Abstract
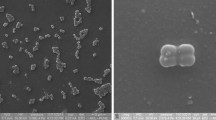
Strain 16F3HT, a Gram-negative, aerobic, non-motile, and oval-shaped bacterium, was isolated from river water collected from the Han River in South Korea. Growth of strain 16F3HT was observed at 10–42 °C (optimum at 25–30 °C), but no growth occurred at 4 °C. The strain is able to grow at pH 4–10 (optimum at pH 7–8) and tolerates up to 4% NaCl (w/v), with optimum growth at 0.5% NaCl. The isolate was found to be resistant to UV irradiation. Based on 16S rRNA gene sequence analysis, it is closely related to ‘Deinococcus seoulensis’ 16F1E (98.8%), Deinococcus aquaticus PB314T (98.1%) and Deinococcus caeni Ho-08T (98.0%). The level of DNA–DNA homology between the novel strain and the three related strains was 57.4, 41.2, and 35.8%, respectively. Chemotaxonomic data revealed that strain 16F3HT possesses MK-8 as the predominant respiratory quinone, an unidentified phosphoglycolipid as the major polar lipid, and C15:1 ω6c and C16:1 ω7c as the major fatty acids. The DNA G + C content was determined to be 65.7 mol%. Based on polyphasic evidence, strain 16F3HT (=KCTC 33794T = JCM 31406T) should be classified as the type strain of a novel Deinococcus species, for which the name Deinococcus knuensis sp. nov. is proposed.


Similar content being viewed by others
References
Ahmed I, Abbas S, Kudo T, Iqbal M, Fujiwara T, Ohkuma M (2014) Deinococcus citri sp. nov., isolated from citrus leaf canker lesions. Int J Syst Evol Microbiol 64:4134–4140
Asker D, Awad TS, McLandsborough L, Beppu T, Ueda K (2011) Deinococcus depolymerans sp. nov., a gamma and UV-radiation-resistant bacterium, isolated from a naturally radioactive site. Int J Syst Evol Microbiol 61:1448–1453
Brooks BW, Murray RGE (1981) Nomenclature for ‘‘Micrococcus radiodurans’’ and other radiation-resistant cocci: Deinococcaceae fam. nov. and Deinococcus gen. nov., including five species. Int J Syst Evol Microbiol 31:353–360
Buck JD (1982) Non-staining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Callegan RP, Nobre MF, McTernan PM, Battista JR, Navarro-Gonzalez R, McKay CP, da Costa MS, Rainey FA (2008) Description of four novel psychrophilic, ionizing radiation-sensitive Deinococcus species from alpine environments. Int J Syst Evol Microbiol 58:1252–1258
Cappuccino JG, Sherman N (2002) Microbiology: a laboratory manual, 6th edn. Pearson Education, Inc., Benjamin Cummings
Casanellas M, Fernández-Sánchez J (2008) Geometry of the Kimura 3-parameter model. Adv Appl Math 41:265–292
Cha S, Srinivasan S, Seo T, Kim MK (2014a) Deinococcus radiotolerans sp. nov., a gamma-radiation resistant bacterium isolated from gamma ray-irradiated soil. Antonie Van Leeuwenhoek 105:229–235
Cha S, Srinivasan S, Seo T, Kim MK (2014b) Deinococcus soli sp. nov., a gamma-radiation-resistant bacterium isolated from rice field soil. Curr Microbiol 68:777–783
Counsell TJ, Murray RGE (1986) Polar lipid profiles of the genus Deinococcus. Int J Syst Bacteriol 36:202–206
Ezaki T, Hashimoto Y, Yabuuchi E (1989) Fluorometric deoxyribonucleic acid-deoxyribonucleic acid hybridization in microdilution wells as an alternative to membrane-filter hybridization in which radioisotopes are used to determine genetic relatedness among bacterial strains. Int J Syst Bacteriol 39:224–229
Felsenstein J (1985) Confidence limit on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specified tree topology. Syst Zool 20(4):406–416
Groot A, Chapon V, Servant P, Christen R, Saux MFL, Sommer S, Heulin T (2005) Deinococcus deserti sp. nov., a gamma-radiation tolerant bacterium isolated from the Sahara Desert. Int J Syst Evol Microbiol 55:2441–2446
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Im WT, Jung HM, Ten LN, Kim MK, Bora N, Goodfellow M, Lim S, Jung J, Lee ST (2008) Deinococcus aquaticus sp. nov., isolated from fresh water, and Deinococcus caeni sp. nov., isolated from activated sludge. Int J Syst Evol Microbiol 58:2348–2353
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1981) Estimation of evolutionary distances between homologous nucleotide sequences. Proc Natl Acad Sci 78:454–458
Komagata K, Suzuki K (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–207
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lee JJ, Lee YH, Park SJ, Lim S, Jeong SW, Lee SY, Cho YJ, Kim MK, Jung HY (2016) Deinococcus seoulensis sp. nov., a bacterium isolated from sediment at Han River in Seoul, Republic of Korea. J Microbiol 54:537–542
Mattimore V, Battista JR (1996) Radioresistance of Deinococcus radiodurans: functions necessary to survive ionizing radiation are also necessary to survive prolonged desiccation. J Bacteriol 178:633–637
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G + C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Minnikin DE, Patel PV, Alshamaony L, Goodfellow M (1977) Polar lipid composition in the classification of Nocardia and related bacteria. Int J Syst Bacteriol 27:104–117
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Rainey FA, Ray K, Ferreira M, Gatz BZ, Nobre MF, Bagaley D, Rash BA, Park MJ, Earl AM, Shank NC, Small AM, Henk MC, Battista JR, Ka¨mpfer P, da Costa MS (2005) Extensive diversity of ionizing-radiation-resistant bacteria recovered from Sonoran Desert soil and description of nine new species of the genus Deinococcus obtained from a single soil sample. Appl Environ Microbiol 71:5225–5235
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Bio Evol 4:406–425
Sasser M (1990) Identification of Bacteria by Gas Chromatography of Cellular Fatty Acids. MIDI Technical Note 101. Newark, DE: MIDI Inc
Tamaoka J, Komagata K (1984) Determination of DNA base composition by reversed phase high-performance liquid chromatography. FEMS Microbiol Lett 25:125–128
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Weisburg WG, Barns SM, Pellerier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Wilson K (1997) Preparation of genomic DNA from bacteria. In: Frederick M et al (eds) Current protocols in molecular biology. Wiley, Australia, pp 2.4.1–2.4.5
Acknowledgements
This work was supported by the Human Resource Training Program for Regional Innovation and Creativity through the Ministry of Education and National Research Foundation (NRF) of Korea (NRF-2014H1C1A1066929), and by the Brain Pool Program of 2016 funded by the Ministry of Science, Republic of Korea (Grant 162S-4-3-1727).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, JJ., Lee, YH., Park, SJ. et al. Deinococcus knuensis sp. nov., a bacterium isolated from river water. Antonie van Leeuwenhoek 110, 407–414 (2017). https://doi.org/10.1007/s10482-016-0813-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-016-0813-3