Abstract
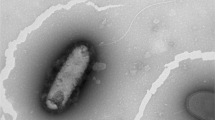
A novel potential plant growth promoting bacterium, designated OPS13-3T, was isolated from rhizosphere soil of citrus in Aotou Town of Guangzhou, Guangdong Province, PR China. It showed high ability to dissolve insoluble inorganic phosphate and organic phosphorus and to produce 3-indoleacetic acid (IAA) and siderophore. Cells of the novel strain were Gram-stain-negative, rod-shaped, aerobic and motile with polar flagellum. It shared the highest 16S rRNA gene similarity with Pseudomonas mucoides CCUG 74874T (98.7%) and P. bijieensis LMG 31948T (98.7%). Phylogenetic analyses based the 16S rRNA gene and genome sequences revealed that strain OPS13-3T belonged to the genus Pseudomonas, and was most closely related to P. mediterranea ICMP 14184T and P. corrugate ICMP 5819T. The average nucleotide identity (ANI) and DNA–DNA hybridization (dDDH) values between the novel strain and closely relatives with high 16S rRNA gene similarities were 80.8‒87.5% and 24.7‒34.6%, respectively, which were much below the threshold values for species delimitation. The major fatty acids included C16:0, C10:0 3-OH and summed feature 3 (C16:1ω7c and/or C16:1ω6c). It took ubiquinone 9 as the predominant respiratory quinone and the polar lipids contained phosphatidylglycerol (PG), diphosphatidylglycerol (DPG), phosphatidylethanolamine (PE), three unidentified phospholipids, an unidentified aminophospholipid and an unidentified lipid. Based on the phylogenetic, phenotypic and chemotaxonomic analyses and genome comparison, strain OPS13-3T should be considered as a novel species of the genus Pseudomonas, for which the name Pseudomonas citri sp. nov. is proposed (type strain OPS13-3T = GDMCC 1.3118T = JCM 35385T).


Similar content being viewed by others
Data availability
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene and draft genome sequences of strain OPS13-3T are OM327744 and JAODAB000000000, respectively.
References
Ames BN (1966) Assay of inorganic phosphate, total phosphate and phosphatases. Methods Enzymol 8:115–118. https://doi.org/10.1016/0076-6879(66)08014-5
Anwar N, Rozahon M, Zayadan B et al (2017) Pseudomonas tarimensis sp. nov., an endophytic bacteria isolated from Populus euphratica. Int J Syst Evol Microbiol 67:4372–4378. https://doi.org/10.1099/ijsem.0.002295
Aziz RK, Bartels D, Best AA et al (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Bric JM, Bostock RM, Silverstone SE (1991) Rapid in situ assay for indoleacetic acid production by bacteria immobilized on a nitrocellulose membrane. Appl Environ Microbiol 57:535–538. https://doi.org/10.1128/aem.57.2.535-538.1991
Buck JD (1982) Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993. https://doi.org/10.1128/aem.44.4.992-993.1982
Collins MD, Pirouz T, Goodfellow M et al (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230. https://doi.org/10.1099/00221287-100-2-221
Delorme S, Lemanceau P, Christen R et al (2002) Pseudomonas lini sp. nov., a novel species from bulk and rhizospheric soils. Int J Syst Evol Microbiol 52:513–523. https://doi.org/10.1099/00207713-52-2-513
Duman M, Mulet M, Altun S et al (2019) Pseudomonas piscium sp. nov., Pseudomonas pisciculturae sp. nov., Pseudomonas mucoides sp. nov. and Pseudomonas neuropathica sp. nov., isolated from rainbow trout. Int J Syst Evol Microbiol 71:004714. https://doi.org/10.1099/ijsem.0.004714
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Fierer N, Hamady M, Lauber CL et al (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999. https://doi.org/10.1073/pnas.0807920105
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Biol 20:406–416. https://doi.org/10.1093/sysbio/20.4.406
González AJ, Cleenwerck I, De Vos P et al (2013) Pseudomonas asturiensis sp. nov., isolated from soybean and weeds. Syst Appl Microbiol 36:320–324. https://doi.org/10.1016/j.syapm.2013.04.004
Goris J, Konstantinidis KT, Klappenbach JA et al (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Hiraishi A, Ueda Y, Ishihara J et al (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J General Appl Microbiol 42:457–469. https://doi.org/10.2323/jgam.42.457
Höfte M, Altier N (2010) Fluorescent pseudomonads as biocontrol agents for sustainable agricultural systems. Res Microbiol 161:464–471. https://doi.org/10.1016/j.resmic.2010.04.007
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Kujur RRA, Das SK (2022) Pseudomonas phenolilytica sp. nov., a novel phenol-degrading bacterium. Arch Microbiol 204:320. https://doi.org/10.1007/s00203-022-02912-y
Kumar A, Verma H, Singh VK et al (2017) Role of Pseudomonas sp. in sustainable agriculture and disease management. In: Meena VS, Mishra PK, Bisht JK, Pattanayak A (eds) Agriculturally important microbes for sustainable agriculture. Springer Singapore, Singapore. https://doi.org/10.1007/978-981-10-5343-6_7
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lee I, Chalita M, Ha SM et al (2017) ContEst16S: an algorithm that identifies contaminated prokaryotic genomes using 16S RNA gene sequences. Int J Syst Evol Microbiol 67:2053–2057. https://doi.org/10.1099/ijsem.0.001872
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nature Commun 10:1–10. https://doi.org/10.1038/s41467-019-10210-3
Meier-Kolthoff JP, Auch AF, Klenk HP et al (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinf 14:60. https://doi.org/10.1186/1471-2105-14-60
Migula W (1894) Über ein neues System der Bakterien. Arbeiten Aus Dem Bakteriologischen Institut Der Technischen Hochschule Zu Karlsruhe 1:235–238
Na SI, Kim YO, Yoon SH et al (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Nicklasson M, Martín-Rodríguez AJ, Thorell K et al (2022) Pseudomonas boanensis sp. nov., a bacterium isolated from river water used for household purposes in Boane District. Mozambique Int J Syst Evol Microbiol 72:005461. https://doi.org/10.1099/ijsem.0.005461
Nübel U, Garcia-Pichel F, Muyzer G (1997) PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl Environ Microbiol 63:3327–3332. https://doi.org/10.1128/aem.63.8.3327-3332.1997
Overbeek R, Olson R, Pusch GD et al (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42:D206–D214. https://doi.org/10.1093/nar/gkt1226
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP et al (2020) List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pérez-Miranda S, Cabirol N, George-Téllez R et al (2007) O-CAS, a fast and universal method for siderophore detection. J Microbiol Methods 70:127–131. https://doi.org/10.1016/j.mimet.2007.03.023
Qin J, Hu Y, Wu W et al (2020) Pseudomonas defluvii sp. nov., isolated from hospital sewage. Int J Syst Evol Microbiol 70:4199–4203. https://doi.org/10.1099/ijsem.0.004272
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Bacterial identification by gas chromatographic analysis of fatty acid methyl esters (GC-FAME). Technical note 101. http://www.midi-inc.com/pdf/MIS_Technote_101.pdf
Timsy Spanner T, Ulrich A et al (2021) Pseudomonas campi sp. nov., a nitrate-reducing bacterium isolated from grassland soil. Int J Syst Evol Microbiol 71:004799. https://doi.org/10.1099/ijsem.0.004799
Tindall BJ, Sikorski J, Smibert RA et al (2007) Phenotypic characterization and the principles of comparative systematics. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf GA, Schmidt TM, Synder LR (eds) Methods for general and molecular microbiology, 3rd edn. ASM Press, Washington, pp 330–393
Wahab MA, Sijam K, Ismail MR et al (2015) Phenotypic characterization and molecular identification of Malaysian Pseudomonas fuscovaginae isolated from rice plants. Asian J Plant Pathol 9:112–123. https://doi.org/10.3923/ajppaj.2015.112.123
Wang MQ, Sun L (2016) Pseudomonas oceani sp. nov., isolated from deep seawater. Int J Syst Evol Microbiol 66:4250–4255. https://doi.org/10.1099/ijsem.0.001343
Weisburg WG, Barns SM, Pelletier DA et al (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Widnyana IK, Javandira C (2016) Activities Pseudomonas spp. and Bacillus sp. to stimulate germination and seedling growth of tomato plants. Agric Agric Sci Proc 9:419–423. https://doi.org/10.1016/j.aaspro.2016.02.158
Yoon SH, Ha SM, Kwon S et al (2017a) Introducing Ezbiocloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yoon SH, Ha SM, Lim JM et al (2017b) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Funding
This work was jointly supported by the Natural Science Foundation of China (32170118), the Science and Technology Program of Guangdong Province (2021B1212050022), the GDAS’ Project of Science and Technology Development (2020GDASYL-20200301003) and the Guangdong Special Support Program (2021JC06N628).
Author information
Authors and Affiliations
Contributions
HG and GF drafted the manuscript; HG, JL and XD performed isolation, deposition and identifications; ZF and QY performed the genome and data analysis; GF, XL and HZ designed the experiments and improved the manuscript. All authors have approved the manuscript for submission.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical approval
This study does not describe any experimental work related to human.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gao, H., Feng, GD., Feng, Z. et al. Pseudomonas citri sp. nov., a potential novel plant growth promoting bacterium isolated from rhizosphere soil of citrus. Antonie van Leeuwenhoek 116, 281–289 (2023). https://doi.org/10.1007/s10482-022-01803-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-022-01803-y