Abstract
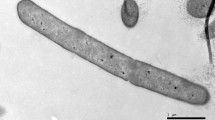
A Gram-stain positive, non-motile, non-sporogenous, aerobic, rod-shaped and halophilic bacterium, designated LAM0015T, was isolated from a saline sediment sample collected from Yantai City in China. The isolate was found to be able to grow at NaCl concentrations of 5–25 % (w/v) (optimum: 7–12 %), 15–45 °C (optimum: 35 °C) and pH 5.0–9.0 (optimum: 7.0). The major fatty acids were determined to be anteiso-C15:0 and anteiso-C17:0. The predominant respiratory quinone was identified as MK-7. The cell wall peptidoglycan was determined to contain meso-diaminopimelic acid. The polar lipids were found to be diphosphatidyglycerol, phosphatidylglycerol, five phospholipids and one glycolipid. The DNA G+C content was 43.1 mol% as determined by the T m method. Analysis of the 16S rRNA gene sequence indicated that the isolate belongs within the genus Lentibacillus and is closely related to Lentibacillus persicus DSM 22530T, Lentibacillus salicampi JCM 11462T and Lentibacillus jeotgali JCM 15795T with 97.3, 96.7 and 96.4 % sequence similarity, respectively. The DNA–DNA hybridization value between LAM0015T and L. persicus DSM 22530T was 51.2 ± 1.4 %. Based on its phenotypic, phylogenetic and chemotaxonomic characteristics, strain LAM0015T is concluded to represent a novel species of the genus Lentibacillus, for which the name Lentibacillus amyloliquefaciens sp. nov. is proposed. The type strain is LAM0015T (=ACCC 06401T = JCM 19838T).

Similar content being viewed by others
References
Buck JD (1982) Nonstaining (KOH) method for determination of gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Cowan ST, Steel KJ (1965) Manual for the identification of medical bacteria. Cambridge University Press, London
De LJ, Cattoir H, Reynaert A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Harrigan WF, McCance ME (1976) Book Laboratory Methods in Food and Dairy Microbiology. Academic Press, London
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol 4:184–192
Jeon CO, Lim JM, Lee JC, Lee GS, Lee JM, Xu LH, Jiang CL, Kim CJ (2005) Lentibacillus salarius sp. nov., isolated from saline sediment in China, and emended description of the genus Lentibacillus. Int J Syst Evol Microbiol 55:1339–1343
Jung MJ, Roh SW, Kim MS, Bae JW (2010) Lentibacillus jeotgali sp. nov., a halophilic bacterium isolated from traditional Korean fermented seafood. Int J Syst Evol Microbiol 60:1017–1022
Jung WY, Lee SH, Jin HM, Jeon CO (2015) Lentibacillus garicola sp. nov., isolated from myeolchi-aekjeot, a Korean fermented anchovy sauce. Antonie Van Leeuwenhoek 107:1569–1576
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Komagata K, Suzuki K-I (1987) Lipid and Cell-Wall Analysis in Bacterial Systematics. Methods Microbiol 19:161–207
Lányi B (1987) Classical and rapid identification methods for medically important bacteria. Methods Microbiol 19:1–67
Lee JC, Li WJ, Xu LH, Jiang CL, Kim CJ (2008) Lentibacillus salis sp. nov., a moderately halophilic bacterium isolated from a salt lake. Int J Syst Evol Microbiol 58:1838–1843
Logan NA, Berge O, Bishop AH, Busse HJ, De Vos P, Fritze D, Heyndrickx M, Kampfer P, Rabinovitch L, Salkinoja-Salonen MS, Seldin L, Ventosa A (2009) Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int J Syst Evol Microbiol 59:2114–2121
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J Mol Biol 5:109–118
Minnikin DE, Odonnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Ruan Z, Wang Y, Song J, Jiang S, Wang H, Li Y, Zhao B, Jiang R, Zhao B (2014) Kurthia huakuii sp. nov., isolated from biogas slurry, and emended description of the genus Kurthia. Int J Syst Evol Microbiol 64:518–521
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sakamoto M et al (2002) Reclassification of Bacteroides forsythus (Tanner et al. 1986) as Tannerella forsythensis corrig., gen. nov., comb. nov. Int J Syst Evol Microbiol 52:841–849
Sánchez-Porro C, Amoozegar MA, Fernandez AB, Babavalian H, Ramezani M, Ventosa A (2010) Lentibacillus persicus sp. nov., a moderately halophilic species isolated from a saline lake. Int J Syst Evol Microbiol 60:1407–1412
Schleifer KH (1985) Analysis of the Chemical Composition and Primary Structure of Murein. Methods Microbiol 18:123–156
Sehgal SN, Gibbons NE (1960) Effect of metal ions on the growth of Halobacterium cutirubrum. Can J Microbial 6:165–169
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for General and Molecular Bacteriology, pp 647–654. American Society for Microbiology, Washington, DC
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Xu XW, Huo YY, Wang CS, Oren A, Cui HL, Vedler E, Wu M (2011) Pelagibacterium halotolerans gen. nov., sp. nov. and Pelagibacterium luteolum sp. nov., novel members of the family Hyphomicrobiaceae. Int J Syst Evol Microbiol 61:1817–1822
Yoon JH, Kang KH, Park YH (2002) Lentibacillus salicampi gen. nov., sp. nov., a moderately halophilic bacterium isolated from a salt field in Korea. Int J Syst Evol Microbiol 52:2043–2048
Acknowledgments
We are grateful to Professor Min Wu from Zhejiang University for his technical assistance. This work was supported by Open Funds of Xinjiang Production & Construction Corps Key Laboratory of Protection and Utilization of Biological Resources in Tarim Basin (Tarim University, BRZD1403), National Nonprofit Institute Research Grant of CAAS (No. 2014-30), National Key Technology R&D Program of China (No. 2013BAD05B04F02).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Jing-Li Wang and Ke-Dong Ma have contributed equally to the work and share first authorship.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, JL., Ma, KD., Wang, YW. et al. Lentibacillus amyloliquefaciens sp. nov., a halophilic bacterium isolated from saline sediment sample. Antonie van Leeuwenhoek 109, 171–178 (2016). https://doi.org/10.1007/s10482-015-0618-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0618-9