Abstract
Human-predator conflicts are frequently caused by livestock and/or game depredation. The golden jackal’s (Canis aureus) range expansion in Europe, as well as the recent re-expansion of several gray wolf (Canis lupus) populations, might increase risk of such conflicts. In Austria, golden jackal presence has been increasing since the 1990s including reports of wildlife and livestock kills, frequently occurring in the provinces Styria and Burgenland. We developed a rapid, two-step genetic screening protocol to (1) detect canid mtDNA from non-invasively collected samples like swabs from kills using diagnostic PCRs, and (2) assign this DNA to red fox (Vulpes vulpes), golden jackal, or gray wolf/dog. To monitor golden jackal presence in the region, a total of 167 signs of presence were collected over a period of 30 months throughout the Styrian province. Among these, 14 non-invasive genetic samples (13 swabs from kill sites and one scat) were screened with the developed protocol. Four of these samples revealed golden jackal mtDNA and six samples red fox mtDNA. The developed genetic screening protocol represents a quick and inexpensive method to assess canid presence, e.g., at kill sites, and therefore possesses high value for the conservation and wildlife management community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Development of diagnostic PCRs for golden jackal, red fox, gray wolf, and domestic dog
Molecular genetic–based protocols can provide a valuable tool for predator identification at kill sites (Harms et al. 2015) and can support human-wildlife conflict mitigation (Sundqvist et al. 2008). Screening protocols for predator identification from kill sites require molecular markers that occur in high copy numbers, such as the mitochondrial Cytochrome Oxidase Subunit I (or COI) gene.
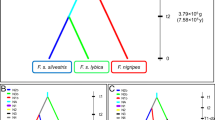
To date, four wild canids occur in Central Europe: red fox (Vulpes vulpes), gray wolf (Canis lupus), golden jackal (Canis aureus), and raccoon dog (Nyctereutes procyonoides) (IUCN 2020). The raccoon dog largely feeds on small vertebrates, invertebrates, and plant material, while the diets of the remaining three species include larger mammals such as European roe deer (Capreolus capreolus) or domestic animals (Demeter and Spassov 1993). The recent range expansion of the golden jackal and the re-expansion of the wolf are likely to lead to increased human-wildlife conflicts that require genetic monitoring (Trouwborst et al. 2015; Stronen et al. 2020). Livestock and game depredation is a major cause of such conflicts and skewed predator perception might lead to misguided mitigation measures (Chase Grey et al. 2017). Negative attitudes like fear or distrust of wolves or jackals might further skew biased predator perception (Ericsson and Heberlein 2003; Chavez et al. 2005). Therefore, ungulates and/or livestock might be an overestimated part of the jackal’s diet (McShane and Grettenberger 1984; Markov 2012), while red foxes might actively predate on ungulates more frequently than generally assumed (Liberg et al. 1993; Lindström et al. 1994).
Molecular-based species identification protocols have been developed for canids (e.g., Khatoon et al. 2019; Kheidorova et al. 2018; Stronen et al. 2020) but these methods mostly rely on DNA sequencing, or target nuclear markers such as the Zfy gene or microsatellites, making the protocols less suited for degraded samples. Species-specific PCRs only require a conventional PCR and visualization step, representing a cost-effective alternative to DNA sequencing (Thalinger et al. 2016).
We developed a rapid screening protocol to detect mtDNA (targeting the COI gene) of red fox, golden jackal, and gray wolf or dog from canid scats or swabs of kill sites, consisting of two separate PCR steps. The first step targets canid mtDNA (using primer pair CanCOI—to confirm successful isolation of canid DNA from kill sites). The second step involves species-specific PCRs to selectively amplify (a) red fox, (b) golden jackal, and (c) wolf/dog (primer pairs VvCOI, CaCOI, and ClCOI, respectively) to confirm target taxon presence at the investigated sites (Table 1). Due to the phylogenetic position of domestic dog (Canis lupus familiaris) nested within the gray wolf lineage, we were not able to design reliable primers to distinguish these two taxa.
For in silico primer design and evaluation, sequences (3 to 7 per species) of the mtDNA COI fragment were downloaded from GenBank for the following carnivores, as well as relevant wild and domestic prey species: red fox, golden jackal, gray wolf, domestic dog, raccoon dog, Eurasian lynx (Lynx lynx), European wildcat (Felis silvestris), domestic cat (Felis catus), red deer (Cervus elaphus), European roe deer, brown hare (Lepus europaeus), European rabbit (Oryctolagus cuniculus), domestic goat (Capra hircus), domestic sheep (Ovis aries), cattle (Bos taurus), and domestic pig (Sus scrofa domesticus) (Table S1, Online Resource 1).
Primer design was conducted using the R package DECIPHER (Wright 2015) in R 3.6.0 (R Core Team 2018), as well as MEGA 10.0.5 (Kumar et al. 2018), targeting autapomorphic sites for target taxa at the primer’s 3′ end, maximizing additional primer mismatches in non-target species, and aiming for amplicon sizes between 50 and 200 bp. Selected primers (Table 1) were checked for annealing temperature and potential secondary structure and dimer formations with Multiple Primer Analyzer (Thermo Fisher Scientific).
Primer pair specificity was validated in silico using PrimerBlast (Ye et al. 2012), using GenBank’s nucleotide database, and in vitro using tissue samples of four golden jackals, three red foxes, two wolves, and one dog, as well as seven target prey species (two samples each of European rabbit, roe deer, and red deer, and one sample each of domestic pig, goat, sheep, and cattle). Optimal annealing temperatures were determined via gradient PCRs (see Online Resource 1).
Final PCR reactions (volume 10 μl) contained 2 μl DNA template, 2 μl Phusion HF Buffer, 500 nM of forward and reverse primer, respectively, 200 μM of each dNTP, and 0.5U of Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific). Final PCR cycling conditions were 98 °C for 30 s (1 cycle); 98 °C for 10 s, 15 s at primer-specific annealing temperature (Table 1), 72 °C for 15 s (40 cycles); and 72 °C for 10 min (1 cycle). PCR products were visualized on a 2% agarose gel stained with PeqGREEN (VWR). At least two replicates per assay and sample are recommended when using degraded DNA in order to compensate for false negatives due to stochastic PCR dropout.
Application of developed protocol to identify potential golden jackal kills
The golden jackal’s range expansion throughout Europe since early twentieth century has led to intense, ongoing political debates about its native status in many European countries (Trouwborst et al. 2015). In Austria, golden jackal was first recorded in 1987 (Hoi-Leitner and Kraus 1989) with records steadily increasing until today (Plass 2007; Hatlauf and Hackländer 2016). The spread of this mesocarnivore may lead to increased human-wildlife conflict, as seen with other predators such as wolf, bear, and lynx (Breitenmoser 1998), as it might be perceived as a competitor or even threat to prey species. From April 2018 to October 2020, reports of potential golden jackal presence were collected throughout the Austrian province of Styria (16,401 km2) and classified in accordance to the SCALP (Status and Conservation of the Alpine Lynx Population) criteria. A total of 167 reports were collected (58 C1, 55 C2, 54 C3, Fig. 1; Online Resource 2), including the first clear documentation of golden jackal reproduction in Styria. At 14 of these records, genetic samples (13 swabs from bite marks on prey carcasses and one scat sample) were taken. Swab sampling was performed by thoroughly wiping the surface of clear bite marks on prey carcasses with a Buccal Collection Brush of the Gentra PureGene Kit (Qiagen). The brush’s head was immediately transferred into a microcentrifuge tube containing the cell lysis solution of the kit and stored at ambient temperature until arrival at the laboratory. The scat sample was collected in a clean tube and kept cool until arrival at the laboratory. Once in the laboratory, all samples were stored at – 20 °C.
DNA was extracted using either Gentra PureGene Cell or QIAamp Fast DNA Stool Mini Kits (Qiagen) following the kits’ guidelines and screened with our protocol. For four swab samples, amplification with both CanCOI and CaCOI primers confirmed the presence of golden jackal mtDNA. Seven (6 swabs, 1 scat) samples showed amplification with CanCOI and VvCOI. The three remaining swab samples showed no amplification with any primer pair. Sanger sequencing of the first seven successful amplifications confirmed the result of the diagnostic PCRs (GenBank sequences MT680188 – MT680194); therefore, Sanger sequencing was omitted for the remaining samples.
Assuming canid presence at all 13 sites where swab samples were taken, the success rate of 74% (10 of 13 swab samples) is relatively high. All samples were taken from bite marks of medium- and large-sized prey, and the only non-canid predator that could take down prey of this size, the Eurasian lynx, has not been reported in the area in the last 30 years. Therefore, the failure to obtain amplicons in the remaining three samples is assumed to be due to unsuccessful application of the protocol. The success of a non-invasive genetic screening protocol depends on (1) PCR efficiency, and good primer fit on the target sequence, both of which we optimized, and (2) DNA integrity of the sample, which decreases with time after sample deposition. For our samples, the timespan between kill and sampling was mostly longer than 48 h, suggesting degraded DNA as the main cause of PCR failure.
One limitation of mitochondrial genetic markers is that they only depict the maternal lineage preventing the identification of hybrids. Three hybrids between golden jackals and domestic dogs have been reported from Croatia (Galov et al. 2015) and jackals might also hybridize with wolves (Stronen et al. 2020). To date, no records of such hybrids have been reported from Austria. Feral or stray dogs are much rarer in Austria than in Croatia, rendering the possibility of a hybrid occurring in our dataset unlikely.
Our genetic screening protocol represents a quick and low-cost diagnostic tool allowing the efficient processing of scattered, individual samples delivered from stakeholders outside academia. However, batch-processing and the application of multiplex real-time PCR might allow for higher detection sensitivity and faster throughput when working with a larger number of samples. With our protocol, we received trustworthy results with short turnaround times. Therefore, this diagnostic PCR protocol could be of high value for the conservation and wildlife management community. In our study, over 50% of the screened samples taken from presumed golden jackal kill sites revealed red fox mtDNA. Our data indicate a potentially skewed perception of the extent of the golden jackal’s role in the currently reported prey kills in Styria and suggest that the red fox is a potential predator as well as the primary canine scavenger in the area.
Availability of data and material
All created DNA sequences have been submitted to the NCBI GenBank database (Accession Numbers MT680184 – MT680194).
Code availability
N/A.
References
Breitenmoser U (1998) Large predators in the Alps: the fall and rise of man’s competitors. Biol Conserv 83:279–289. https://doi.org/10.1016/S0006-3207(97)00084-0
Chase Grey JN, Bell S, Hill RA (2017) Leopard diets and landowner perceptions of human wildlife conflict in the Soutpansberg Mountains, South Africa. J Nat Conserv 37:56–65. https://doi.org/10.1016/j.jnc.2017.03.002
Chavez AS, Gese EM, Krannich RS (2005) Attitudes of rural landowners toward wolves in northwestern Minnesota. Wildl Soc Bull 33:517–527. https://doi.org/10.2193/0091-7648(2005)33[517:AORLTW]2.0.CO;2
Demeter A, Spassov N (1993) Canis aureus L. 1758-Schakal, Goldschakal. In: Niethammer J, Krapp F (eds) Handbuch Der Säugetiere Europas; Band 5/I Raubsäuger (Teil I). AULA-Verlag, Wiesbaden, pp 107–138
Ericsson G, Heberlein TA (2003) Attitudes of hunters, locals, and the general public in Sweden now that the wolves are back. Biol Conserv 111:149–159. https://doi.org/10.1016/S0006-3207(02)00258-6
Galov A, Fabbri E, Caniglia R et al (2015) First evidence of hybridization between golden jackal (Canis aureus) and domestic dog (Canis familiaris) as revealed by genetic markers. R Soc Open Sci 2. https://doi.org/10.1098/rsos.150450
Harms V, Nowak C, Carl S, Munoz-Fuentes V (2015) Experimental evaluation of genetic predator identification from saliva traces on wildlife kills. J Mammal 96:138–143. https://doi.org/10.1093/jmammal/gyu014
Hatlauf J, Hackländer K (2016) Current status of a spreading meso-carnivore in Austria, the golden jackal (Canis aureus). Mamm Biol 81:3–18
Hoi-Leitner M, Kraus E (1989) Der Goldschakal, Canis aureus (Linnaeus 1758), in Österreich (Mammalia austriaca 17). Bonn zool Beitr 40:197–204
IUCN (2020) The IUCN Red List of Threatened Species. Version 2020-3. http://www.iucnredlist.org. Accessed 2 Jan 2020
Khatoon R, Mehmood T, Anwar M, Habiba U, Eggert LS, Gompper ME (2019) A field and laboratory-based assessment of the distribution of large- and meso-carnivore species in the newly established Murree, Kotli Sattian, and Kahuta National Park, Pakistan. MAMMAL Res 64:411–422. https://doi.org/10.1007/s13364-019-00428-3
Kheidorova EE, Shpak VA, Homel VK et al (2018) Molecular genetic identification of the invasive species - Asian Jackal (Canis aureus) in the territory of Belarus. Dokl NATSIONALNOI Akad Nauk BELARUSI 62:86–92. https://doi.org/10.29235/1561-8323-2018-62-1-86-92
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Liberg O, Johansson A, Lockowandt S, Wahlström LK (1993) Red fox predation as a dominant cause of neonatal mortality in roe deer fawns. In: Swedusg Gzbters’ Association Conference. Lund, Sweden, pp 12–13
Lindström E, Andrén H, Angelstam P et al (1994) Disease reveals the predator: sarcoptic mange, red fox predation, and prey populations. Ecology 75:1042–1049. https://doi.org/10.2307/1939428
Markov G (2012) Golden jackal (Canis aureus L.) in Bulgaria: what is going on. Acta Zool Bulg Suppl 4:67–71
McShane TO, Grettenberger JF (1984) Food of the golden jackal (Canis aureus) in central Niger. Afr J Ecol 22:49–53. https://doi.org/10.1111/j.1365-2028.1984.tb00673.x
Plass J (2007) Dokumentation einer zweiten Einwanderungswelle des Goldschakals Canis aureus Linnaeus 1758 in Österreich aus den Jahren 2003-2006. Beitr Naturk Oberösterreichs 17:55–68
Stronen AV, Bartol M, Boljte B et al (2020) “Passive surveillance” across species with cross-amplifying molecular markers: the potential of wolf (Canis lupus) genetic monitoring in tracking golden jackal (C. aureus) colonization and hybridization. Hystrix It J Mammal 31:74–76. https://doi.org/10.4404/hystrix-00259-2019
Sundqvist A-K, Ellegren H, Vila C (2008) Wolf or dog? Genetic identification of predators from saliva collected around bite wounds’ on prey. Conserv Genet 9:1275–1279. https://doi.org/10.1007/s10592-007-9454-4
R Core Team (2018) R: A language and environment for statistical computing
Thalinger B, Oehm J, Mayr H, Obwexer A, Zeisler C, Traugott M (2016) Molecular prey identification in Central European piscivores. Mol Ecol Resour 16(1):123–137
Trouwborst A, Krofel M, Linnell JDC (2015) Legal implications of range expansions in a terrestrial carnivore: the case of the golden jackal (Canis aureus) in Europe. Biodivers Conserv 24:2593–2610. https://doi.org/10.1007/s10531-015-0948-y
Wright ES (2015) DECIPHER: harnessing local sequence context to improve protein multiple sequence alignment. BMC Bioinformatics 16:322. https://doi.org/10.1186/s12859-015-0749-z
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13:134
Acknowledgements
We thank Stephan Koblmüller and Lukas Zangl for providing mammal tissue samples from the Austrian bar-coding of life (ABOL) collection. We are further grateful to the Steirische Landesjägerschaft (Styrian hunting association) for funding initial data collection, and the Styrian hunters who contributed to the collection of the golden jackal records.
Funding
Open access funding provided by University of Graz. M.K. was funded by the Styrian hunting association (Steirische Landesjägerschaft) to collect the majority of the raw data of this project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
N/A.
Consent to participate
N/A.
Consent for publication
N/A.
Conflict of interest
M.K. was funded by the Styrian hunting association (Steirische Landesjägerschaft) to collect the majority of the raw data of this project and is currently employed by the Styrian chamber of agriculture (Landwirtschaftskammer Steiermark).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schenekar, T., Karrer, M., Karner, I. et al. Non-invasive diagnostic PCRs for rapid detection of golden jackal, red fox, and gray wolf/domestic dog and application to validate golden jackal presence in Styria, Austria. Eur J Wildl Res 67, 42 (2021). https://doi.org/10.1007/s10344-021-01488-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-021-01488-0