Abstract
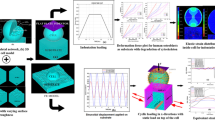
Cell’s shape is dependent on the cytoskeleton mechanical properties. Hybrid models were developed that combine the discrete structure for the cytoskeleton and continuum parts for other cell organelles. Tensegrity-based structures that consist of tensile and compression elements are useful models to understand the cytoskeleton mechanical behavior. In this study, we are looking to examine the reaction of the cell to a variety of substrate stiffnesses and explain the relationship between cell behavior and substrate mechanical properties. However, which tensegrity structure is appropriate for modeling a living cell? Is the structure’s complexity play a major role? We used two spherical tensegrities with different complexities to assess the impact of the structure on the cell’s mechanical response versus substrate’s stiffness. Six- and twelve-strut tensegrities together with membrane, cytoplasm, nucleoskeleton, and nucleus envelope were assembled in Abaqus package to create a hybrid cell model. A compressive load was applied to the cell model and the reaction forces versus deflection curves were analyzed for number of substrate stiffness values. By analyzing the difference due to two different tensegrities it became clear that the lower density structure is a better choice for modeling stiffer cells. It was also found that the six-strut tensegrity is sensitive to higher range of substrate stiffness.








Similar content being viewed by others
Data Availability
There are no datasets for the current study.
References
Coughlin, M. A tensegrity model of the cytoskeleton in spread and round cells. J. Biomech. Eng.. 120(6):770–777, 1998.
Wang, N., et al. Mechanical behavior in living cells consistent with the tensegrity model. Proc. Natl. Acad. Sci.. 98(14):7765–7770, 2001.
Chicurel, M. E., C. S. Chen, and D. E. Ingber. Cellular control lies in the balance of forces. Curr. Opini. Cell Biol.. 10(2):232–239, 1998.
Ingber, D. E., and I. Tensegrity. Cell structure and hierarchical systems biology. J. Cell Sci.. 116(7):1157–1173, 2003.
Ingber, D. E., and I. I. Tensegrity. How structural networks influence cellular information processing networks. J. Cell Sci.. 116(8):1397–1408, 2003.
Ingber, D. E., et al. Cellular tensegrity: exploring how mechanical changes in the cytoskeleton regulate cell growth, migration, and tissue pattern during morphogenesis. In: International review of cytology, Elsevier, 1994, pp. 173–224.
Satcher, R., C. F. Dewey, and J. H. Hartwig. Mechanical remodeling of the endothelial surface and actin cytoskeleton induced by fluid flow. Microcirculation. 4(4):439–453, 1997.
Stamenovic, D., et al., Cell prestress. II. Contribution of microtubules. Am. J. Physiol. Cell Physiol. 282(3): C617-C624, 2002.
Khismatullin, D. B. The cytoskeleton and deformability of white blood cells. Curr. Top. Membr.. 64:47–111, 2009.
Ingber, D. E. Tensegrity-based mechanosensing from macro to micro. Prog. Biophys. Mol.Biol.. 97(2–3):163–179, 2008.
Wendling, S., P. Cañadas, C. Oddou, and A. Meunier. Interrelations between elastic energy and strain in a tensegrity model: contribution to the analysis of the mechanical response in living cells. Comput. Methods Biomech. Biomed. Eng.. 5(1):1–6, 2002.
Wendling-Mansuy, S., P. Cañadas, and P. Chabrand. Toward a generalized tensegrity model describing the mechanical behaviour of the cytoskeleton structure. Comput. Methods Biomech. Biomed. Eng.. 6(1):45–52, 2003.
Prendergast, P.J., Computational modelling of cell and tissue mechanoresponsiveness. Gravit. Space Res.. 20(2):43, 2007.
Kardas, D., U. Nackenhorst, and D. Balzani. Computational model for the cell-mechanical response of the osteocyte cytoskeleton based on self-stabilizing tensegrity structures. Biomech. Model. Mechanobiol.. 12(1):167–183, 2013.
Ingber, D. E. Cellular tensegrity: defining new rules of biological design that govern the cytoskeleton. J. Cell Sci.. 104(3):613–627, 1993.
Ingber, D. E. Tensegrity: the architectural basis of cellular mechanotransduction. Annu. Rev. Physiol.. 59(1):575–599, 1997.
Bursa, J. and V. Fuis. Finite element simulation of mechanical tests of individual cells. in World Congress on Medical Physics and Biomedical Engineering, September 7-12, 2009, Munich, Germany. Springer, 2009
De Santis, G., et al. How can cells sense the elasticity of a substrate?: an analysis using a cell tensegrity model. Eur. Cells Mater.. 22:202–213, 2011.
Chen, T.-J., et al. Complexity of the tensegrity structure for dynamic energy and force distribution of cytoskeleton during cell spreading. PloS one.5(12):e14392, 2010.
Bansod, Y., V.V. Jakka, and J. Burša, Bendo-tensegrity model simulates compression test of animal cell
Mehrbod, M., and M. R. Mofrad. On the significance of microtubule flexural behavior in cytoskeletal mechanics. PLoS one.6(10):e25627, 2011.
Khunsaraki, G. M., H. N. Oscuii, and A. Voloshin. Study of the mechanical behavior of subcellular organelles using a 3D finite element model of the tensegrity structure. Appl. Sci.. 11(1):249, 2020.
Zhang, Y., et al. Novel MEMS-based thermometer with low power consumption for health-monitoring network application. in Device and Process Technologies for Microelectronics, MEMS, Photonics, and Nanotechnology IV. 2008. SPIE.
Kenner, H. Geodesic math and how to use it. Univ of California Press, 1993
Alieva, I. B., et al. Microtubules growth rate alteration in human endothelial cells. J. Biomed.Biotechnol..2010:671536, 2010.
Micheletti, A. and D. Cadoni. Design of single-layer floating-compression tensegrities. in 10e colloque national en calcul des structures. 2011.
Barreto, S., et al. A multi-structural single cell model of force-induced interactions of cytoskeletal components. Biomaterials. 34(26):6119–6126, 2013.
Stricker, J., T. Falzone, and M. L. Gardel. Mechanics of the F-actin cytoskeleton. J. Biomech.. 43(1):9–14, 2010.
Ladjal, H., et al. Atomic force microscopy-based single-cell indentation: Experimentation and finite element simulation. in 2009 IEEE/RSJ International Conference on Intelligent Robots and Systems. 2009. IEEE.
Simulia, D., ABAQUS 6.11 analysis user's manual. Abaqus. 6: 2.2, 2011.
Nguyen, T. D., and Y. Gu. Determination of strain-rate-dependent mechanical behavior of living and fixed osteocytes and chondrocytes using atomic force microscopy and inverse finite element analysis. J .Biomech. Eng..136(10):101004, 2014.
Voloshin, A. Modeling cell movement on a substrate with variable rigidity. Int. J. Biomed. Eng. Sci. (IJBES). 3(1):19–36, 2016.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by all authors. The first draft of the manuscript was written by Gholamreza Mohammadi Khunsaraki and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Additional information
Associate Editor Chiara Bellini oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khounsaraki, G.M., Movahedi, M., Oscuii, H.N. et al. Analysis of the Adherent Cell Response to the Substrate Stiffness Using Tensegrity. Ann Biomed Eng 52, 1213–1221 (2024). https://doi.org/10.1007/s10439-024-03447-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-024-03447-7