Abstract
Climatic variation along the elevation gradient promotes the natural parapatric occurrence of the European hare (Lepus europaeus) and Alpine mountain hare (Lepus timidus varronis) in the Alps. Recent data indicate a displacement of mountain hares caused by competition with the European hare. Competitive exclusion might take place at a fine spatial scale and hybrids may sharpen competition. Genetic non-invasive sampling (gNIS) demonstrates to be effective to retrieve information from wild animals. However, based on the accuracy of the differing genetic analysis methods, the selection of the method might decisively influence results. To examine habitat preferences of Alpine mountain hares, European hares and their hybrids with particular interest in the influence of the accuracy of the genetic analysis method on the results, we performed gNIS in Grisons (Switzerland) for four years and compared habitat associations of the genotyped samples. We recorded 137 individuals (i.e., 35 hybrids, 49 European hares, 53 Alpine mountain hares). Combined nuclear and mitochondrial DNA analysis including individual identification revealed to be the most accurate indirect method for the study of habitat preferences of hares. Alpine mountain hares had a narrow habitat breadth and used little habitat diversity. Hybrids showed great similarities in their habitat preferences to European hares. Hybrids might increase the competition in favour of European hares and the displacement of Alpine mountain hares, since they show similar patterns of habitat use to European hares. Ongoing climate change potentiate the niche overlap between species, increasing the risk of Alpine hare decline due to hybridisation and displacement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The European hare (Lepus europaeus) is an important prey and game species in Europe and numerous studies have been investigating the biology and ecology of this species (for an overview see Hackländer 2022). Inhabiting originally the steppes of the Middle East, the European hare has been spreading in the agricultural landscapes of Europe. As a habitat generalist, this lagomorph species inhabits a range of different habitats (Hackländer 2022). In contrast, another Lepus species living in Europe, the mountain hare (Lepus timidus), is a habitat specialist and well adapted to cold and harsh climate (Angerbjörn and Schai-Braun 2023). After the end of the last glacial period (Weichsel) when the ice cover over Europe slowly melted, the mountain hares colonised Scandinavia (Angerbjörn and Schai-Braun 2023), but an isolated population remained in the Alps in Central Europe, where the subspecies Alpine mountain hare (L. t. varronis) has been described (Angerbjörn and Schai-Braun 2023).
Although mountain hares and European hares in general occur allopatrically (for an overview see Schai-Braun and Hackländer 2016), the climatic variation along the elevation gradient resulted in a natural parapatric occurrence of the two congeneric hare species in the Alps. Since the beginning of the 20th century, the European hare distribution is expanding to the north in Scandinavia (Thulin 2003) and displacing the mountain hare distribution further north (Thulin 2003). In line with this, an evaluation of hunting bag statistics collected over 30 years in Grisons (Switzerland) showed that both hare species shifted their minimum elevations towards higher elevations in the Alps (Schai-Braun et al. 2021). Whether the possible displacement of mountain hares to the north and to higher elevations is caused by climate change or by direct competition with the European hare is widely discussed and still under investigation (see Schai-Braun and Hackländer 2016) but climate data confirm that climate warming has occurred within the last 100 years (IPCC et al. 2014, 2021). As well as spatial displacement, introgressive hybridisation poses a threat to the Alpine mountain hare since introgression, i.e. transfer of genes between species induced principally by backcrossing, reduces the subspecies’ genetic integrity. Hybridisation has been confirmed in hares from Grisons (Marques et al. 2017; Zachos et al. 2010) and South Tyrol (Italy), where also F2 hybrids were recorded (Schai-Braun et al. 2023).
Interspecific competition between the two congeneric hare species is expected to be strong due to the high potential for niche overlap between closely related species (Gidoin et al. 2015). Stable coexistence of species requires that species possess ecologically different niches, to that they can avoid or reduce interspecific competition for essential resources (Hardin 1960). Ecological niche theory predicts that two similar and closely related species in sympatry must reduce interspecific competition by partitioning at least one resource (Hutchinson 1959). By differing in their ecological niche, competing species develop divergence in resource-exploiting traits (Brown, JR. and Wilson 1956), which may include morphological, ecological, behavioural, or physiological characters. The coexistence of some sympatric species can be explained by their temporal (mammals: Thorén et al. 2011) or spatial separation (birds: Holmes and Robinson 1988; mammals: Rakotondranary et al. 2011). Understanding the mechanisms by which competition takes place is crucial, both for predicting individual behaviour and resource use and for understanding community processes (Tilman 1987).
Both hare species are mostly nocturnal (Schai-Braun and Hackländer 2016) and showed similar circadian activity patterns (mountain hare: Cederlund and Lemnell 1980; European hare: Holley 2001; Pépin and Cargnelutti 1994; Schai-Braun et al. 2012). Hence, the two lagomorph species exhibit probably not a strong niche differentiation by temporal separation. A spatial separation of the two hare species along the elevation gradient in the Alps has been demonstrated (Schai-Braun et al. 2023). However, competitive exclusion might take place at a fine spatial scale. Consistently reported habitat preferences of European hares living at low elevations are different types of fallow land (Bertolino et al. 2011a, b; Schai-Braun et al. 2013; Smith et al. 2004) due to its characteristic open vegetation. However, no knowledge exists about habitat preferences of this species in the Alps.
Alpine mountain hares select dwarf mountain pine as habitat throughout the year (Bisi et al. 2013). But as this subspecies lives in areas not readily accessible to humans, more detailed studies on habitat preferences are missing. Not only more details on selected habitat types but also other habitat characteristics such as land use and ground cover, habitat breadth and habitat diversity might reveal processes of competitive exclusion. Furthermore, the plant vegetation growth period in the Alps is much shorter than in lowland areas (Alps: middle of May-middle of September, 4 months; lowland: April-October, 7 months, Calanca and Holzkämper 2010) which might additionally influence the competition between the two lagomorph species.
Being a habitat generalist, the European hare may have some advantages over the Alpine mountain hare, a habitat specialist, also in the Alps. Generalists have a wider habitat breadth and, thus, a higher used habitat diversity, which might counterbalance especially in lower Alpine elevation ranges the disadvantage of being not specifically adapted to cold and rough climate. On the other hand, suitable food sources for hares are less available already in September (i.e., at the end of the plant vegetation growth period) in Alpine habitat (Schai-Braun et al. 2020). European hares that are not adapted to the short plant vegetation growth period in the Alps might show a wider habitat breadth to include suitable food sources at the end of the Alpine plant vegetation growth period than Alpine mountain hares that are adapted to the early start in food reduction.
Hybrids can shape evolutionary processes and contribute to species and subspecies loss (Allendorf et al. 2001; Laikre et al. 2010; Sakai et al. 2001). Hence, it is important to gain information on how hybrids shape competition between these two lagomorphs. Yet information on the ecology of the hybrids between these two hare species is unknown. It remains unclear, whether hybrids are ecologically similar to one of the parental hare species or intermediate. The understanding of these questions will reveal insights on adaptation of mammalian hybrids to their environment, but also will give further insights for species conservation, namely for the Alpine mountain hare.
Genetic non-invasive sampling (gNIS) is a widely used method to retrieve information from wild living animal species without disturbing them (Schwartz et al. 2007). For instance, gNIS might be used for providing species identification, assessing population structure and individuals (Ferreira et al. 2018), namely in rare and illusive species, as well as when densities are low or capturing and handling the individuals demand intense human resources (Sabino-Marques et al. 2018). Moreover, gNIS provides additional relevant information on population parameters, namely genetic diversity, inbreeding, dispersal, parental relationships, among others. Furthermore, the samples are obtained without the need to capture, handle and manipulate the individuals, thus avoiding stress and producing harmful situations.
Often wildlife biologists and stakeholders have to decide on the genetic analysis methods based on a cost-benefit consideration when planning a project. The cost of a genetic analysis increases steeply as more precise and more detailed information is pursued. Among the used molecular markers and type of samples, fragments of mitochondrial DNA (mtDNA) from faeces show a high success rate and are cost-efficient, namely to species identification. One of the reasons of the higher success rate is that mtDNA extraction of older faeces is more likely to occur, simply because numerous mitochondria containing mtDNA are present in a cell. Hence, more samples are included in the analysis producing ample data. However, because mtDNA is maternally inherited, the solely use of mtDNA does not allow the individual identification and it is imprecise for detecting hybridisation (Brito et al. 2020; Ferreira et al. 2018). Consequently, hybrids are not perceived in regions in which hybridisation takes place and samples of hybrids are identified erroneously as one of the hybridising species.
The results of mtDNA analysis might be the cause of other errors when individuals are recaptured unwittingly several times and bias the results of further analysis. On the other hand, analysis of nDNA is much more expensive and more successful when using samples with higher quality, i.e. fresh samples (Rehnus and Bollmann 2016). When using faeces, this means a lower number of successful genotyped samples can be included in the analysis or collection effort must be increased substantially in order to collect the fresher samples. Both high cost of analysis and large effort for collecting samples enhance the budget considerably. Nevertheless, the use of nDNA, namely microsatellites or SNPs (Single Nucleotide Polymorphisms), are highly precise for detecting hybridisation, individual and gender identification, and enabling further kinship analyses (e.g., Ferreira et al. 2018; Jones et al. 2010; Jones and Wang 2010) or estimation of population densities using capture-recapture methods (e.g., Borchers and Efford 2008; Efford and Fewster 2013). Thus, the selection of the genetic analysis methods, i.e., the selection of genetic markers, might decisively influence the outcome of the research questions of a study, especially in hybridising species.
The goal of this study was to investigate habitat preferences and spatial distribution of Alpine mountain hares, European hares and their hybrids in the Alps with particular interest in the influence of the accuracy of the genetic analysis method on the results. Our hypotheses were: (1) the European hare, a habitat generalist, exhibits a greater habitat breadth than hybrids, and hybrids a greater habitat breadth than the Alpine mountain hare, a habitat specialist; (2) habitat breadth of European hares at the end of the plant vegetation growth period is greater than habitat breadth of Alpine mountain hares; (3) spatial overlap between the European hare and the Alpine mountain hare is lower than between each hare species and the hybrids; (4) the habitat generalist European hare uses a higher habitat diversity than hybrids and hybrids a higher habitat diversity than the habitat specialist Alpine mountain hare; (5) the accuracy of the genetic analysis method (mtDNA vs. mtDNA/nDNA vs. individual identification based on microsatellites) has a decisive effect on the outcome of the research questions, especially in hybridising species. To test these hypotheses, we performed genetic analysis of hare faecal samples collected on seven transects along the altitudinal gradient in the Alps in Grisons (Switzerland) for four years in the middle and at the end of the plant vegetation growth period. We then investigated habitat preferences, spatial overlap and used habitat diversity of the two hare species and their hybrids using habitat types and land use and ground cover maps.
Materials and methods
Study area
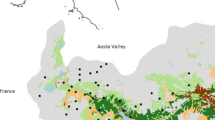
The study area comprised seven transects between the villages of Susch and Ramosch in Grisons, Switzerland (Susch 46°44′N, 10°4′E, Ramosch 46°49′N, 10°23′E, Online Resource Fig. 1). These cover an altitude range between 1,000 and 2,600 m a.s.l. We used two data sets describing the habitat types (Bronwyn Price et al. 2021) and the land use and ground cover of Grisons as habitat maps for our study area. A rough categorisation (habitat type rough categorisation) containing nine categories pooled all the 45 habitat types of the study area into broader groups (Table 1a). The land use and ground cover of our study area consisted of 73 variables (Table 1a).
Data collection
Faecal samples were collected yearly between 2019 and 2022 on the five transects of the south exposed mountain slopes along the altitudinal gradient. We additionally included two transects on the north exposed mountain slopes in the same valley in the year 2022. Each transect was searched twice in the middle and at the end of the plant vegetation growth period (beginning of July and end of August/beginning of September). Faeces with the same freshness and size within two meters or less from each other were considered as the same sample. Preference was given to fresh faecal pellets as older pellets have significantly lower amplification rates (Rehnus and Bollmann 2016). One to six pellets were collected for each sample and stored in 50-ml tubes filled with Silica or in ethanol for the genetic analysis. Each sample location was recorded with a GPS device (Garmin GPSMAP 60Cx) and consisted of a minimum of one pellet. The faeces were collected with gloves to avoid contamination with human DNA.
Around each point where a faecal sample was mapped, a circle with the size of 10 ha was drawn. We chose 10 ha circles since small home range sizes of European hares in agricultural and Alpine landscapes are recorded to be around 10 ha (Schai-Braun and Hackländer 2014 and unpublished data). Subsequently, the landscape composition (habitat type, land use and ground cover) within the circle was evaluated assuming that the landscape within the circle was actually being used by the individual hare. Habitat availability was determined using the two habitat maps. The availability of each habitat variable was calculated in percent for every transect. Note that we performed the same analysis with larger assumed home-ranges (25 ha circles), but the results did not differ substantially (data not shown). No smaller (< 10 ha) or larger (> 25 ha) home range sizes were considered as our data of recaptured individuals does not support it.
Genetic analysis
DNA was extracted from the faecal pellets using the QIAamp DNA Stool Mini Kit (Qiagen, Valencia, California, USA). A molecular marker (219 bp) of the mitochondrial control region (mtDNA) was amplified using the primers Lmtof1 and LmtNr1 (Palumbi et al. 2002), and sequencing was performed after purification of the amplified samples using the Lmtof1 primer. All faecal samples with a successful species determination based on mtDNA were pooled subsequently in the mtDNA group. Accordingly, the mtDNA group was based on mtDNA data only and assigned the species without being able to detect introgression.
Two short nuclear DNA markers (nDNA) containing species-specific nuclear SNPs were then amplified only in samples for which the mtDNA marker had been successfully amplified. Fragments of SMCX and Albumin nuclear genes were amplified following the procedures described by Melo-Ferreira et al. (2009). Sanger sequencing was performed after purification of amplified products (176 bp for SMCX and 137 bp for Albumin). These fragments of nuclear genes have been shown to present fixed speciesspecific SNPs in both species (Melo-Ferreira et al. 2009), and thus allow the species diagnosis and detection of hybridization. This analysis, together with the inspection of the mtDNA lineage, also allowed determination of mtDNA introgression. We also genotyped microsatellites in those samples with successful mtDNA amplification. We used a battery of 16 hare microsatellites (Le2, Le3, Le4, Le6, Le15, Le17, Le19, Le23, Le28, Le30, Le43, Le45, Le46, Le47, Le48 and Le51) developed in the CIBIO/BIOPOLIS laboratory, which revealed to be highly polymorphic and informative for identifying both species and hybrids (Costa et al. 2023). These markers were amplified in three multiplex PCR reactions. The first screening was done with one multiplex, and only those samples that amplified for this first multiplex, were genotyped for the remaining two multiplexes. All faecal samples with a successful species determination based on mtDNA and nDNA (SMCX and Albumin genes, microsatellites) were pooled subsequently in the mtDNA/nDNA group. Correspondingly, the mtDNA/nDNA group was based on mtDNA and nDNA and assigned the species identity including the detection of introgression (= hybrids).
Based on CIBIO’s reference microsatellite dataset for the mountain and the European hare (39 Alpine mountain hares from Switzerland and 54 European hares from Germany, France, Greece, Austria and Spain), the individual probability of identity (PI) and sibling identity (Plsibs) were inferred using GENALEX, version 6.503 (Peakall and Smouse 2012). The values of PI per population varied from 4.5 × 10-10 to 9.8 × 10-16, while Plsibs ranged from 1.2 × 10-5 to 5.9 × 10-6. Individual identification was performed using IRMACRON (Amos et al. 2001). All faecal samples with a successful individual determination based on microsatellites were pooled subsequently in the individual group. Thus, the individual group was based on microsatellites and assigned individual identity.
In addition to individual identification, the microsatellites were further used to assess the degree of genetic admixture between species using STRUCTURE software, version 2.3 (Pritchard et al. 2000). Default STRUCTURE parameters were set together with an admixture model in combination with correlated allele frequencies and no prior-information about population origin. The log likelihood of the data In (P(X│K) was calculated for K = 1 to K = 4, with 5 repetitions of 106 MCMC iterations following a burn-in period of 105 steps. Moreover, ΔK was calculated following (Evanno et al. 2005) using STRUCTURE HARVESTER 0.6.94 (Earl and vonHoldt 2012), and revealed ΔK = 2 as the best model. To determine the probable generation of the recent hybrid, we simulated 30 genotypes in the HibridLab software (Nielsen et al. 2006) for each hybrid class (F1, F2, BC1, BC2) using 66 parental European hares and 40 parental mountain hares from the Alps, and then performed the STRUCTURE analysis with the same settings as above to infer the range values of the membership proportion of each hybrid class (Queirós et al. 2020).
Statistical analyses
All statistical analyses were carried out using the software R 4.3.1 (R Core Team 2023).
Preference indices
The hares’ habitat preferences were measured by using Chesson’s electivity index ε (Chesson 1983), an index based on Manly’s alpha (Manly et al. 1972) which can be used to analyse habitat preferences (Krebs 1989), among others. We chose Chesson’s electivity index, because it has the advantage that results between cases for which the number of available habitat categories vary are comparable. The Chesson’s electivity index ranges between − 1 and + 1, with negative values showing a negative selection, whereas positive values signify a positive selection. If the index value is zero, the habitat variable concerned is used in the same proportion as it is available. We calculated the Chesson’s electivity indices for each habitat map (habitat type, habitat type rough categorisation, land use and ground cover) for the three groups mtDNA, mtDNA/nDNA, and individuals. For individual hares, the mean of all habitat categories was determined before calculating preference indices. Note that due to the pooling into the three groups, particular faecal samples were included in several of the three groups. Additionally, we examined the Chesson’s Electivity Indices in the middle and at the end of the plant vegetation growth period.
The reliability of the electivity indices were tested using the bootstrap method (Dixon 1993). The original εi values (εi = Chesson’s electivity index for the habitat variable i) were resampled 1000 times with replacement and an accelerated bootstrap confidence interval (CI) was calculated. The accelerated bootstrap adjusted the CI for bias and skewness (Efron and Tibshirani 1993). If the two values of the lower and upper boundary (95% CI) featured the same algebraic sign, the selection for this habitat variable was significant. If the lower and upper boundary featured different algebraic signs, the selection of the respective habitat variable was not significant (n.s.). εi values for habitat categories were only bootstrapped if they were selected by six or more hares, as smaller sample sizes provide unreliable results. Bootstrapping was done using the R package boot (Canty and Ripley 2019).
Spatial overlap
The degree of spatial overlap between the two hare species and their hybrids was estimated with Pianka’s index of niche overlap (Pianka 1973) using the EcoSimR package (Gotelli et al. 2015). We tested for significance by comparing used habitat resources with values obtained by randomizing the original matrices (1000 iterations), using the default procedure ra3 as randomisation algorithm. Pianka’s index of niche overlap was calculated for each habitat map (habitat type, habitat type rough categorisation, land use and ground cover) separately, and used habitat was defined as the landscape composition within a 10 ha circle around each faecal sample. Pianka’s index ranges from 0 (exclusive resource categories) to 1 (similar resource categories). It is considered that values of Pianka’s index higher than 0.6 means that there is a biologically significant niche overlap for the resources (Wallace 1981). This procedure was conducted for the mtDNA, mtDNA/nDNA and individual group.
Habitat diversity
We used Shannon Wiener index to estimate available and used habitat diversity of the hares. Shannon Wiener index measures diversity in the sense of number of types and number of samples with 0 describing no diversity (Spellerberg and Fedor 2003). We calculated Shannon Wiener index for each habitat map (i.e., habitat type, habitat type rough categorisation, land use and ground cover) and group (i.e., mtDNA, mtDNA/nDNA, individuals) with the vegan package (Oksanen et al. 2022). Generalized linear mixed-effects models were fitted using the package lme4 (Bates et al. 2015). The response variable Shannon Wiener index was investigated with models including the covariate type (used by species/hybrids vs. available).
All models included the variable transect as random factor in order to account for the repeated measurements collected from the different transects. As the program R does not directly provide p-values for such models, the p-values were extracted by likelihood ratio tests (Faraway 2006). Residuals of the models were checked for normal distribution by QQ-plots and histograms. The homogeneity of variances and goodness of fit were examined by plotting residuals versus fitted values (Faraway 2006).
Results
Faecal pellet sampling
A total of 1257 faecal samples were collected between 2019 and 2022. Of these, 776 were collected in the middle of the plant vegetation growth period (July), and 481 at the end (August/September). From the initially tested faecal pellet samples, a total of 1079 (86%) amplified for Cytochrome b (see Online Resource Appendix S1). Hence in the mtDNA group, a total of 318 (29%) were assigned to the European hare, while 761 (71%) were assigned to the Alpine mountain hare. From the 1079 samples of the mtDNA group, 423 (39%) amplified successfully for the SMCX and/or Albumin nuclear genes and/or for the microsatellites. Hence in the mtDNA/nDNA group, 165 were assigned to the European hare, 149 to the Alpine mountain hare, and 109 samples showed signs of mtDNA introgression, and thus hybridization (see as an example Online Resource Appendix S1, Sample ID #56 having as Species ID L. europaeus based on mtDNA and L. timidus based on the nDNA gene Albumina). The great majority of the introgressed individuals (107) had mtDNA of the Alpine mountain hare and nDNA of the European hare.
A total of 137 different individuals (European hare n = 49, Alpine mountain hares n = 53, hybrids n = 35) were identified from 423 samples genotyped for microsatellites (see Online Resource Appendices S2 and S3). Only one hybrid showed an admixture pattern between species in the nDNA (see Online Resource Appendix S1, Sample ID #115, having as Species ID L. europaeus and L. timidus based on nuclear microsatellites). The STRUCTURE results of the simulated genotypes for each hybrid class: F1, F2, BC1, BC2 suggested that this individual (qi-EUR = 0.742 and qi-TIM = 0.258) falls within the membership (qi) range values of the cross between F1 hybrid and a European hare (average qi-EUR 0.749, 95%CI 0.715–0.843) or between F2 hybrid and European hare (average 0.721, 95CI 0.688–0.815). All other hybrids showed discordant species assignment between mtDNA and nDNA, hence, were characterized by an older interspecific gene flow. Recapture rate was on average 2.48 (± 2.06 SD, min = 1, max = 12) for European hares, 1.70 (± 1.81 SD, min = 1, max = 13) for Alpine mountain hares, and 2.54 (± 3.10 SD, min = 1, max = 13) for hybrids (Online Resource Appendix S4).
European hare faecal pellets were collected within an altitudinal range of 1,215–2,345 m a.s.l. (mean = 1,689.94 m a.s.l., SD = 270.21 m a.s.l.), Alpine mountain hare within 1,604–2,570 m a.s.l. (mean = 2,149.34 m a.s.l., SD = 150.13 m a.s.l.), and hybrids within 1,106–2,198 m a.s.l. (mean = 1,575.54 m a.s.l., SD = 261.56 m a.s.l., for an overview of the elevation ranges see Online Resource Appendix S5).
Preference indices
A comparison of the preference indices between the three groups (i.e., mtDNA, mtDNA/nDNA, individuals) revealed that differences were much higher between the individual and mtDNA/nDNA group than between the mtDNA/nDNA and mtDNA group (Table 2) except for habitat type preferences of Alpine mountain hares. Based on the comparison, only preference indices of individual hares are reported in detail, whereas preference indices of the mtDNA and mtDNA/nDNA group are available as Online Resource Appendices S6, S7, S8, S9, S10, S11 and Figs. 2, 3 and 4).
Chesson’s electivity indices in individual (a) European hares (n = 49), (b) hybrids (n = 35), and (c) Alpine mountain hares (n = 53; species determined by SMCX and Albumin genes, and microsatellites) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat types of the habitat type map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type). Faecal samples were collected in Grisons (Switzerland) during the years 2019, 2020, 2021, and 2022. Non-significant results cross the vertical line at zero. See text for statistical details
Chesson’s electivity indices in individual (a) European hares (n = 49), (b) hybrids (n = 35), and (c) Alpine mountain hares (n = 53; species determined by SMCX and Albumin genes, and microsatellites) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat type categories of the habitat type rough categorisation map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type category). Faecal samples were collected in Grisons (Switzerland) during the years 2019, 2020, 2021, and 2022. Non-significant results cross the vertical line at zero. See text for statistical details
Chesson’s electivity indices in individual (a) European hares (n = 49), (b) hybrids (n = 35), and (c) Alpine mountain hares (n = 53; species determined by SMCX and Albumin genes, and microsatellites) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat variables of the land use and ground cover map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat variable). Faecal samples were collected in Grisons (Switzerland) during the years 2019, 2020, 2021, and 2022. Non-significant results cross the vertical line at zero. See text for statistical details
For the habitat type map, 24 different habitat types were found in more than six faecal pellet samples and thus provided reliable Chesson’s Electivity Indices in the European hare, 23 different habitat types provided reliable indices in the hybrids, and 19 different habitat types provided reliable indices in the Alpine mountain hare (Fig. 2). Of these, five habitat types were avoided (buildings; natural grassland; ruderal meadows; scree vegetation; watercourses) and nine selected (central European semi-arid grassland; forests; improved grassland (mountain, sea milk wort); individual trees; larch-pine forest; rough track/unpaved road; shrubs; thermophile dry grasslands; variegated fescue heap) by the European hare, four habitat types were avoided (buildings; dwarf shrub heaths; subcontinental calcareous pine forest; watercourses) and five selected (central European semi-arid grassland; improved grassland (mountain, sea milk wort); larch-pine forest; thermophile dry grasslands; variegated fescue heap) by the hybrids, and two habitat types were avoided (buildings; watercourses) and ten selected (bluegrass meadow; dwarf shrub heaths; forests; larch-pine forest; nardus grasslands; pioneer grasslands on rocky soils; rough track/unpaved road; shrubs; stagnant waters; variegated fescue heap) by the Alpine mountain hare.
For the habitat type rough categorisation map, seven habitat types provided reliable Chesson’s Electivity Indices in the European hare, seven habitat types in the hybrids, and six habitat types in the Alpine mountain hare (Fig. 3). Of these, three habitat types were avoided (ruderal sites; sand/gravel/rock/crushed rock; watercourse) and one selected (natural grassland) by the European hare, one habitat type was avoided (watercourse) and one selected (natural grassland) by the hybrids, and two habitat types were avoided (sand/gravel/rock/crushed rock; watercourse) and three selected (forests; natural grassland; shelterbelt/tall forb communities/shrubs) by the Alpine mountain hare.
Finally for the land use and ground cover map, 21 habitat variables provided reliable Chesson’s Electivity Indices in the European hare, 17 habitat variables in the hybrids, and 16 habitat variables in the Alpine mountain hare (Fig. 4). Of these, two habitat variables were avoided (forest; rock) and eight habitat variables were selected (brushland; bushes and trees; extensively used pastures; groves; less intensively cultivated meadows; overgrown Alpine pasture; pastures; summer grazing pastures) by the European hare, one habitat variable was avoided (forest) and six habitat variables were selected (brushland; extensively used pastures; groves; natural meadows; pastures; biodiversity promotion areas) by the hybrids, and one habitat variable was avoided (permanent meadows) and eight habitat variables were selected (Alpine meadows; brushland; extensively farmed meadows; extensively used pastures; forest; grassy and herbaceous vegetation; less intensively cultivated meadows; biodiversity promotion grassland) by the Alpine mountain hare.
In summary, the European hare exhibited a greater habitat breadth than hybrids and hybrids exhibited a greater habitat breadth than the Alpine mountain hare irrespective of the habitat map (habitat type map: 24 vs. 23 vs. 19; habitat type rough categorisation map: 7 vs. 7 vs. 6; land use and ground cover map: 21 vs. 17 vs. 16) Furthermore, the hares had more positive than negative associations with habitat variables irrespective of the habitat map except European hares with habitat types of the rough categorisation (Table 3).
Seasonal preference indices
There were more reliable preference indices in the middle of the plant vegetation growth period than at the end of the plant vegetation growth period for European hares, Alpine mountain hares, and their hybrids regardless which habitat map (i.e., habitat type, habitat type rough categorisation, land use and ground cover) was used for calculating the indices (Table 4). Hence, habitat breadth was always greater in the middle of the plant vegetation growth period than at the end of the plant vegetation growth period.
At the end of the plant vegetation growth period, the hares chose more habitat variables neutrally than in the middle of the plant vegetation growth period, whereas selections and avoidances of habitat variables were more numerous in the middle than at the end of the plant vegetation growth period irrespective of the habitat map. European hares showed the highest seasonal differences in selection (i.e., selection, avoidance, neutral) of habitat type, whereas Alpine mountain hares showed the highest seasonal differences in selection of land use and ground cover (Table 5).
Spatial overlap
The spatial overlap of habitat types between the European hare and the Alpine mountain hare was greatest in the mtDNA group with 0.84, slightly lower with 0.71 in the individual group and in the mtDNA/nDNA group with 0.65. The overlap between European hares and hybrids was significantly stronger than between Alpine mountain hares and hybrids, both in the mtDNA/nDNA group (European hares vs. hybrids = 0.86, Alpine mountain hares vs. hybrids = 0.34) and in the individual group (European hares vs. hybrids = 0.88, Alpine mountain hares vs. hybrids = 0.43, see Fig. 5). The spatial overlap of the habitat type rough categorisation ranged between 0.97 and 1.00 for all hare species/hybrids and groups, whereas the spatial overlap of the land use and ground cover ranged between 0.92 and 0.98 for all hare species/hybrids and groups.
Spatial overlap of habitat types based on Pianka’s index of niche overlap for mtDNA (dark grey), mtDNA/nDNA (light grey) and the individual group (white). Arrows represent variance of simulated index. Faecal samples were collected in Grisons (Switzerland) during the years 2019, 2020, 2021, and 2022. See text for statistical details
Habitat diversity
In the mtDNA group, the habitat diversity for each habitat map (i.e., habitat type, habitat type rough categorisation, land use and ground cover) was by far the highest in the available landscape compared to the used landscape by the hares, with European hares using a higher habitat diversity than Alpine mountain hares (p < 0.001, for an overview of all p-values see Online Resource Appendix S12). In the mtDNA/nDNA group, the habitat diversity for each habitat map was again much higher in the available landscape compared to the used landscape by the hares, with European hares and hybrids using a comparable higher habitat diversity than Alpine mountain hares (p < 0. 05).
In the individual group, the habitat diversity for each habitat map was again much higher in the available landscape compared to the used landscape by the hares (Fig. 6). Habitat type diversity was significantly higher in European hares than in Alpine mountain hares with hybrids in between the two hare species (Fig. 6a), whereas the habitat type diversity of the rough categorisation between the two hare species and their hybrids did not differ (Fig. 6b). The land use and ground cover diversity was higher in European hares and hybrids compared to Alpine mountain hares (Fig. 6c).
Habitat diversity available to and used by individual European hares (n = 49), Alpine mountain hares (n = 53), and their hybrids (n = 35). Habitat diversity was measured by the Shannon Wiener index (medians with 25th/75th and 10th/90th percentiles) using the (a) habitat type, (b) habitat type rough categorisation, and (c) land use and ground cover map. Faecal samples were collected in Grisons (Switzerland) during the years 2019, 2020, 2021, and 2022. See text for statistical details
Discussion
In the Alps, competitive exclusion between the European hare and the Alpine mountain hare might take place at a fine spatial scale. Hybrids may sharpen the competition between the two lagomorph species. Although gNIS is a suitable method to collect information, the accuracy of the differing genetic analysis methods and, thus, the selection of the method might decisively influence results. We recorded 137 individuals, i.e., 35 hybrids, 49 European hares, 53 Alpine mountain hares by performing gNIS and determined habitat preferences. The combination of nuclear and mitochondrial DNA analysis including individual identification revealed to be the most accurate indirect method for the study of habitat preferences of hares. Alpine mountain hares had a narrower habitat breadth and used less habitat diversity than European hares. Hybrids showed great similarities in their habitat preferences to European hares.
Species and hybrid assignment
The considerable change in the proportion of species assignment between the mtDNA and mtDNA/nDNA group was, on the one hand, due to the missing of hybrids in the mtDNA group but, on the other hand, due to predominantly unidirectional hybridisation (i.e., female Alpine mountain hares mating primarily male European hares) and introgression of mtDNA between these two lagomorph species (Thulin 2003). Accordingly, a large part of the Alpine mountain hares of the mtDNA group in our study were in fact backcrossed European hares, i.e., hybrids. This is consistent with the finding in the individual group that only two out of 35 introgressed individuals were backcrossed Alpine mountain hares.
We found a higher proportion of hybrids (26%) than have been recorded in Sweden (Thulin and Tegelström 2002; in European hares about 15% hybrids), in the Swiss Alps (Zachos et al. 2010; in Alpine mountain hares about 4% hybrids) or in South Tyrol (Schai-Braun et al. 2023; about 9% hybrids between Alpine mountain hare and European hare) but less hybrids than in Ireland (Reid et al. 2022: 34% hybrids between native Irish hare Lepus timidus hibernicus and non-native European hare). The high proportion of hybrids found in this study may be explained by our study area having almost equal high abundances of European hares and Alpine mountain hares. Recapture rates of European hares and hybrids were similar but much higher than the recapture rate of Alpine mountain hares. Recapture rate of individuals in Grisons in summer was approximately the same as in South Tyrol in winter (Schai‐Braun et al. 2023: on average 2.18).
Preference indices
Although proportions of species assignment changed the most between the mtDNA and mtDNA/nDNA group, differences of preference indices were highest between the individual and mtDNA/nDNA group. Hence, repeatedly sampled individuals biased preference indices more than erroneously assigned hybrids to Alpine mountain hares. Therefore, the combined nuclear and mitochondrial DNA analysis including individual identification is the preferable analysis method to investigate habitat preferences of species.
Some avoided or selected habitat variables were used similarly by the two hare species and their hybrids in Alpine habitat (e.g., avoidance: buildings, watercourses; selection: larch pine forest, natural grassland, brushland). Other habitat variables were used equally by Alpine mountain hares and European hares (e.g., avoidance: sand/gravel/rock/crushed rock; selection: shrubs, rough track/unpaved road), or European hares and hybrids (e.g., avoidance: forest; selection: pastures, improved grassland). In contrast, no habitat variables were used similarly by Alpine mountain hares and hybrids. Our results suggest that the two lagomorph species and their hybrids have a lot of commonly shared habitat requirements in the Alps but hybrids and Alpine mountain hares are the most ecologically dissimilar.
As a habitat generalist, the European hare exhibited a greater habitat breadth than the Alpine mountain hare, a habitat specialist, with the hybrids in between the two species. This is in line with the wider niche breadth of the European hare than of the Irish hare in northern Ireland, where both species occur in sympatry (Caravaggi et al. 2015). The more positive than negative associations with habitat variables may indicate that all hares had access to the habitat types they require. These results imply that the European hare is well adapted to inhabit Alpine habitat and, thus, may be a strong competitor for the congeneric Alpine mountain hare in the Alps.
Seasonal preference indices
Habitat breadth was wider in the middle of the plant vegetation growth period than at the end of the plant vegetation growth period. We explain this finding with the reproductive season being at its highest during early summer and declining in late summer for both hare species (Angerbjörn and Schai-Braun 2023; Hackländer 2022). Hares are most active during high peak of the reproductive season (Lincoln 1974) which might be the reason for the animals’ wider habitat breadth earlier than later in the reproductive season.
More selected and avoided habitat variables in the middle of the plant vegetation growth period but more neutrally selected habitat variables at the end of the plant vegetation growth period may be explained by the food availability for the hares. Diet preferences analysed for Alpine mountain hares suggested that more suitable food sources were available in the middle than at the end of the plant vegetation growth period in the Alps (Schai-Braun et al. 2020). Overall, we did not find large seasonal differences in habitat preferences between the European hare, the Alpine mountain hare and their hybrids.
Spatial overlap
When examining the habitat type spatial overlap of European hares and Alpine mountain hares, the results of the mtDNA/nDNA group and the individual group were lying closer together than the results of the mtDNA. When comparing the spatial overlap of the hybrids with each of the two hare species, both the mtDNA/nDNA group and the individual group showed similar results. Hence, it is strongly recommended to use combined nuclear and mitochondrial DNA analysis to investigate spatial overlap of habitat types, whereas additional analysis of individual identification does not reveal much more information.
Hybrids and European hares showed a very high spatial overlap of habitat types, whereas hybrids and Alpine mountain hares had a very weak spatial overlap in the Alps. This fits our findings that almost all hybrids were backcrossed European hares and, thus, are ecologically inhabiting similar habitat types than European hares.
When examining a broader categorisation of habitat types or land use and ground cover, the two hare species and their hybrids had almost a complete spatial overlap. Accordingly, a categorisation into too broad habitat types does not capture the slight ecological differences in habitat types used by the two hare species and their hybrids. To avoid a broad habitat categorisation is in line with a study about habitat preferences of European hares in an agricultural area (Schai-Braun et al. 2013).
Habitat diversity
The habitat diversity results of the mtDNA and mtDNA/nDNA group were similar except that in the mtDNA/nDNA group additionally habitat diversity of hybrids was revealed. However, results of the individual group exposed subtle details between the European hare, Alpine mountain hare and their hybrids. Consequently, combined nuclear and mitochondrial DNA analysis including individual identification is preferable for habitat diversity investigations.
The hares used little of the available habitat diversity along the transects. Hares’ locomotor behaviour is more localised when resource availability is high (Schai-Braun and Hackländer 2014). The Alpine habitat in the study area seems to provide access to food and shelter to the hares when using only a small part of the available habitat diversity.
In all three groups, Alpine mountain hares used less habitat diversity than European hares. As a habitat specialist, the Alpine mountain hare may be confined to specific habitats and, thus, used habitat diversity is lower than of the European hare, a habitat generalist. The habitat diversity analysis of the individual group revealed different use of habitat diversity of hybrids. The hybrids’ requirement of habitat type diversity laid in-between the two parental species, whereas the land use and ground cove diversity of hybrids was the same as of European hares. In comparison what was available and what was used by the other two hare species, hybrids expressed a relative high requirement of land use and ground cover diversity. Again, when pooling habitat types in a rough categorisation, information was lost and no differences in habitat diversity use between European hares, Alpine mountain hares and their hybrids was discernible.
Hybridisation events
We recorded only one hybrid from a recent hybridisation event out of 35 individual hybrids. This is less than was found in South Tyrol (four F2 hybrids out of 14 hybrids, Schai-Braun et al. 2023). Accordingly, the majority of the hybrids of our study seems to result from older hybridisation events. Our results suggest that differences in habitat use of the two lagomorph species and their hybrids occur at a fine spatial scale in the Alps. This might be the reason that there are not numerous recent hybridisation events noticeable. The two hare species seem to occupy habitat niches differing on the small spatial scale and, thus, avoiding frequent hybridisation. However, as ongoing global warming might favour the European hare in the Alpine habitat, these fine habitat niche differences might be threatened in future.
As both hare species live parapatrically in the study area since hundreds or thousands of years, we hypothesise that the hybridisation events of our hybrids might have taken place in the last decades and, thus, might represent contemporary hybridisation, but further genetic studies using genomic methods should be conducted to test this hypothesis. This inference is in contrast to ancestral hybridisation found on the Iberian Peninsula (Melo-Ferreira et al. 2005). In this case, mtDNA of mountain hares was recorded in introgressed European hares, Iberian hares (Lepus granatensis), and broom hares (Lepus castroviejoi) although mountain hares retreated from this region at the end of the last ice age (Melo-Ferreira et al. 2005). Consequently, the hybridisation events took place more than thousands of years ago when mountain hares were still inhabiting the Iberian Peninsula (Melo-Ferreira et al. 2005). Nevertheless, introgressed mtDNA seems to affect physiological traits in today’s Iberian hares carrying mountain hare mtDNA (Cardoso et al. 2024).
Conclusions
Combined nuclear and mitochondrial DNA analysis including individual identification revealed to be the most precise indirect method for the study of habitat preferences of hares using faecal samples. Not only subtle details were exposed, also the bias of repeatedly sampled individuals was avoided and the investigation of hybrids was possible. In particular, when studying species with known mtDNA introgression genetic analysis including nDNA is mandatory.
The Alpine mountain hare underlined in its habitat use to be a habitat specialist by having a narrower habitat breadth and using less habitat diversity than the European hare. The hybrids showed greater similarities in their habitat preferences to the European hare than to the Alpine mountain hare reflecting the predominantly unidirectional hybridisation and introgression of mtDNA between the two lagomorph species. Our findings imply that the European hare is well adapted to inhabit Alpine habitat and, thus, both hare species change from being parapatric to sympatric. European hares may be a strong competitor for the congeneric Alpine mountain hare in the Alps. As the hybrids are in terms of habitat use similar to the European hare, hybrids may increase the competition in favour of the European hare and to the disadvantage of the Alpine mountain hare. Moreover, as hybrids showed mostly an older interspecific gene flow, backcrossed European hares may even be better adapted to life at high elevations than native European hares under global warming.
Data availability
No datasets were generated or analysed during the current study.
References
Allendorf FW, Leary RF, Spruell P, Wenburg JK (2001) The problems with hybrids: setting conservation guidelines. Trends Ecol Evol 16:613–622. https://doi.org/10.1016/S0169-5347(01)02290-X
Amos W, Wilmer JW, Fullard K, Burg TM, Croxall JP, Bloch D, Coulson T (2001) The influence of parental relatedness on reproductive success. Proc Biol Sci 268:2021–2027. https://doi.org/10.1098/rspb.2001.1751
Angerbjörn A, Schai-Braun SC (2023) Mountain Hare Lepus timidus Linnaeus, 1758. In: Hackländer K, Zachos FE (eds) Handbook of the Mammals of Europe. Springer International Publishing, Cham, 1–29. In press
Bates DM, Maechler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48
Bertolino S, Cordero di Montezemolo N, Perrone A (2011a) Daytime habitat selection by introduced eastern cottontail Sylvilagus floridanus and native European hare Lepus europaeus in Northern Italy. Zool Sci 28:414–419. https://doi.org/10.2108/zsj.28.414
Bertolino S, Perrone A, Gola L, Viterbi R (2011b) Population density and habitat use of the introduced eastern cottontail (Sylvilagus floridanus) compared to the native European hare (Lepus europaeus). Zool Stud 50:315–326
Bisi F, Nodari M, Dos Santos Oliveira NM, Ossi F, Masseroni E, Preatoni DG, Wauters LA, Martinoli A (2013) Habitat selection and activity patterns in Alpine mountain hare (Lepus timidus varronis). Mamm Biol 78:28–33
Borchers DL, Efford MG (2008) Spatially explicit maximum likelihood methods for capture-recapture studies. Biometrics 64:377–385. https://doi.org/10.1111/j.1541-0420.2007.00927.x
Brito C, Vilaça ST, Lacerda AL, Maggioni R, Marcovaldi MÂ, Vélez-Rubio G, Proietti MC (2020) Combined use of mitochondrial and nuclear genetic markers further reveal immature marine turtle hybrids along the South Western Atlantic. Genet Mol Biol 43:e20190098. https://doi.org/10.1590/1678-4685-GMB-2019-0098
Brown WL, Wilson JR EO (1956) Character displacement. Syst Zool 49–64. https://doi.org/10.2307/2411924
Calanca P, Holzkämper A (2010) Agrarmeteorologische Bedingungen Im Schweizer Mittelland Von 1864 Bis 2050. In: pp 320–325
Canty A, Ripley B (2019) boot: Bootstrap R (S-Plus) Functions. R package version 1.3–22. http://CRAN.R-project.org/package=boot
Caravaggi A, Montgomery WI, Reid N (2015) Range expansion and comparative habitat use of insular, congeneric lagomorphs: invasive European hares Lepus europaeus and endemic Irish hares Lepus timidus hibernicus. Biol Invasions 17:687–698. https://doi.org/10.1007/s10530-014-0759-1
Cardoso B, Carpio AJ, Martinez-Haro M, Beltrán‐Beck B, Alzaga V, Farelo L, Campos R, Queirós J, Melo‐Ferreira J, Alves PC, Acevedo P (2024) Eco‐physiological arguments on the functional impact of Lepus timidus mitochondrial DNA introgression in Iberian hares (Lepus granatensis). J Zool 322:281–289. https://doi.org/10.1111/jzo.13142
Cederlund G, Lemnell PA (1980) Activity recording of radio-tagged animals. Biotelemetry Patient Monit 7:206–214
Chesson J (1983) The estimation and analysis of preference and its relationship to foraging models. Ecol 64:1297–1304
R Core Team (2023) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. www.R-project.org
Costa J, Queirós J, Ballesteros F, Mucci N, Souto J, Silva E, Melo-Ferreira J, Alves PC (2023) Low genetic diversity and shallow population structure in the broom hare, Lepus castroviejoi (Lagomorpha: Leporidae). Biol J Linn Soc. https://doi.org/10.1093/biolinnean/blad080
Dixon PM (1993) The bootstrap and the Jackknife: describing the precision of ecological indices. In: Scheiner SM, Gurevitch J (eds) Design and analysis of ecological experiments. Chapman and Hall, pp 290–318
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Efford MG, Fewster RM (2013) Estimating population size by spatially explicit capture-recapture. Oikos 122:918–928. https://doi.org/10.1111/j.1600-0706.2012.20440.x
Efron B, Tibshirani RJ (1993) An introduction to the bootstrap. – monographs on statistics and applied probability 57. Chapman and Hall
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Faraway JJ (2006) Extending the linear model with R. texts in statistical science. Chapman and Hall/CRC
Ferreira CM, Sabino-Marques H, Barbosa S, Costa P, Encarnação C, Alpizar-Jara R, Pita R, Beja P, Mira A, Searle JB, Paupério J, Alves PC (2018) Genetic non-invasive sampling (gNIS) as a cost-effective tool for monitoring elusive small mammals. Eur J Wildl Res 64. https://doi.org/10.1007/s10344-018-1188-8
Gidoin C, Roques L, Boivin T (2015) Linking niche theory to ecological impacts of successful invaders: insights from resource fluctuation-specialist herbivore interactions. J Anim Ecol 84:396–406
Gotelli NJ, Hart EM, Ellison AM (2015) Ecosimr-Alpha. Zenodo
Hackländer K (2022) European Hare Lepus europaeus Pallas, 1778. In: Hackländer K, Zachos FE (eds) Handbook of the mammals of Europe. Springer International Publishing, Cham, pp 1–36
Hardin G (1960) The competitive exclusion principle. Science 131:1292–1297. https://doi.org/10.1126/science.131.3409.1292
Holley AJF (2001) The daily activity period of the brown hare (Lepus europaeus). Mamm Biol 66:357–364
Holmes RT, Robinson SK (1988) Spatial patterns, foraging tactics, and diets of ground-foraging birds in a northern hardwoods forest. Wilson Bull 100:377–394
Hutchinson GE (1959) Homage to Santa Rosalia or why are there so many kinds of animals? Am Nat 93:145–159. https://doi.org/10.1086/282070
IPCC (2014) In: Field C, Barros V, Dokken D, Mach K, Mastrandrea M, Bilir T, Chatterjee M, Ebi K, Estrada Y, Genova R, Girma B, Kissel E, Levy A, MacCracken S, Mastrandrea P, White L (eds) Climate change 2014: impacts, adaptation and vulnerability. Contribution of working group II to the fifth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA
IPCC (2021) In: Masson-Delmotte V, Zhai P, Pirani A, Connors SL, Péan C, Berger S, Caud N, Chen Y, Goldfarb L, Gomis MI, Huang M, Leitzell K, Lonnoy E, Matthews JB, Maycock TK, Waterfield T, Yelekçi O, Yu R, Zhou B (eds) Climate change 2021: the physical science basis. Contribution of working group I to the sixth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge, UK, New York, NY
Jones OR, Wang J (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555. https://doi.org/10.1111/j.1755-0998.2009.02787.x
Jones AG, Small CM, Paczolt KA, Ratterman NL (2010) A practical guide to methods of parentage analysis. Mol Ecol Resour 10:6–30. https://doi.org/10.1111/j.1755-0998.2009.02778.x
Krebs CJ (1989) Measurement of dietary preferences. In: Wilson MW, Pisano S (eds) Ecological methodology. HarperCollins, New York, pp 392–407
Laikre L, Schwartz MK, Waples RS, Ryman N (2010) Compromising genetic diversity in the wild: unmonitored large-scale release of plants and animals. Trends Ecol Evol 25:520–529. https://doi.org/10.1016/j.tree.2010.06.013
Lincoln G (1974) Reproduction and March madness in the Brown hare, Lepus europaeus. J Zool 174:1–14. https://doi.org/10.1111/j.1469-7998.1974.tb03140.x
Manly B, Miller P, Cook L (1972) Analysis of a selective predation experiment. Am Nat 106:719–736
Marques JP, Ferreira MS, Farelo L, Callahan CM, Hackländer K, Jenny H, Montgomery WI, Reid N, Good JM, Alves PC, Melo-Ferreira J (2017) Mountain hare transcriptome and diagnostic markers as resources to monitor hybridization with European hares. Sci Data 4:170178. https://doi.org/10.1038/sdata.2017.178
Melo-Ferreira J, Boursot P, Suchentrunk F, Ferrand N, Alves PC (2005) Invasion from the cold past: extensive introgression of mountain hare (Lepus timidus) mitochondrial DNA into three other hare species in northern Iberia. Mol Ecol 14:2459–2464. https://doi.org/10.1111/j.1365-294x.2005.02599.x
Melo-Ferreira J, Alves PC, Freitas H, Ferrand N, Boursot P (2009) The genomic legacy from the extinct Lepus timidus to the three hare species of Iberia: contrast between mtDNA, sex chromosomes and autosomes. Mol Ecol 18:2643–2658. https://doi.org/10.1111/j.1365-294x.2009.04221.x
Nielsen EE, Bach LA, Kotlicki P (2006) Hybridlab (version 1.0): a program for generating simulated hybrids from population samples. Mol Ecol Notes 6:971–973. https://doi.org/10.1111/j.1471-8286.2006.01433.x
Oksanen A, Simpson G, Blanchet F, Kindt R, Legendre P, Minchin P, O’Hara R, Solymos P, Stevens M, Szoecs E, Wagner H, Barbour M, Bedward M, Bolker B (2022) vegan: Community Ecology Package. https://CRAN.R-project.org/package=vegan
Peakall R, Smouse PE (2012) GenAIEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research–an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pépin D, Cargnelutti B (1994) Individual variations of daily activity patterns in radiotracked European hares during winter. Acta Theriol 39:399–409
Pianka ER (1973) The structure of lizard communities. Annu Rev Ecol Syst 4:53–74. https://doi.org/10.1146/annurev.es.04.110173.000413
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Queirós J, Gortázar C, Alves PC (2020) Deciphering Anthropogenic effects on the genetic background of the red deer in the Iberian Peninsula. Front Ecol Evol 8. https://doi.org/10.3389/fevo.2020.00147
Rakotondranary SJ, Hapke A, Ganzhorn JU (2011) Distribution and morphological variation of Microcebus spp. along an environmental gradient in southeastern Madagascar. Int J Primatol 32:1037–1057. https://doi.org/10.1007/s10764-011-9521-z
Rehnus M, Bollmann K (2016) Non-invasive genetic population density estimation of mountain hares (Lepus timidus) in the Alps: systematic or opportunistic sampling? Eur J Wildl Res 62:737–747. https://doi.org/10.1007/s10344-016-1053-6
Reid N, Hughes MF, Hynes RA, Montgomery WI, Prodöhl PA (2022) Bidirectional hybridisation and introgression between introduced European brown hare, Lepus europaeus and the endemic Irish hare, L. Timidus Hibernicus. Conserv Genet 23:1053–1062. https://doi.org/10.1007/s10592-022-01471-5
Sabino-Marques H, Ferreira CM, Paupério J, Costa P, Barbosa S, Encarnação C, Alpizar-Jara R, Alves PC, Searle JB, Mira A, Beja P, Pita R (2018) Combining genetic non-invasive sampling with spatially explicit capture-recapture models for density estimation of a patchily distributed small mammal. Eur J Wildl Res 64. https://doi.org/10.1007/s10344-018-1206-x
Sakai AK, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA, Baughman S, Cabin RJ, Cohen JE, Ellstrand NC, McCauley DE, O’Neil P, Parker IM, Thompson JN, Weller SG (2001) The Population Biology of Invasive species. Annu Rev Ecol Syst 32:305–332. https://doi.org/10.1146/annurev.ecolsys.32.081501.114037
Schai-Braun SC, Hackländer K (2014) Home range use by the European hare (Lepus europaeus) in a structurally diverse agricultural landscape analysed at a fine temporal scale. Acta Theriol 59:277–287. https://doi.org/10.1007/s13364-013-0162-9
Schai-Braun S, Hackländer K (2016) Family Leporidae. Hares and rabbits. In: Wilson D, Lacher T, Mittermeier R, JR. (eds) Handbook of the mammals of the world: volume 6. Lagomorphs and rodents I. Lynx Editions, Barcelona, pp 62–148
Schai-Braun S, Rödel H, Hackländer K (2012) The influence of daylight regime on diurnal locomotor activity patterns of the European hare (Lepus europaeus) during summer. Mamm Biol 77:434–440. https://doi.org/10.1016/j.mambio.2012.07.004
Schai-Braun SC, Weber D, Hackländer K (2013) Spring and autumn habitat preferences of active European hares (Lepus europaeus) in an agricultural area with low hare density. Eur J Wildl Res 59:387–397. https://doi.org/10.1007/s10344-012-0684-5
Schai-Braun SC, Lapin K, Bernhardt K-G, Alves PC, Hackländer K (2020) Effect of landscape type, elevation, vegetation period, and taxonomic plant identification level on diet preferences of Alpine mountain hares (Lepus timidus varronis). Eur J Wildl Res 66. https://doi.org/10.1007/s10344-020-01398-7
Schai-Braun SC, Jenny H, Ruf T, Hackländer K (2021) Temperature increase and frost decrease driving upslope elevational range shifts in Alpine grouse and hares. Glob Change Biol 27:6602–6614. https://doi.org/10.1111/gcb.15909
Schai-Braun SC, Schwienbacher S, Smith S, Hackländer K (2023) Coexistence of European hares and Alpine mountain hares in the Alps: what drives the occurrence and frequency of their hybrids? J Zool. https://doi.org/10.1111/jzo.13067
Schwartz MK, Luikart G, Waples RS (2007) Genetic monitoring as a promising tool for conservation and management. Trends Ecol Evol 22:25–33. https://doi.org/10.1016/j.tree.2006.08.009
Smith RK, Jennings NV, Robinson A, Harris S (2004) Conservation of European hares Lepus europaeus in Britain: is increasing habitat heterogeneity in farmland the answer? J Appl Ecol 41:1092–1102. https://doi.org/10.1111/j.0021-8901.2004.00976.x
Spellerberg IF, Fedor PJ (2003) A tribute to Claude Shannon (1916–2001) and a plea for more rigorous use of species richness, species diversity and the ‘Shannon-Wiener’ Index. Glob Ecol Biogeogr 12:177–179. https://doi.org/10.1046/j.1466-822X.2003.00015.x
Thorén S, Quietzsch F, Schwochow D, Sehen L, Meusel C, Meares K, Radespiel U (2011) Seasonal changes in feeding ecology and activity patterns of two sympatric mouse lemur species, the gray mouse lemur (Microcebus murinus) and the golden-brown mouse lemur (M. ravelobensis), in northwestern Madagascar. Int J Primatol 32:566–586. https://doi.org/10.1007/s10764-010-9488-1
Thulin C-G (2003) The distribution of mountain hares Lepus timidus in Europe: a challenge from brown hares L. Europaeus? Mamm Rev 33:29–42. https://doi.org/10.1046/j.1365-2907.2003.00008.x
Thulin C-G, Tegelström H (2002) Biased geographical distribution of mitochondrial DNA that passed the species barrier from mountain hares to brown hares (genus Lepus): an effect of genetic incompatibility and mating behaviour? J Zool (London) 258:299–306. https://doi.org/10.1017/s0952836902001425
Tilman D (1987) The importance of the mechanisms of interspecific competition. Am Nat 129:769–774
Wallace RK (1981) An Assessment of Diet-Overlap indexes. Trans Am Fish Soc 110:72–76. https://doi.org/10.1577/1548-8659(1981)110%3C;72:AAODI%3E;2.0.CO;2
Zachos FE, Slimen HB, Hackländer K, Giacometti M, Suchentrunk F (2010) Regional genetic in situ differentiation despite phylogenetic heterogeneity in Alpine mountain hares. J Zool (London) 282:47–53. https://doi.org/10.1111/j.1469-7998.2010.00710.x
Acknowledgements
We are grateful to Tobias Schiller, Lukas Scharinger, and Greta Oberhofer for their help with data collection. We thank the hunters of Grison as licence holders and the municipalities of Scuol and Valsot for their cooperation, and Urs Monstein for his valuable support.
Funding
The study was funded by the following foundations or organisations: Stiftung Temperatio, Graf Fabrice, von Gundlach und Payne Smith-Stiftung, Verein Grünes Kreuz, Legat Dr. Joachim de Giacomi, Basler Stiftung für biologische Forschung, Swiss CIC Delegation, Amt für Jagd und Fischerei Graubünden, Swiss National Park, Institute for Wildlife Biology and Game Management, BOKU.
Open access funding provided by University of Natural Resources and Life Sciences Vienna (BOKU).
Author information
Authors and Affiliations
Contributions
SS and KH conceived the ideas and designed methodology. SS and NC collected data. SS, JQ, and PCA analysed the data. SS led the writing of the manuscript. SS, NC, FF, HJ, JQ, PCA, and KH contributed critically to the drafts and gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1: Appendix S1
. STRUCTURE diagram based on the analysis of the hare faecal samples’ microsatellite composition assuming K = 2 clusters (species level). The diagram includes the reference dataset of each hare species. Each column symbolises a hare faecal or tissue (i.e., reference dataset) sample. Each colour corresponds to the genetic proportion of a given species. Red corresponds to the Alpine mountain hare, blue to the European hare. Faecal samples (n = 137) were collected in the years 2019–2022 in the Alps in Grisons (Switzerland)
Supplementary Material 2: Appendix S2
. List of sample ID, x-coordinate, y-coordinate, transect, sampling date, season, elevation, species ID (mtDNA), species ID (SMCX), species ID (Albumina), species ID (microsatellites), species consensus/hybridisation, and individual ID for the faecal samples collected in the years 2019–2022 in the Alps in Grisons (Switzerland)
Supplementary Material 3: Appendix S3
. Chesson’s electivity indices in (a) European hare faeces (n = 165), (b) hybrid faeces (n = 109), and (c) Alpine mountain hare faeces (n = 149; species determined by mtDNA/nDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat variables of the land use and ground cover map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat variable). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Supplementary Material 4: Appendix S4
. Microsatellite composition and results of the STRUCTURE analysis of faecal samples (n = 137) and the reference dataset of each hare species assuming K = 2 clusters (species level). Faecal samples were collected in the years 2019–2022 in the Alps in Grisons (Switzerland)
Supplementary Material 5: Appendix S5
. Recapture frequency of individuals genotyped by microsatellites. Faecal samples were collected in the years 2019–2022 in the Alps in Grisons (Switzerland). European hare is shown in dark grey, Alpine mountain hare in white, and hybrids in light grey
Supplementary Material 6: Appendix S6
. Chesson’s electivity indices in (a) European hare faeces (n = 318) and (b) Alpine mountain hare faeces (n = 761; species determined by mtDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat variables of the land use and ground cover map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat variable). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Supplementary Material 7: Appendix S7
. Elevation range and number of faecal pellet locations collected in the years 2019–2022 in the Alps in Grisons (Switzerland) separated according to mtDNA and mtDNA/nDNA analysis
Supplementary Material 8: Appendix S8
. Chesson’s electivity indices in (a) European hare faeces (n = 165), (b) hybrid faeces (n = 109), and (c) Alpine mountain hare faeces (n = 149; species determined by mtDNA/nDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat types of the habitat type map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Supplementary Material 9: Appendix S9
. Post-hoc test results of the Shannon Wiener indices (parameter estimates β and p-values) of the (a) habitat type, (b) habitat type rough categorisation, and (c) land use and ground cover map using the Tukey’s all-pair comparisons method. Indices were calculated for each faecal pellet collected in the years 2019–2022 in the Alps in Grisons (Switzerland) separated by group (i.e., mtDNA, mtDNA/nDNA, individuals)
Supplementary Material 10: Appendix S10
. Chesson’s electivity indices in (a) European hare faeces (n = 318) and (b) Alpine mountain hare faeces (n = 761; species determined by mtDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat types of the habitat type map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Supplementary Material 11: Appendix S11
. Chesson’s electivity indices in (a) European hare faeces (n = 165), (b) hybrid faeces (n = 109), and (c) Alpine mountain hare faeces (n = 149; species determined by mtDNA/nDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat type categories of the habitat type rough categorisation map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type category). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Supplementary Material 12: Appendix S12
. Chesson’s electivity indices in (a) European hare faeces (n = 318) and (b) Alpine mountain hare faeces (n = 761; species determined by mtDNA) and their distributions of 1000 bootstrap resamples (mean and 95% confidence interval) for all habitat type categories of the habitat type rough categorisation map which were selected by n ≥ 6 hares (sample size in brackets is the number of hares selecting the respective habitat type category). Faecal samples were collected in Grisons (Switzerland) during the years 2019–2022. Non-significant results cross the vertical line at zero
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schai-Braun, S.C., Cybulska, N., Filli, F. et al. Competition between sympatric hare species in the Alps is boostered by climate change and hybridisation. Eur J Wildl Res 70, 83 (2024). https://doi.org/10.1007/s10344-024-01830-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-024-01830-2