiidae). Syst Appl Acarol 22(5):749–758. https://doi.org/10.11158/saa.22.5.12
Seniczak S, Stefaniak O (1981) The effect of fungal diet on the development of Oppia nitens (Acari, Oribatei) and on the microflora of its alimentary tract. Pedobiologia 21:202–210
Siepel H, de Ruiter-Dijkman EM (1993) Feeding guilds of oribatid mites based on their carbohydrase activities. Soil Biol Biochem 25(11):1491–1497. https://doi.org/10.1016/0038-0717(93)90004-U
Siepel H, Maaskamp F (1994) Mites of different feeding guilds affect decomposition of organic matter. Soil Biol Biochem 26(10):1389–1394. https://doi.org/10.1016/0038-0717(94)90222-4
Smrž J (1991) Some aspects of the microanatomy of soil sarcoptiform mites (Acari: Oribatida and Acaridida). In: Dusbábek F, Bukva V (eds) Modern Acarology. Academia, Prague and SPB Academic Publishing, The Hague, pp 319–322
Smrž J, Norton R (2004) Food selection and internal processiong in Archegozetes longisetosus (Acari: Oribatida). Pedobiologia 48:111–120
Smrž J, Soukalová H, Čatská V, Hubert J (2016) Feeding patterns of Tyrophagus putrescentiae (sarcoptiformes: Acaridae) indicate that mycophagy is not a single and homogeneous category of nutritional biology. J Insect Sci. https://doi.org/10.1093/jisesa/iew070
Smrž J (2002) The excrement analysis - the useful tool for the biological and autoecological studies in soil zoology. In: Tajovsky K, Batik V, Pizl V (ed), Studies on soil fauna in central Europe, Ceske Budejovice, pp 185-189. https://doi.org/10.1016/j.pedrobi.2003.09.003
Stott DE, Martin JP (1989) Organic matter decomposition and retention in arid soils. Arid Soil Res Rehabil 3:115–148. https://doi.org/10.1080/15324988909381195
Subias L (2022) Listado sistemático, sinonímico y biogeográfico de los ácaros oribátidos (Acariformes: Oribatida) del mundo (excepto fósiles). Monografías electrónicas S. E. A. http://sea-entomologia.org/MeSEA_12_Listado_mundial_Acaros_Oribatidos_L_Subias.pdf
Todria N, Murvanidze M, Mumladze L (2021) Oribatid (Acari: Oribatida) diversity in natural and altered open arid ecosystems of South-Eastern Caucasus. Pedobiologia J Soil Ecol. https://doi.org/10.1016/j.pedobi.2021.150750
van Bronswijk J (1971) Food preference of pyroglyphid house-dust mites (Acari). Neth J Zool 22:335–340
Vašutová M, Mleczko P, López-García A, Maček I, Gergely Boros G, Ševčík J, Fujii S, Hackenberger D, Tuf I, Hornung E, Páll-Gergely B, Kjøller R (2019) Taxi drivers: the role of animals in transporting mycorrhizal fungi. Mycorrhiza 29:413–434
Velez P, Ojeda M, Espinosa-asuar L, Pérez T, Eguiarte LE, Souza V (2018) Experimental and molecular approximation to microbial niche: trophic interactions between Oribatid mites and microfungi in an oligotrophic freshwater system. Peer. https://doi.org/10.7717/peerj.5200
Visser S, Whittaker JB, Parkinson D (1981) Effects of collembolan grazing on nutrient release and respiration of a leaf litter inhabiting fungus. Soil Biol Biochem 13(3):215–218. https://doi.org/10.1016/0038-0717(81)90023-7
Walter DE, Proctor HC (2013) Mites: Ecology, Evolution & Behaviour: Life at a Microscale. Springer, Netherlands. https://doi.org/10.1007/978-94-007-7164-2
Wardle DA, Bardgett RD, Klironomos JN, Setälä H, Van Der Putten WH, Wall DH (2004) Ecological linkages between aboveground and belowground biota. Science 304(5677):1629–1633. https://doi.org/10.1126/science.1094875
Werner S, Persoh D, Rambold G (2016) New aspects of the biology of Mortierella alliacea. Mycol Prog 15:1293–1301. https://doi.org/10.1007/s11557-016-1243-3
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR Protocols: A Guide to Methods and Applications. Academic Press, Inc, New York, pp 315–322
Whitford WG, Duval BD (2019) Conceptual framework, paradigms, and models. In: Whitford WD, Duval BD (eds) Ecology of Desert Systems. Elsevier Ltd, London, pp 1–20. https://doi.org/10.1016/B978-0-12-815055-9.00001-1
Wickings K, Grandy AS (2011) The oribatid mite Scheloribates moestus (Acari: Oribatida) alters litter chemistry and nutrient cycling during decomposition. Soil Biol Biochem 43(2):351–358. https://doi.org/10.1016/j.soilbio.2010.10.023
Williams RH, Whipps JM, Cooke RC (1998) Role of soil mesofauna in dispersal of Coniothyrium minitans: transmission to sclerotia of Sclerotinia sclerotiorum. Soil Biol Biochem 30(14):1929–1935. https://doi.org/10.1016/S0038-0717(98)00063-7
Yoder JA, Krieger M, Oakley M, Trotter J et al (2019) Growth characteristics and pathogenic consequences of predominant entomopathogenic Yukon soil fungi Mortierella alpina and Penicillium expansum, and effectiveness of Met52®, against larvae of the winter Tick Dermacentor Albipictus. Studies Fungi 4(1):101–110. https://doi.org/10.5943/sif/4/1/13
Zak J, Whitford W (1988) Interactions among soil biota in desert ecosystems. Agric Ecosyst Environ 24(1–3):87–100. https://doi.org/10.1016/0167-8809(88)90058-8
Acknowledgements
Daniel I. Sánchez-Chávez acknowledges Posgrado en Ciencias Biológicas, UNAM and CONACyT fellowship (CVU No. 545397) for their support during the doctoral program. The present article is a requirement to obtain the Doctor in Science degree. All authors acknowledge Dra. Rosa Gabriela Castaño-Meneses for her valuable and constructive suggestions, advice and assistance on the planning and development of this research study, and Dr. Mark Maraun for the insightful comments offered to substantially improve the quality of the work. We also express our gratitude to the Zapotitlán (Salinas) authorities for all the help provided to carry on this study in the field, to Lidia I. Cabrera Martínez for technical support during molecular work at the Laboratorio de Sistemática Molecular del Departamento de Botánica (Instituto de Biología, UNAM), and to Laura Márquez and Nelly López for their assistance during sequencing procedures at the Laboratorio Nacional de Biodiversidad (Instituto de Biología, UNAM).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. All authors performed the material preparation and analysis. Data collection was performed by DISC and SRZ. The first draft of the manuscript was written by DISC and SRZ and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare they have no financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sánchez-Chávez, D.I., Rodríguez-Zaragoza, S., Velez, P. et al. Fungal feeding preferences and molecular gut content analysis of two abundant oribatid mite species (Acari: Oribatida) under the canopy of Prosopis laevigata (Fabaceae) in a semi-arid land. Exp Appl Acarol 89, 417–432 (2023). https://doi.org/10.1007/s10493-023-00790-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10493-023-00790-7