Abstract
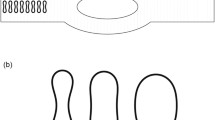
Movement, deformation, and partitioning of mammalian red blood cells (RBCs) in diverging microvessel bifurcations are simulated using a two-dimensional, flexible-particle model. A set of viscoelastic elements represents the RBC membrane and the cytoplasm. Motion of isolated cells is considered, neglecting cell-to-cell interactions. Center-of-mass trajectories deviate from background flow streamlines due to migration of flexible cells towards the mother vessel centerline upstream of the bifurcation and due to flow perturbations caused by cell obstruction in the neighborhood of the bifurcation. RBC partitioning in the bifurcation is predicted by determining the RBC fraction entering each branch, for a given partition of total flow and for a given upstream distribution of RBCs. Typically, RBCs preferentially enter the higher-flow branch, leading to unequal discharge hematocrits in the downstream branches. This effect is increased by migration toward the centerline but decreased by the effects of obstruction. It is stronger for flexible cells than for rigid circular particles of corresponding size, and decreases with increasing parent vessel diameter. For unequally sized daughter vessels, partitioning is asymmetric, with RBCs tending to enter the smaller vessel. Partitioning is not significantly affected by branching angles. Model predictions are consistent with previous experimental results.







Similar content being viewed by others
References
Ascher U. M., L. R. Petzold. Computer Methods for Ordinary Differential Equations and Differential-Algebraic Equations. Philadelphia: Society for Industrial and Applied Mathematics, 1998, pp. 73–82
Audet D. M., W. L. Olbricht 1987 The motion of model cells at capillary bifurcations. Microvasc. Res. 33, 377–396. doi:10.1016/0026-2862(87)90029-X
El-Kareh A. W., T. W. Secomb 2000 A model for red blood cell motion in bifurcating microvessels. Int. J. Multiphase Flow 26, 1545–1564. doi:10.1016/S0301-9322(99)00096-8
Enden G., A. S. Popel 1992 A numerical study of the shape of the surface separating flow into branches in microvascular bifurcations. J. Biomech. Eng. 114, 398–405. doi:10.1115/1.2891401
Enden G., A. S. Popel 1994 A numerical study of plasma skimming in small vascular bifurcations. J. Biomech. Eng. 116, 79–88. doi:10.1115/1.2895708
Fung Y. C. 1973 Stochastic flow in capillary blood vessels. Microvasc. Res. 5, 34–48. doi:10.1016/S0026-2862(73)80005-6
Fung, Y. C. Biodynamics: Circulation. New York: Springer-Verlag, pp. 241–249, 1984.
Goldsmith, H. L. 1971 Red cell motions and wall interactions in tube flow. Fed. Proc. 30, 1578–1590
Leal, L. G. 1980 Particle motions in a viscous-fluid. Annu. Rev. Fluid Mech. 12, 435–476. doi:10.1146/annurev.fl.12.010180.002251
Mayrovitz H. N., J. Roy 1983 Microvascular blood flow: evidence indicating a cubic dependence on arteriolar diameter. Am. J. Physiol. 245, H1031–H1038
Pries A. R., K. Ley, M. Claassen, P. Gaehtgens 1989 Red cell distribution at microvascular bifurcations. Microvasc. Res. 38, 81–101. doi:10.1016/0026-2862(89)90018-6
Pries A. R., K. Ley, P. Gaehtgens 1986 Generalization of the Fahraeus principle for microvessel networks. Am. J. Physiol. 251, H1324–H1332
Pries, A. R. and T. W. Secomb. Blood flow in microvascular networks. In: Handbook of Physiology: Section 2: The Cardiovascular System, Vol. 4. Microcirculation, edited by R. F. Tuma, K. Ley, and W. N. Duran. Academic Press, in press, 2008
Pries A. R., T. W. Secomb 2005 Microvascular blood viscosity in vivo and the endothelial surface layer. Am. J. Physiol. Heart Circ. Physiol. 289, H2657–H2664. doi:10.1152/ajpheart.00297.2005
Pries A. R., T. W. Secomb, P. Gaehtgens 1996 Biophysical aspects of blood flow in the microvasculature. Cardiovasc. Res. 32, 654–667. doi:10.1016/0008-6363(96)00065-X
Pries A. R., T. W. Secomb, P. Gaehtgens 2000 The endothelial surface layer. Pflugers Arch. 440, 653–666. doi:10.1007/s004240000307
Schmid-Schonbein G. W., R. Skalak, S. Usami, S. Chien 1980 Cell distribution in capillary networks. Microvasc. Res. 19, 18–44. doi:10.1016/0026-2862(80)90082-5
Secomb T. W., B. Styp-Rekowska, A. R. Pries 2007 Two-dimensional simulation of red blood cell deformation and lateral migration in microvessels. Ann. Biomed. Eng. 35, 755–765. doi:10.1007/s10439-007-9275-0
Acknowledgments
This work was supported by NIH grants HL034555 and HL07249, DOE Grant DEFG0202ER25533, NSF Grant DMS-0602173 (VIGRE) and the ARCS Foundation.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
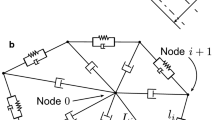
Finite elements are used to express Eqs. (5–6) as a system of linear equations. This system is coupled with the 2(n + 1) equations arising from the force and moment balances for each node and the appropriate boundary conditions. The coupled linear system is solved to obtain the instantaneous velocities of the nodes on the cell. The finite element package FlexPDE (version 5.0.19) was used. In the simulations, the RBC membrane is discretized using n = 20 nodes. Between 100 and 400 fluid elements are used. The finite element package uses adaptive meshing to resolve narrow gaps between cells and walls, according to a user-specified tolerance. On a 1.66 GHz processor machine, computing the instantaneous velocities takes 10–20 s.
Given cell nodal positions at time t n, the nodal positions at some subsequent time t n+1 = t n + dt are found using an explicit trapezoidal, order-2 method.1 In this scheme, the instantaneous velocities, u n i , are obtained using the finite element package (superscript corresponds to time t n). The nodal positions at time t n+1 are predicted using a forward Euler estimate:
These new nodal positions are used to obtain the instantaneous velocities \( \tilde{\user2{u}}^{{n + 1}}_{i} \) of the cell nodes at time t n+1. The final estimates of the nodal positions at time t n+1 are given by:
A time step of 1 ms was found to give numerical stability as well as good convergence, whereas the forward Euler method required a time step of around 0.2 ms for convergence. Generating a cell trajectory with the given method and time step usually takes 10–40 min on a 1.66 GHz processor machine.
To find y c for a given vessel geometry and Ψ1 using a bisection algorithm, quantities y below and y above are found such that y c is in the interval [y below, y above]. The particle trajectory for (y below + y above)/2 is then found, and the interval is halved, so that y c is in the new interval. The algorithm is repeated until the size of the interval was less than ε bisection = 0.05w 0. The midpoint of this final interval is then used as an estimate for y c; ε bisection can be viewed as an estimate of the error on this y c. If a particle starting within 0.05 μm of either the top or bottom mother vessel wall enters branch 1 or branch 2 (respectively), all cells are presumed to enter only one branch and y c(Ψ1) is taken as y t or y b, as appropriate.
To compute the upstream particle velocity profile, the instantaneous velocities of circular particles placed at nine locations across the channel are computed. The arithmetic average of the external node velocities is fitted with a quartic function (r 2 > 0.98) to obtain an estimate for u d(y), for use in Eq. (7).
Rights and permissions
About this article
Cite this article
Barber, J.O., Alberding, J.P., Restrepo, J.M. et al. Simulated Two-dimensional Red Blood Cell Motion, Deformation, and Partitioning in Microvessel Bifurcations. Ann Biomed Eng 36, 1690–1698 (2008). https://doi.org/10.1007/s10439-008-9546-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-008-9546-4