Abstract
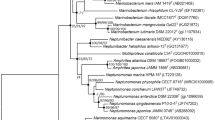
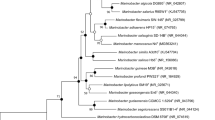
A Gram-stain negative, aerobic, motile, short rod-shaped bacterium, designated as B2.3T, was isolated from coal bed water collected from Jincheng, Shanxi Province, China. The strain was able to grow at 10–40 °C (optimum 28–30 °C), pH 4.0–10.0 (optimum 7.0), and in the presence of 0–5.0% NaCl (optimum 3.0%, w/v). Phylogenetic analysis based on the 16S rRNA and concatenated housekeeping gene recA, atpD and glnA sequences showed strain B2.3T belongs to the genus Mesorhizobium, with Mesorhizobium oceanicum B7T as the closely related type strain. Strain B2.3T exhibited ANI value of 77.5% and GGDC value of 21.5% to M. oceanicum B7T. The major fatty acids were identified as summed feature 8 (C18:1ω7c and/or C18:1ω6c) and 11-methyl C18:1ω7c. The major polar lipids were found to consist of diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, phosphatidylcholine and an unidentified aminophospholipid. The predominant ubiquinone was identified as Quinone 10. Phenotypic and biochemical analysis results indicated that strain B2.3T can be distinguished from closely related type strains. On the basis of phenotypic, genotypic and chemotaxonomic characteristics, strain B2.3T is concluded to represent a novel species in the genus Mesorhizobium, for which the name Mesorhizobium carbonis sp. nov. is proposed. The type strain is B2.3T (=CGMCC 1.15730T = KCTC 52461T).


Similar content being viewed by others
Abbreviations
- ANI:
-
Average nucleotide identity
- DDH:
-
DNA–DNA hybridization
References
Chen WM, Zhu WF, Bontemps C, Young JP, Wei GH (2011) Mesorhizobium camelthorni sp. nov., isolated from Alhagi sparsifolia. Int J Syst Evol Microbiol 61:574–579
Choma A, Komaniecka I (2002) Analysis of phospholipids and ornithine-containing lipids from Mesorhizobium spp. Syst Appl Microbiol 25:326–331
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Collins MD, Dorothy J (1980) Lipids in the classification and identification of coryneform bacteria containing peptidoglycans based on 2, 4-diaminobutyric acid. J Appl Microb 48:459–470
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Collins MD, Goodfellow M, Minnikin DE, Alderson G (1985) Menaquinone composition of mycolic acid-containing actinomycetes and some sporoactinomycetes. J Appl Bacteriol 58:77–86
De Meyer SE, Tan HW, Heenan PB, Andrews M, Willems A (2015) Mesorhizobium waimense sp. nov. isolated from Sophora longicarinata root nodules and Mesorhizobium cantuariense sp. nov. isolated from Sophora microphylla root nodules. Int J Syst Evol Microbiol 65:3419–3426
De Meyer SE, Tan HW, Andrews M, Heenan PB, Willems A (2016) Mesorhizobium calcicola sp. nov., Mesorhizobium waitakense sp. nov., Mesorhizobium sophorae sp. nov., Mesorhizobium newzealandense sp. nov. and Mesorhizobium kowhaii sp. nov. isolated from Sophora root nodules. Int J Syst Evol Microbiol 66:786–795
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23:673–679
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Biol 20:406–416
Fu GY, Yu XY, Zhang CY, Zhao Z, Wu D, Su Y, Wang RJ, Han SB, Wu M, Sun C (2017) Mesorhizobium oceanicum sp. nov., isolated from deep seawater. Int J Syst Evol Microbiol 67:2739–2745
Gaunt M, Turner S, Rigottier-Gois L, Lloyd-Macgilp S, Young J (2001) Phylogenies of atpD and recA support the small subunit rRNA-based classification of rhizobia. Int J Syst Evol Microbiol 51:2037–2048
Ghosh W, Roy P (2006) Mesorhizobium thiogangeticum sp. nov., a novel sulfur-oxidizing chemolithoautotroph from rhizosphere soil of an Indian tropical leguminous plant. Int J Syst Evol Microbiol 56:91–97
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Jarvis B, Van Berkum P, Chen W, Nour S, Fernandez M, Cleyet-Marel J, Gillis M (1997) Transfer of Rhizobium loti, Rhizobium huakuii, Rhizobium ciceri, Rhizobium mediterraneum, and Rhizobium tianshanense to Mesorhizobium gen. nov. Int J Syst Evol Microbiol 47:895–898
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Küster E (1959) Outline of a comparative study of criteria used in characterization of the actinomycetes. Int J Syst Evol Microbiol 9:97–104
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lorite MJ, Flores-Felix JD, Peix A, Sanjuan J, Velazquez E (2016) Mesorhizobium olivaresii sp. nov. isolated from Lotus corniculatus nodules. Syst Appl Microbiol 39:557–561
Marcos-Garcia M, Menendez E, Ramirez-Bahena MH, Mateos PF, Peix A, Velazquez E, Rivas R (2017) Mesorhizobium helmanticense sp. nov., isolated from Lotus corniculatus nodules. Int J Syst Evol Microbiol 67:2301–2305
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3:208-IN201
Martínez-Hidalgo P, Ramírez-Bahena MH, Flores-Félix JD, Rivas R, Igual JM, Mateos PF, Martínez-Molina E, León-Barrios M, Peix A, Velázquez E (2015) Revision of the taxonomic status of type strains of Mesorhizobium loti and reclassification of strain USDA 3471T as the type strain of Mesorhizobium erdmanii sp. nov. and ATCC 33669T as the type strain of Mesorhizobium jarvisii sp. nov. Int J Syst Evol Microbiol 65:1703–1708
Martínez-Hidalgo P, Florés-Felix JD, Igual JM, Sanjuán J, León-Barrios M, Peix A, Velázquez E (2016) Reclassification of strains MAFF 303099T and R7A into Mesorhizobiumjaponicum sp. nov. Int J Syst Evol Microbiol 66:4936–4941
Meier-Kolthoff JP, Auch AF, Klenk H-P, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Minnikin D, Collins M, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Microb 47:87–95
Mohamad R, Willems A, Le Quere A, Maynaud G, Pervent M, Bonabaud M, Dubois E, Cleyet-Marel JC, Brunel B (2017) Mesorhizobium delmotii and Mesorhizobium prunaredense are two new species containing rhizobial strains within the symbiovar anthyllidis. Syst Appl Microbiol 40:135–143
Richter M, Rossello-Mora R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131
Rossello-Mora R, Trujillo ME, Sutcliffe IC (2017) Introducing a digital protologue: a timely move towards a database-driven systematics of archaea and bacteria. Antonie Van Leeuwenhoek 110:455–456
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101 MIDI Inc
Smibert R, Krieg N (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, pp 607–654
Tighe S, De Lajudie P, Dipietro K, Lindström K, Nick G, Jarvis B (2000) Analysis of cellular fatty acids and phenotypic relationships of Agrobacterium, Bradyrhizobium, Mesorhizobium, Rhizobium and Sinorhizobium species using the Sherlock Microbial Identification System. Int J Syst Evol Microbiol 50:787–801
Vinuesa P, Silva C, Werner D, Martinez-Romero E (2005) Population genetics and phylogenetic inference in bacterial molecular systematics: the roles of migration and recombination in Bradyrhizobium species cohesion and delineation. Mol Phylogenet Evol 34:29–54
Xu CF, Zhang L, Huang JW, Chen K, Li SP, Jiang JD (2017) Aquamicrobium soli sp. nov., a bacterium isolated from a chlorobenzoate-contaminated soil. Antonie Van Leeuwenhoek 110:305–312
Xu L, Zhang Y, Mohamad OA, Jiang C, Friman VP (2018) Mesorhizobium zhangyense sp. nov., isolated from wild Thermopsis lanceolate in northwestern China. Arch Microbiol 200:603–610
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Yuan CG, Jiang Z, Xiao M, Zhou EM, Kim CJ, Hozzein WN, Park DJ, Zhi XY, Li WJ (2016) Mesorhizobium sediminum sp. nov., isolated from deep-sea sediment. Int J Syst Evol Microbiol 66:4797–4802
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18:821–829
Zhang JJ, Liu TY, Chen WF, Wang ET, Sui XH, Zhang XX, Li Y, Li Y, Chen WX (2012) Mesorhizobium muleiense sp. nov., nodulating with Cicer arietinum L. Int J Syst Evol Microbiol 62:2737–2742
Zhang J, Yang X, Guo C, de Lajudie P, Singh RP, Wang E, Chen W (2017) Mesorhizobium muleiense and Mesorhizobium gsp. nov. are symbionts of Cicer arietinum L. in alkaline soils of Gansu, Northwest China. Plant Soil 410:103–112
Zhang J, Guo C, Chen W, de Lajudie P, Zhang Z, Shang Y, Wang ET (2018) Mesorhizobium wenxiniae sp. nov., isolated from chickpea (Cicer arietinum L.) in China. Int J Syst Evol Microbiol 68:1930–1936
Zheng WT, Li Y Jr, Wang R, Sui XH, Zhang XX, Zhang JJ, Wang ET, Chen WX (2013) Mesorhizobium qingshengii sp. nov., isolated from effective nodules of Astragalus sinicus. Int J Syst Evol Microbiol 63:2002–2007
Zhou S, Li Q, Jiang H, Lindstrom K, Zhang X (2013) Mesorhizobium sangaii sp. nov., isolated from the root nodules of Astragalus luteolus and Astragalus ernestii. Int J Syst Evol Microbiol 63:2794–2799
Acknowledgements
This work was financially supported by the National Natural Science Foundation (30901108 and 21776310), the Shandong Provincial Natural Science Foundation (ZR2009EQ007 and ZR2017MB019) and the Fundamental Research Funds for the Central Universities (15CX02014A and 16CX02044A).
Author information
Authors and Affiliations
Contributions
JL designed the study and wrote the manuscript. WX, ZZX and FQX performed the experiments. JL and LJX analyzed the data. JJZ contributed to the polar lipid analysis. JBQ and JGL revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, J., Xin, W., Xu, ZZ. et al. Mesorhizobium carbonis sp. nov., isolated from coal bed water. Antonie van Leeuwenhoek 112, 1221–1229 (2019). https://doi.org/10.1007/s10482-019-01254-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01254-y