Abstract
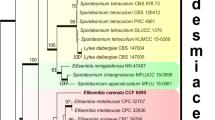
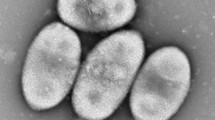
A Gram-stain-negative, aerobic, pear to oval shaped, rosette forming bacterium with crateriform structures well distributed on the cell surface designated as strain JC647T was isolated from a sponge specimen belonging to the genus Spongia. Strain JC647T reproduces through budding. Strain JC647T shared highest 16S rRNA gene sequence identity of 99.9% with “Crateriforma conspicua” Mal65T (not a valid species name). The genome size of strain JC647T is 6.9 Mb with a G + C content of 57.8 mol %. For the resolution of the phylogenetic congruence of the novel strain, the phylogeny was also constructed with the sequences of ninety-two housekeeping genes. Based on the phylogenetic analyses, low dDDH value (51.0%), low gANI (93.2%), low AAI (94.9%) results, chemotaxonomic characteristics and differential physiological properties, strain JC647T is recognized as a new species of the genus “Crateriforma”, for which we propose the name Crateriforma spongiae sp. nov. The type species is JC647T (= KCTC 72176T = NBRC 114068T).




Similar content being viewed by others
Abbreviations
- NCBI:
-
National Centre for Biotechnology Information
- gANI:
-
Genome Average Nucleotide Identity
- AAI:
-
Average Amino Acid Identity
- dDDH:
-
Digital DNA–DNA Hybridization
- POCP:
-
Percentage Of Conserved Proteins
- HPLC:
-
High-Pressure Liquid Chromatography
- KCTC:
-
Korean Collection for Type Cultures
- NBRC:
-
Biological Resource Centre, NITE
References
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genom 9:75
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Bondoso J, Harder J, Lage OM (2013) rpoB gene as a novel molecular marker to infer phylogeny in Planctomycetales. Antonie Van Leeuwenhoek 104:477–488
Bondoso J, Albuquerque L, Nobre MF, Lobo-da-Cunha A, da Costa MS, Lage OM (2015) Roseimaritima ulvae gen. nov., sp. nov. and Rubripirellula obstinate gen. nov., sp. nov. two novel planctomycetes isolated from the epiphytic community ofmacroalgae. Syst Appl Microbiol 37:8–15
Cai HY, Yan ZS, Wang AJ, Krumholz LR, Jiang HL (2013) Analysis of the attached microbial community on mucilaginous cyanobacterial aggregates in the eutrophic lake taihu reveals the importance of planctomycetes. Microb Ecol 66:73–83
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Darzi Y, Letunic I, Bork P, Yamada T (2018) iPath3. 0: interactive pathways explorer v3. Nucleic Acids Res 46:W510–W513
Dedysh SN, Kulichevskaya IS, Beletsky AV, Ivanova AA, Rijpstra WIC, Damsté JSS, Ravin NV (2020) Lacipirellula parvula gen. nov., sp. nov., representing a lineage of planctomycetes widespread in low-oxygen habitats, description of the family Lacipirellulaceae fam. nov. and proposal of the orders Pirellulales ord. nov., Gemmatalesord. nov. and Isosphaerales ord. nov. Syst Appl Microbiol 43:126050
DeLong EF, Franks DG, Alldredge AL (1993) Phylogenetic diversity of aggregate-attached vs. free-living marine bacterial assemblages. Limnol Oceanogr 38:924–934
Eisenreich W, Bacher A, Arigoni D, Rohdich F (2004) Biosynthesis of isoprenoids via the non-mevalonate pathway. Cell Mol Life Sci 61:1401–1426
Graca AP, Calisto R, Lage OM (2016) Planctomycetes as novel source of bioactive molecules. Front Microbiol 7:1241
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, Mering C, Bork P (2017) Fast genome-wide functional annotation through orthology assignment by egg NOG-mapper. Mol Biol Evol 34:115–2122
Imhoff JF (1984) Quinones of phototrophic purple bacteria. FEMS Microbiol Lett 25:85–89
Izumi H, Sagulenko E, Webb RI, Fuerst JA (2013) Isolation and diversity of planctomycetes from the sponge Niphates sp., seawater, and sediment of Moreton Bay. Australia. Antonie van Leeuwenhoek 104:533–546
Jeske O, Schüler M, Schumann P, Schneider A, Boedeker C, Jogler M, Bollschweiler D, Rohde M, Mayer C, Engelhardt H, Spring S, Jogler C (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:7116
Kaboré OD, Godreuil S, Drancourt M (2020) Planctomycetes as host-associated bacteria: a perspective that holds promise for their future isolations, by mimicking their native environmental niches in clinical microbiology laboratories. Front Cell Infect Microbiol 10:729
Kallscheuer N, Jeske O, Sandargo B, Boedeker C, Wiegand S, Bartling P, Jogler M, Rohde M, Petersen J, Medema MH, Surup F, Jogler C (2020) The planctomycete Stieleriamaiorica Mal15T employs stieleriacines to alter the species composition in marine biofilms. Commun Biol 3:303
Kates M (1972) Isolation, analysis and identification of lipids. Tech Lipidol, pp 268–618
Köhler T, Stingl U, Meuser K, Brune A (2008) Novel lineages of Planctomycetes densely colonize the alkaline gut of soil-feeding termites (Cubitermes spp.). Environ Microbiol 10:1260–1270
Kohn T, Wiegand S, Boedeker C, Rast P, Heuer A, Jetten MSM, Schüler M, Becker S, Rohde C, Müller RW, Brümmer F (2020) Planctopirus ephydatiae, a novel Planctomycete isolated from a freshwater sponge. Syst Appl Microbiol 43:126022
Kumar D, Gaurav K, Jagadeshwari U, Sasikala Ch, Ramana Ch (2020a) Roseimaritima sediminicola sp. nov., a new member of Planctomycetaceae isolated from Chilika lagoon. Int J Syst Evol Microbiol 70:2616–2623
Kumar D, Gaurav K, Sreya PK, Shabbir A, Jagadeshwari U, Sasikala Ch, Ramana C (2020b) Gimesia chilikensis sp. nov., a haloalkali-tolerant planctomycete isolated from Chilika lagoon and emended description of the genus Gimesia. Int J Syst Evol Microbiol 70:3647–3655
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Luo C, Rodriguez-r LM, Konstantinidis KT (2014) MyTaxa: an advanced Taxonomicclassifier for genomic and metagenomic sequences. Nucleic Acids Res 42:e73
Meier-Kolthoff JP, Klenk HP, Göker M (2014) Taxonomic use of DNA G + C content and DNA–DNA hybridization in the genomic age. Int J Syst Evol Microbiol 64:352–356
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285
Najafi A, Moradinasab M, Nabipour I (2018) First record of microbiomes of sponges collected from the Persian Gulf, using tag pyrosequencing. Front Microbiol 9:1500
Oren A, Duker S, Ritter S (1996) The polar lipid composition of Walsby’s square bacterium. FEMS Microbiol Lett 138:135–140
Peeters SH, Wiegand S, Kallscheuer N, Jogler M, Heuer A, Jetten MS, Rast P, Boedeker C, Rohde M, Jogler C (2020) Three marine strains constitute the novel genus and species Crateriforma conspicua in the phylum Planctomycetes. Antonie Van Leeuwenhoek 10:1–13
Pruesse E, Peplies J, Glockner FO (2012) SINA: accurate high throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol Res 196:2210–2215
Rodriguez-R LM, Konstantinidis KT (2014) Bypassing cultivation to identify bacterial species. Microbe 9:111–118
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids
Stackebrandt E, Liesack W, Goebel BM (1993) Bacterial diversity in a soil sample from a subtropical Australian environment as determined by 16S rDNA analysis. FASEB J 7:232–236
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Vacelet J, Donadey C (1977) Electron microscope study of the association between some sponges and bacteria. J Exp Mar Biol Ecol 30:301–314
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiaeand sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Wattam AR, Davis JJ, Assaf R, Boisvert S, Brettin T, Bun C, Conrad N, Dietrich EM, Disz T, Gabbard JL, Gerdes S, Henry CS, Kenyon RW, Machi D, Mao C, Nordberg EK, Olsen GJ, Murphy-Olson DE, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Vonstein V, Warren A, Xia F, Yoo H, Steven RL (2017) Improvements to PATRIC, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res 45:D535–D542
Wick RR, Judd LM, Gorrie CL, Holt KE (2017) Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol 13:e1005595
Wiegand S, Jogler M, Jogler C (2018) On the maverick Planctomycetes. FEMS Microbiol Rev 42:739–760
Wiegand S, Jogler M, Boedeker C, Pinto D, Vollmers J, Rivas-Marín E, Kohn T, Peeters SH, Heuer A, Rast P, Oberbeckmann S, Bunk B, Jeske O, Meyerdierks A, Storesund JE, Kallscheuer N, Lucker S, Lage OM, Pohl T, Merkel BJ, Hornburger P, Muller R, Brummer F, Labrenz M, Sporemann AM, Op den Camp H, Overmann J, Amann R, Jetten MSM, Mascher T, Medema MH, Devos DP, Kaster A-K, Øvreås L, Rohde M, Galperin MY, Jogler C (2020) Cultivation and functional characterization of 79 Planctomycetes uncovers their unique biology. Nat Microbiol 5:126–140
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613
Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, Busk PK, Xu Y, Yin Y (2018) dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 46:W95–W101
Zhao L, Chang WC, Xiao Y, Liu HW, Liu P (2013) Methylerythritol phosphate pathway of isoprenoid biosynthesis. Annu Rev Biochem 82:497–530
Acknowledgements
Gaurav and Ramana thank DBT, New Delhi for the award of SRF and TATA innovation fellowship, respectively. Sasikala, Dhanesh Kumar and Shabbir thank UGC for Mid-career award, RA and SRF awards respectively. Sreya and Jagadeeshwari thank CSIR and TEQIP, respectively for JRF. We also thanks Mr. Bernhard Schink for providing species name and etymology for strain JC647T.
Funding
This work received specific grant from Department of Biotechnology, Government of India, with sanction No. BT/HRD/35/01/02/2015. Financial support received from Council for Scientific and Industrial Research (CSIR), New Delhi is acknowledged. Infrastructure facilities funded by DST-FIST, UGC-SAP (DRS), TEQIP and AICTE are acknowledged.
Author information
Authors and Affiliations
Contributions
GK, Ramana and Sasikala performed sample collection, GK performed media optimisation and polar lipid analysis and wrote manuscript, GK and DK isolated the strain and performed the initial cultivation and strain deposition, GK and SPK performed strain characterisation, DK and GK performed the electron microscopic analysis, JU performed the genomic and phylogenetic analysis, SA performed and analysed the data for polyamines, Ramana and Sasikala supervised the study and contributed to text preparation and revised the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene sequences of the strains JC647T is LR132071. This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBankunder the accession JAAXMS000000000. The version described in this paper is version JAAXMS010000000.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, G., Kumar, D., Jagadeeshwari, U. et al. Crateriforma spongiae sp. nov., isolated from a marine sponge and emended description of the genus “Crateriforma”. Antonie van Leeuwenhoek 114, 341–353 (2021). https://doi.org/10.1007/s10482-020-01515-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-020-01515-1