Abstract
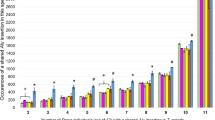
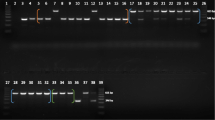
Young Alu elements that have been inserted recently in macaque genomes are still polymorphic for insertion presence/absence at the individual and population levels. They have been considered as a powerful tool in population genetics of primates. Combining a computational approach and a PCR display method based on primers of OWM-specific Alu subfamilies, in the present study, we identified and characterized 26 novel Alu insertions that are polymorphic in the Tibetan macaque (Macaca thibetana). Along with 20 previously reported loci, a total of 96 Alu insertions were found to be present in the Tibetan macaque but absent in rhesus macaques (Macaca mulatta) as validated by computational methods. After PCR verification, 27.1 % (26/96) of the analyzed insertions were confirmed to be heterozygous among Tibetan macaque individuals. These 26 newly identified loci were further genotyped in 41 individuals from two Tibetan macaque populations. The average allele frequencies across the 26 loci varied from 2.4 to 48.8 % in the two populations. The average expected heterozygosity per locus ranged from 25.3 to 50.6 %. The results indicate that the combination of a computational approach and a PCR display method is an efficient way to identify young polymorphic Alu insertions in macaques, and the newly identified polymorphic loci represent valuable genetic markers for the study of Tibetan macaque population genetics in the future.

Similar content being viewed by others
References
Batzer MA, Deininger PL (1991) A human-specific subfamily of Alu sequences. Genomics 9:481–487
Batzer MA, Deininger PL (2002) Alu repeats and human genomic diversity. Nat Rev Genet 3:370–379
Batzer MA, Rubin CM, Hellmann-Blumberg U, Alegria-Hartman M, Leeflang EP, Stern JD, Bazan HA, Shaikh TM, Deininger PL, Schmid CW (1995) Dispersion and insertion polymorphism in two small subfamilies of recently amplified human Alu repeats. J Mol Biol 247:418–427
Bennett EA, Coleman LE, Tsui C, Pittard WS, Devine SE (2004) Natural genetic variation caused by transposable elements in humans. Genetics 168:933–951
Cordaux R, Lee J, Dinoso L, Batzer MA (2006) Recently integrated Alu retrotransposons are essentially neutral residents of the human genome. Gene 373:138–144
Cordaux R, Srikanta D, Lee J, Stoneking M, Batzer MA (2007) In search of polymorphic Alu insertions with restricted geographic distributions. Genomics 90:154–158
Groves CP, Wilson DE, Reeder DM (2005) Order primate. Mammal species of the world: a taxonomic and geographic reference [M], vol 1, 3rd edn. Johns Hopkins University Press, Maryland, pp 111–184
Hamdi H, Nishio H, Zielinski R, Dugaiczyk A (1999) Origin and phylogenetic distribution of Alu DNA repeats: irreversible events in the evolution of primates. J Mol Biol 289:861–871
Han K, Konkel MK, Xing J, Meyer TJ, Huang CT, Sandifer E, Hebert K, Barnes EW (2007) Mobile DNA in old world monkeys: a glimpse through the rhesus macaque genome. Science 316:238–240
Jia X, Yang B, Yue B, Yin H, Wang H, Zhang X (2011) Isolation and characterization of twenty-one polymorphic microsatellite loci in the Tibetan macaque (Macaca thibetana). Russ J Genet 47:884–887. doi:10.1134/S1022795411070088
Jiang X, Wang Y, Wang Q (1996) Taxonomy and distribution of Tibetan macaque (Macaca thibetana). Zool Res 17:361–369
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment
Konkel MK, Wang JX, Liang P, Batzer MA (2007) Identification and characterization of novel polymorphic LINE–1 insertions through comparison of two human genome sequence assemblies. Gene 390:28–38
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Li D, Fan L, Yin H, Wang H, Wu S, Yue B (2008) Genetic diversity analysis of Macaca thibetana based on mitochondrial DNA control region sequences. Mitochondrial DNA 19:446–452
Li J, Han K, Xing JC, Kim HS, Rogers J, Ryder OA, Disotell T, Yue B, Batzer MA (2009) Phylogeny of the macaques (Cercopithecidae): Macaca based on Alu elements. Gene 448:242–249
Liu Y, Li J, Zhao J (2005) Sequence variation of mitochondrial DNA control region and population genetic diversity of Tibetan macaques Macaca thibetana in the Huangshan Mountain. Acta Zool Sin 52:724–730
Munroe DJ, Haas M, Bric E, Whitton T, Aburatani H, Hunter K, Ward D, Housman DE (1994) IRE-bubble PCR: a rapid method for efficient and representative amplification of human genomic DNA sequences from complex sources. Genomics 19:506–514
Nikaido M, Rooney AP, Okada N (1999) Phylogenetic relationships among cetartiodactyls based on insertions of short and long interpersed elements: hippopotamuses are the closest extant relatives of whales. Proc Natl Acad Sci U S A 96:10261–10266
Okada N, Shedlock AM, Nikaido M (2004) Retroposon mapping in molecular systematics. Methods Mol Biol 260:189–226
Ray DA, Walker JA, Hall A, Llewellyn B, Ballantyne J, Christian AT, Turteltaub K, Batzer MA (2005) Inference of human geographic origins using Alu insertion polymorphisms. Forensic Sci Int 153:117–124
Raymond M, Rousset F (2002) GENEPOP version 3.3 (online). Available at http://wbiomed.curtin.edu.au/genepop assignment. Mol Ecol 16:1099–1106
Roos C, Schmitz J, Zischler H (2004) Primate jumping genes elucidate strepsirrhine phylogeny. Proc Natl Acad Sci 101:10650–10654
Roy AM, Carroll ML, Kass DH, Nguyen SV, Salem AH, Batzer MA, Deininger PL (1999) Recently integrated human Alu repeats: finding needles in the haystack. Genetica 107:149–161
Rozen S, Skaletsky HJ (1998) Primer3 code available at http://www-genome.wi.mit.edu/genome_software/other/primer3.html
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Lab, New York
Schmitz J, Roos C, Zischler H (2005) Primate phylogeny: molecular evidence from retroposons. Cytogenet Genome Res 108:26–37
Stoneking M, Fontius JJ, Clifford SL, Soodyall H, Arcot SS, Saha N, Jenkins T, Tahir MA, Deininger PL, Batzer MA (1997) Alu insertion polymorphisms and human evolution: evidence for a larger population size in Africa. Genome Res 7:1061–1071
Sun B, Li J, Zhu Y, Xia D (2010) Mitochondrial DNA variation in Tibetan macaque (Macaca thibetana). Folia Zool 59:301–307
Wang J, Song L, Gonder MK, Azrak S, Ray DA, Batzer MA, Tishkoff SA, Liang P (2006a) Whole genome computational comparative genomics: a fruitful approach for ascertaining Alu insertion polymorphisms. Gene 365:11–20
Wang J, Song L, Grover D, Azrak S, Batzer MA, Liang P (2006b) dbRIP: a highly integrated database of retrotransposon insertion polymorphisms in humans. Hum Mutat 27:323–329
Watkins WS, Rogers AR, Ostler CT, Wooding S, Bamshad MJ, Brassington AME, Carroll ML, Nguyen SV, Walker JA, Prasad BVR (2003) Genetic variation among world populations: inferences from 100 Alu insertion polymorphisms. Genome Res 13:1607–1618
Xing J, Wang H, Han K, Ray DA, Huang CH, Chemnick LG, Stewart CB (2005) A mobile element based phylogeny of old world monkeys. Mol Phylogenet Evol 37:872–880
Zhong L, Zhang M, Yao Y, Ni Qing MJ, Li Ch XH (2013) Genetic diversity of two Tibetan macaque (Macaca thibetana) populations from Guizhou and Yunnan in China based on mitochondrial DNA D-loop sequences. Genes Genom 35:205–214
Acknowledgments
We are grateful to Prof. Xu Huailiang in Sichuan Agricultural University for providing samples, Prof. Timothy Moermond in Sichuan University for English revision. This research was funded by the National Natural Science Foundation of China (No. 31270431), this Project Sponsored by the Scientific Research Foundation for the Returned Overseas Scholars, State Education Ministry (20111568-8-3).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Scandura
Huawei Guo and Juan Jiang contributed equally to the work.
Rights and permissions
About this article
Cite this article
Guo, H., Jiang, J., Cui, Y. et al. Identification and characterization of polymorphic Alu insertions in the Tibetan macaque (Macaca thibetana). Eur J Wildl Res 61, 143–149 (2015). https://doi.org/10.1007/s10344-014-0887-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10344-014-0887-z