Abstract
Due to its productivity and potential to adapt to the expected climate change, the Douglas-fir is one of the most important commercial non-native forest tree species in Europe. Currently, seeds from both non-native European and native American seed stands are used for plantations. In this study, we investigate European seed lots for their native origin (variety and potential geographic origin in America) and assess the adaptability, growth and survival potential of European versus American Douglas-fir seed lots. We compare the genetic diversity, morphological characteristics such as height (h), root collar diameter (rcd) and the ratio of h/rcd, and the timing of bud burst. We investigate 852 1-year-old seedlings from 10 different US and European seed lots representing 5 provenance regions which are sold in Germany and Austria. Seedlings are genotyped for 13 nuclear SSRs and analysed together with reference data set and standard genetic structuring and assignment methods. Adaptive traits of morphological characteristics and timing of bud burst of the seedlings are recorded and statistically analysed. The results show that the investigated European seedlings originate from recommended American native seed sources and represent both varieties and inter-varietal admixed individuals. European seedlings have a lower genetic diversity versus the American seedlings and native populations. They show significant differences in the adaptive traits such as morphological characteristics and timing of bud burst. According to the genetic diversity indices, certified North American Douglas-fir seed sources should be preferred for planting in Central Europe.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Douglas-fir (Pseudotsuga menziesii (Mirb.) Franco) is the second most cultivated non-native conifer tree species in Europe after the Sitka spruce (Köble and Seufert 2001), which grows in more than 35 European countries. Within its native distribution range, in western North America, two distinct varieties of the Douglas-fir are known: the coastal variety (P. menziesii var. menziesii) and the interior variety (P. menziesii var. glauca). The coastal variety grows along the Pacific coast and the west facing slopes of the Rocky Mountain range from British Columbia, Canada to California, USA. The interior variety (also called Rocky Mountain variety) grows further east of British Columbia, across the Rocky Mountains to New Mexico (USA) (Eckenwalder 2009). In the contact zone of the two varieties, hybridization is evident (Gugger et al. 2010; van Loo et al. 2015; Wei et al. 2011).
The complex glacial and postglacial history during the Quaternary period accompanied by the adaptation to very different ecological conditions led to specific patterns of genetic diversity and a large genetic variation in adaptive traits (e.g. bud burst, bud set, growth performance, fall and spring frost hardiness) within the native distribution range (Gould et al. 2012; St. Clair 2006; St. Clair et al. 2005).
In Europe, the current distribution of the Douglas-fir is the result of a long introduction history, which started in 1826 (Köble and Seufert 2001). In most European countries, the first Douglas-fir stands were characterized by excellent growth and health conditions. At the beginning of the twentieth century, further imports of Douglas-fir seeds to Germany were less successful. This initiated the first provenance tests in Europe, which confirmed the importance of seed origin for cultivation (Bastien et al. 2013), since it affects the growth, frost sensitivity and tolerance to diseases. As a result, the recommended provenances for Western and Central Europe are the coastal variety from the Western Cascades and the coastal region in Washington as well as Oregon (Barner 1973; Kleinschmit and Bastien 1992). The interior variety is not as productive as the coastal Douglas-fir and has a higher susceptibility to needle cast (Rhabdocline pseudotsugae) (Eilmann et al. 2013). Hence, the interior variety is generally not recommended for cultivation in Europe (Boyle 1999). Only in continental environments (e.g. Sweden, Finland and former Czechoslovakia) does the interior variety from British Columbia perform better than the coastal variety (Kleinschmit and Bastien 1992).
Although European Douglas-fir stands planted prior to the 1980s are of unknown geographic seed origin (Bastien et al. 2013), several of these stands were selected and certified for harvesting and trading forest reproductive material of the Douglas-fir (seeds, parts of plants, planting stocks). The underlying assumption was that Douglas-fir is appropriate for the ecological conditions in Europe. Since these stands have largely not been tested for their genetic origin, an artificial mixture of both varieties and different native origins cannot be excluded (Bastien et al. 2013; Klumpp 1999; Konnert and Ruetz 2006). Such mixtures can influence the genetic diversity in natural regeneration and in seeds, which may decrease due to inbreeding and assortative mating (Fussi et al. 2013; Konnert and Ruetz 2006). Furthermore, the European seed stands are rather small in size (less than 1 ha) and often show a patchy tree distribution, which negatively influences the population structure and mating patterns (Ratnam et al. 2014). In the native distribution range, large Douglas-fir stands ensure a wide-spread population with an extensive pollen gene flow. Today, Douglas-fir seedlings from European stands as well as from native forest stands in North America are planted.
In Europe, two legal frameworks for the trading of forest reproductive material (seeds, parts of plants, planting stocks) are in place: (1) the European Council Directive 1999/105/EC for Members of the European Union; and (2) the regulation of the Organization for Economic Co-operation and Development (OECD) open for all countries who wish to participate (Ackzell 2002). The EU directive differentiates between four categories (1) “source identified”, (2) “selected”, (3) “qualified” and (4) “tested” according to the basic material, e.g. seed sources, seed stands and seed orchards. Each European member state can enforce its own more rigorous rules for the approval of basic material and production of FRM (forest reproductive material) (Konnert et al. 2015). At an international level, the OECD scheme defines the same four categories (OECD 2007).
With the increasing interest in cultivating Douglas-fir as an adaptation option to climate change in Europe, planting suitable FRM is of increasing importance. Thus, assessing the differences in the characteristics between American versus European FRM is important. Commonly, the genetic diversity is seen as a measure of the adaptive potential (Adams et al. 1998; Konnert and Fussi 2012) of the tree population to changing growing conditions. Demographic processes (e.g. bottlenecks, founder events) caused by forest management practices such as selective thinning followed by natural or artificial regeneration may shape the genetic composition of the European seedlings leading to a reduced genetic variation (Ratnam et al. 2014). Adaptive traits are reflected in the morphology and phenology of a plant. The morphological characteristics (1) shoot height and (2) root collar diameter of seedlings are considered as an indicator for the growth and survival potential (Haase 2007; Landis et al. 2010). The timing of bud burst is an important adaptive parameter, as young shoots of the Douglas-fir are highly susceptible to late-frost damages caused by late-spring frost temperatures (Steiner 1979).
In this study, we test European seed sources for their native origin (variety and potential geographic origin in America) and investigate the adaptation, survival and growth potential of European versus American Douglas-fir seed sources. We use cultivated one-year-old seedlings from 10 different American and European seed lots to assess (1) the native origin and genetic diversity of seedlings of a given seed lot, (2) morphological characteristics, such as height, diameter as well as sturdiness (height/diameter) as an indicator for seedling quality and (3) the timing of bud burst, which is an adaptive trait and reflects susceptibility to late frost. Here, we combine two different approaches: firstly population genetics to determine the native origin and genetic diversity and secondly the assessment of adaptive traits to elucidate the growing and survival potential.
Materials and methods
Seedlings
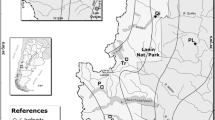
The seedlings for our study are provided by an Austrian company producing containerized forest seedlings. The commercial available seed lots come from 10 different seed stands (S01–S10) (Table 1) and represent 5 provenance regions: two seed stands are located in Austria, three in Germany and five in the US (Fig. 1). Each seedling population consists of 65–98 plants (Table 1). The seed lots are selected according to the demand in Austria and Germany. Since Darrington is currently the most demanded provenance, three seed sources are selected (B. Igler, personal communication, 31 May 2015). The European seed stands grow in the Waldviertel (Austria) and in the Südostdeutsches Berg- und Hügelland (Germany) and belong to the FRM (forest reproductive material) category “selected”. According to national legislation, the minimum number of harvested trees for this category requires 10 individuals within a given seed stand in Austria (Müller and Strohschneider 2004) and 20 in Germany (Behm et al. 2007).
The American seed stands are Darrington, Trout Lake and Randle and belong to the category either “selected” or “source identified” (Table 1). In the native range, the seed stands are harvested with the help of Douglas squirrels (Tamiasciurus douglasi), which collect and store Douglas-fir cones in holes. These cones are collected within the harvesting area which can encompass several hundred hectares (Konnert and Ruetz 2011).
The category “source identified” defines FRM as material from a seed stand located within a single region of provenance, where little or no phenotypic selection has taken place. The FRM category “selected” defines material from a phenotypically selected stand (Council Directive 1999/105/EC). In Austria, both categories are common; in Germany, only the category “selected” (see § 13 FoVG) (Forstvermehrungsgutgesetz (FoVG) 2002) is permitted.
Experimental set-up
In April 2012, the commercial seeds were germinated in pots with a volume of 50 cm3 and the components of the soil substrate were white peat with fraction 0–5 mm (90%) and horticultural pearlite with fraction 0–3 mm (10%). After germination, the seedlings were separated and in December 2012 transported to the laboratory of the Institute of Silviculture in Vienna, Austria. Leaf samples of 30 (10 small, 10 medium and 10 large) seedlings per seed lot (in total 300 samples) were collected, dried and stored in silica gel prior to the DNA extraction. The morphological characteristics height and root collar diameter of each individual seedling were recorded.
The seedlings were further cultivated in the same pots with a distance of 3 cm to each other as a randomized block design and placed at the experimental forest garden “Knödelhütte”. The garden is located in the western part of Vienna at an elevation of 290 m above sea level (N 48°13′ E 16°14′) with a mean annual temperature of 10.3 °C and a mean annual precipitation rate of 603 mm. Three different stages (1) dormant, (2) axillary (lateral) bud burst completed and (3) terminal bud burst completed (when green needles were first visible) were recorded to assess the bud burst development (Fig. 2). The bud stages were recorded every second week from 20 February to 15 April 2013, until the first bud was flushing. From 15 April, the stages were recorded on a daily basis at 9 am until terminal bud burst of each seedling was completed.
DNA extraction and nuSSR genotyping
Total DNA from 25 g of powdered leaf samples per individual was extracted applying the commercial OMEGA E.Z.N.A Plant DNA Kit (OMEGA Bio-Tek, Inc., Norcross, Georgia, USA) according to the manufacturer´s instructions. The extracted DNA samples were amplified and genotyped with 13 nuSSRs. The 13 nuSSRs are identical to those used by van Loo et al. (2015). Details on the PCR procedure are given in van Loo et al. (2015).
Analyses and results
Variety composition and potential native origin
We started our analysis by assessing the variety (coastal or interior) and the potential geographic origin of the seed lots in Northwest America. Both the variety composition and the potential native origin were assessed using multilocus genotype data of the studied seedlings and the reference data set developed by van Loo et al. (2015). Genotypes were not corrected for null alleles. The reference data set represents genotypes of 746 individual Douglas-fir trees from 36 reference populations and covers the natural distribution range in Northwestern America (Figure S1 in the supplementary materials). The reference populations were genotyped with identical nuSSRs to those used in our study.
The variety composition was assessed by applying the software STRUCTURE v.2.3.4, which uses a Bayesian clustering approach by applying the Markov chain Monte Carlo (MCMC) algorithm to allocate individuals to clusters (K) that are genetically similar (Falush et al. 2003, 2007; Pritchard et al. 2000). We used the multilocus data of the reference data set to pre-define 2 clusters (K = 2), representing the two varieties, and probabilistically assigned all individual seedlings to these clusters. Twenty independent runs were applied, one for each cluster (K) with a burn-in period of 50,000, followed by a run length of 100,000 iterations. As recommended by Falush et al. (2003), we used an admixture model that allows the seedlings to have admixed ancestries, permitting it to detect inter-varietal admixed individuals. In addition, we used correlated allele frequencies, as we expected similar frequencies due to the common ancestors of the two varieties. For each independent run, the software STRUCTURE estimates a membership coefficient (Q), which corresponds to the probability of an individual belonging to each cluster. An individual was declared as coastal with Q > 0.90, as Rocky Mountain with Q < 0.10 and inter-varietal admixed with 0.90 > Q > 0.10. To find the optimal alignment of the 20 independent runs, the software CLUMPP v.1.1.2. was used. The estimated individual membership coefficients (Q) were averaged using the greedy algorithm in CLUMPP to correct for discrepancies between runs (Jakobsson and Rosenberg 2007). Then, the average Q values were plotted using DISTRUCT v.1.1 (Rosenberg 2004).
The potential geographic origin of the studied European seedlings in Northwestern America was estimated using genetic assignments of the software GeneClass2 (Piry et al. 2004). Although we knew the origin of the American seedlings, we also applied the analyses of these seedlings to test the accuracy of the applied assignment methodology. Seedlings of all seed stands (here S01-S10) were assigned to the reference populations of the reference data set. Three types of methods for likelihood estimation are available in GeneClass2. In our study, we applied two of them: (1) the frequency-based method after Paetkau et al. (1995) and (2) the distance-based method using Nei’s (1972) standard genetic distance-based criteria according to Takezaki and Nei (1996). The frequency method after Paetkau et al. (1995) assigns an individual or group of individuals to the reference population with the highest likelihood (score) according to its multilocus genotypes and the allele frequencies (Hauser et al. 2006). We applied a threshold likelihood value for assignment of p < 0.05. The distance-based method (Nei 1972; Takezaki and Nei 1996) assigns an individual or group of individuals to the reference population according to the smallest genetic distance (Piry et al. 2004). We use Nei’s (1972) standard genetic distance D S to calculate a rank with the corresponding scores (%) to all reference populations. Commonly, frequency-based methods are more powerful than distance-based methods (Hauser et al. 2006). The distance-based methods tend to be less sensitive to violations of the Hardy–Weinberg Equilibrium (HWE) (Hauser et al. 2006), which could be the case in European populations as they might originate from small, fragmented but also cultivated variety mixed stands. This suggests using both methods.
All seedlings of the stands S03, S04, S05, S07, S08, S09 and S10 were assigned to the coastal variety (Fig. 3). For the Austrian stand S02, three of 30 individuals were assigned to the interior variety and four individuals were allotted to be inter-varietal admixed seedlings. The remaining 23 seedlings were assigned to the coastal variety. For population assignments, S02 was analysed with 30 individuals to test if it is derived from the admixture zone in the natural distribution range. Furthermore, S02 was separated into the 2 varieties (coastal and interior Douglas-fir) to investigate if it was established by two different seed sources. One inter-varietal admixed individual was detected in the US stand S06 and the Austrian stand S01 (Fig. 3).
STRUCTURE results plotted using DISTRUCT v.1.1 for genetic structure (K = 2) representing 746 individuals of the 36 reference populations (marked by R) and 300 individuals of the 10 seed source populations (marked by S) of the coastal (green colour) and Rocky Mountain variety (blue colour). Individuals are grouped by populations. (Color figure online)
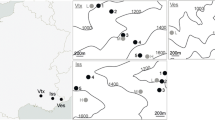
The assignment results for the potential native origin are given in Table 2 and Fig. 4. The frequency-based assignment method of Paetkau et al. (1995) showed score values of correctly assigned populations of 100% in almost all studied populations. Accordingly, seedlings of the European stands (S01, S02, S03, S05) matched the reference populations (R) of the coastal variety R11, R15, R16 and R11 in the Western Cascades in Washington, respectively. The seedlings of S04 were allocated to R32, a coastal variety from Vancouver Island in British Columbia, Canada. Three seedlings of the Rocky Mountain variety of S02 referred to R18 in British Columbia. Results of S02 showed that the population with 30 variety mixed and admixed individuals, and if separated by variety (27 coastal individuals), was allotted to the same reference populations (Table 2). Hence, we conclude that S02 includes two different seed sources and does not originate from the admixture zone. The seedlings of the native US populations were assigned to reference populations in the Western Cascades in Washington and Oregon with the following results: S06–S08 (Darrington) to R15, the seedlings of S09 (Randle) to R11 and those from S10 (Trout Lake) to R05.
Illustration of the assignment results according to the frequency-based approach after Paetkau et al. (1995). Geographic locations of the US populations (S06–S10) and the reference populations are marked with crosses and circles, respectively
When using the second method based on Nei’s distance approach, the same reference populations for S01, S03 and S05-S10 were identified. However, the populations S02 and S04 were assigned to R30/R27 and R30, respectively. The likelihood score values, a measure for the probability of a studied population belonging to the reference population, ranged between 5.2 and 11.1%. We built 10 small subsamples with 3 randomly selected individuals of the reference population R18 and R27 to analyse the reliability of group assignments with 3 interior individuals for S02, (Table 3). The subsamples were assigned to reference populations following identical steps as for the other assignments. The randomly selected individuals were left out in the corresponding reference population. Results showed that the subsamples (1–10) were correctly assigned to R27 and R18 (Table 3).
Genetic diversity
Using the software GenAlEx v. 6.5 (Peakall and Smouse 2006, 2012), we assessed the genetic diversity of the seedlings using the following diversity parameters: (1) allelic diversity (Na), (2) effective number of alleles (Ne), (3) observed frequency of heterozygotes (Ho), (4) expected frequency of heterozygotes (He) and the (5) inbreeding coefficient (Fis). Fis values were tested for significance (p 0.05) of their deviation from the HWE (Hardy–Weinberg Equilibrium), also using the software GENEPOP v.4.1.4 (Raymond and Rousset 1995; Rousset 2008). We further calculated the allelic diversity parameters (6) allelic richness (As) and (7) private allelic richness As(p) (the number of distinct alleles private (specific) to a population based on a standardized population size by rarefaction using ADZE v. 1.0) to allow a comparison of populations with different sample sizes (Szpiech et al. 2008). We applied a standardized population size of 8 individuals, which is the same as used in the reference data set (van Loo et al. 2015).
The procedure allowed us to compare the allelic richness of the European seedlings with their corresponding native reference population since it accounts for differences in the sample size. If a population consists of a mixture of both varieties (as was the case with the Austrian S02), the genetic diversity indices by variety were calculated as well.
An independent t test was applied (with PASW Statistics for Windows Version 18.0) to evaluate the differences between the mean of the two independent groups. We compared the genetic diversity parameters between seedlings from the US and the European stands and the allelic richness parameter (As 8) of the seedlings from European stands with the reference populations from van Loo et al. (2015).
The results for the calculated genetic diversity indices (Na, Ne, Ho, He, Fis, As 8 and As 8 (p)) of the European and American seedlings are given in Table 2. For S02, these indices were additionally calculated for the coastal Douglas-fir variety. The t test revealed that Na (p = 0.021), Ne (p = 0.001), H E (p = 0.023) and As 8 (p = 0.015) were significantly higher within the US populations versus the European populations. Other diversity parameters (Ho, Fis and As 8 (p)) of the European seedlings were within the range of the American seedlings. The Fis were significantly positive, indicating deviations from HWE. This could confirm previously published data on the Douglas-fir. The comparison with the assigned native reference populations R11, R15, R16, R32 (Table S1 in the supplementary materials) of the frequency-based approach, revealed that the allelic diversity parameter As 8 of the European seedlings was significantly lower (p = 0.008).
Morphological characteristics
For the statistical analyses of the morphological characteristics, the PASW Statistics for Windows Version 18.0 was used. The Levene test was applied to verify the assumptions of equal variances. For equal variances, an analysis of variance (one-way ANOVA) using the post hoc test Scheffé for multiple comparisons was used at a significance level of α = 0.05. If the assumption of homogeneity of variances was violated, a one-way ANOVA with Welch’s F-test using the post hoc test Games-Howell for multiple comparisons of unequal variances was performed at a significance level of α = 0.05.
Height
The adjusted Welch ANOVA determined a statistically significant difference between seedlings of the 10 seed populations (p < 0.001). Multiple comparisons (see Table 4; Fig. 5) of the studied populations indicated a significantly higher growth performance of the Austrian seedlings from S02 (µ 16.9 cm) when compared to seedlings from all other studied seed populations, followed by the seedlings of the US population S09 (µ 15.7 cm). The seedlings from the US population S10 (Fig. 6) showed the lowest growth performance with a mean value of 11.5 cm (highly significant). The seedlings from the remaining populations from Europe (S01, S03–S05) and the USA (S06–S08) indicated similar height growth rates ranging from 13.1 to 14.1 cm.
Root collar diameter
The one-way ANOVA determined a statistically significant difference between seedlings of the 10 seed populations (p < 0.001) of the morphological feature root collar diameter (range 1.8 to 2.2 mm) (Table 4; Fig. 5). Again the seedlings of the Austrian population S02 showed the highest growth in diameter (µ 2.2 mm) compared to seedlings from all other populations (significantly, except S01 and S09). The seedlings of the US population S07 revealed the lowest mean diameter at 1.8 mm. The German population S03 (µ 2.1 mm) indicated a significantly better growth performance in diameter compared to the US populations S06, S07 and S10.
Height/diameter
The adjusted Welch ANOVA determined a statistically significant difference between the seedlings of the 10 stands (p < 0.001). Multiple comparisons (see Table 4; Fig. 5) indicated the highest ratio in height to diameter in seedlings from the US population S09 (µ 8.4), which is significantly higher compared to seedlings of all other populations, except S02 (µ 7.9). The seedlings of the US population S10 showed the lowest mean value of the sturdiness quotient at 6.3, followed by S03 at 6.6. In all other populations, similar values ranging between 7.0 and 7.4 were found.
Timing of bud burst
The investigated bud burst development stages showed that within 15 days (15.04.2013 to 29.04.2013), all seedlings completed stage number 3, when the first green needles of the terminal bud were visible. The separate examination of lateral and terminal bud burst indicated that 70% of the auxiliary buds flushed earlier than the terminal buds. Following this, the timing of the terminal buds was evaluated. The same statistical analysis, as described in “Morphological characteristics” section, was applied to compare the timing of bud burst. For estimating the effect of the population, blocks and their interaction, a general linear model (GLM) was applied. Therefore, each population was treated as fixed, and blocks as a random variable. Results of the GLM analysis showed that the main effect of population is significant (p 0.015) (see Table 5). Random block effects and population times block interactions were not significant (p 0.167 and 0.234, respectively) (Table 5). The timing of the terminal buds was further evaluated according to Welch’s one-way ANOVA and the statistically significant difference between seedlings of the 10 seed populations was determined (p < 0.001). The post hoc test Games-Howell for multiple comparisons was used to determine which specific seed lots differ. Results of the multiple comparisons showed the earliest bud burst of seedlings from the European stands S01 and S04 with a mean of 8.3 days, which is significantly earlier compared to seedlings from the German population S05 (µ 9.8 days) and the US population S08 (µ 10.1 days) (Table 6; Fig. 7). Bud flushing within the native US populations (S06–S10) ranged between 8.6 and 10.1 days (Table 6). Multiple comparisons showed a significantly earlier timing of bud burst of S09 (µ 8.6 days) as compared to S08 (µ 10.1 days) (Table 6; Fig. 7). In all other populations, no significant differences were found with mean values ranging between 8.9 and 9.6 days.
Discussion
The assignment tests of the European stands (see Table 2; Fig. 4) showed that they come from the recommended seed zones for Douglas-fir cultivation in Austria and Germany (Kleinschmit et al. 1979; Weißenbacher 2008). However, the assignment results also show that forest stands from the European provenances “Waldviertel” (S01, S02) and “Südostdeutsches Berg und Hügelland” (S03, S04, S05) come from various native provenances (Table 2; Fig. 4). We also found that the Austrian stand (S02) contained a mixture of coastal, interior and inter-varietal admixed seedlings (Fig. 3). This is crucial since the interior variety is not recommended for planting due to its lower growth performance and higher susceptibility to fungi. In Bavaria (Germany), this problem has been recognized and certified seed stands must be genetically tested to ensure that no seed collections come from mixed variety stands (Konnert and Fussi 2012). These genetic tests are planned for other German states (BLE 2016) and should be extended on an European level to ensure the best growing potential.
By assigning the native US seed lots to the reference populations, the performance of the applied assignment methods revealed the validity of the procedure since seed lots were correctly referenced to populations of similar geographic locations in the USA (Fig. 4). Nevertheless, further testing of the European planting material is suggested because (1) our analyses addressed the population level only and not the individual level and (2) all genetic analyses were restricted to 30 individuals. Analyses of larger sets might also reveal a larger number of interior variety and inter-varietal admixed individuals, and (3) European FRM might originate from variety mixed stands or be influenced by gene flow from neighbouring Douglas-fir stands.
The genetic diversity indices of Na (p = 0.021), Ne (p = 0.001), H E (p = 0.023) and As 8 (p = 0.015) were significantly higher within the US populations versus the European populations (Table 2). The private allelic richness (As(p)) of the European seed lots revealed overlapping values compared to American seed lots indicating distinct alleles specific to each population. In the European population S02, the highest As(p) value was found (Table 2), most probable as a result of the detected interior and inter-varietal admixed individuals which account for the distinct alleles. The comparison of the assigned native reference populations R11, R15, R16, R32 (Table S1 in the supplementary materials) with the European seed lots showed significantly higher values of allelic richness (As 8 ) (p = 0.008). The allelic richness is an estimator for the genetic diversity within populations and may decline for smaller populations (bottleneck or founder effect) (El Mousadik and Petit 1996). This indicates that the European seed lots come from stands with assortative mating, where the small sized, and sometimes patchily distributed seed stands can limit gene flow or have been collected from a small number of trees.
The seedling height can be related to the photosynthetic capacity and the transpiration, leading to a competitive advantage of taller seedlings within the shrub layer, but decreasing the survival rate due to drought stress and/or wind damage (Haase 2007). A larger root collar diameter suggests a larger root system, which is important for tree growth (Haase 2007). Blake et al. (1989) reported increased survival rates of Douglas-fir seedlings and a correlation with increased stem diameter. According to Rose and Ketchum (2003), a difference of 2 mm leads to an increase in stem volume of 35–43% after 4 years.
Although we could only obtain data from one growing season, the height and root collar diameter suggest that the best growing potential was evident for the Austrian population S02 (Table 4; Fig. 5), while the seedlings from the US population S10 (Fig. 6) showed the lowest height growth performance of all the investigated populations (p < 0.05). The ratio of height to diameter suggests that seedlings with high ratios are more susceptible to wind, drought and frost damages (Haase 2007). Thus, to reduce the risk, the h/d ratio should range between 5.5 and 7.5 for planted Douglas-fir plants (1.5–2 years old) (Bauer et al. 2009). The highest h/d ratio was evident for the US population S09 (µ 8.4) and was significantly higher versus all other populations except S02 (p < 0.05) (Table 4; Fig. 5). According to Bauer et al. (2009), the US population S09 (µ 8.4) and the European population S02 (µ 7.9), have a higher susceptibility to damages after planting.
Summing up, the European population S02 showed the best growth performance (see Table 4; Fig. 5), but the h/d ratio of 7.9 indicates a higher risk of damages after planting. The US populations S10 and S07 showed the lowest growth performance in terms of height and root collar diameter, respectively. The highest risk for damages after planting is evident for the US population S09.
Selecting the best seed source reduces the risk of frost damage due to the high heritability of bud burst traits by approximately 80% (Janßen and Rau 2008). Multiple comparisons showed that the European populations (S01, S04) exhibited a significantly earlier timing of bud burst (Table 6) as compared to the European population (S05) and the native US population (S08). Within the native range, population S09 exhibited a significantly earlier timing of bud burst as compared to S08 (Table 6). Hence, the European seed sources S01 and S04 and the US seed source S09 might be more susceptible to frost damages. The high range of bud burst within the native populations (µ 8.6 and 10.1 days) confirms the results of a previous study by St. Clair et al. (2015), which also showed a high variation in bud burst and early bud flushing related to latitude and summer drought of coastal Douglas-fir.
Conclusion
European provenance regions consist of different native origins and represent both varieties of Douglas-fir. The genetic diversity of the European seed lots was lower in comparison with seed lots from America and native populations. According to these results, we recommend native FRM from certified North American Douglas-fir seed sources. Provenances from coastal British Columbia, the Western Cascade Mountains and Oregon with a mean annual temperature ranging from 6 to 9.5 °C are predicted to be the most productive for Central Europe under both current and future climate conditions (Chakraborty et al. 2016). They are also characterized by the highest genetic diversity for the Douglas-fir (Table S1, Figure S1 in the supplementary materials), with a peak in Northern Oregon and Southern Washington west of the Cascades and decreasing trends towards south and north of the distribution range (Li and Adams 1989; Neophytou et al. 2016; van Loo et al. 2015).
When using European FRM, only FRM of known variety and native origin should be planted. Hence, we strongly advise to continue testing European seed stands of unknown origin for variety composition and potential native origin. This would be a valuable improvement in FRM certificates provided by/for seed and forest managers and forest owners in Europe.
References
Ackzell L (2002) On the doorstep to new legislation on forest reproductive material—policy framework and legislation on trade with forest reproductive material. For Genet Resour 30:52
Adams WT, Zuo J, Shimizu JY, Tappeiner JC (1998) Impact of alternative regeneration methods on genetic diversity in coastal Douglas-fir. For Sci 44:390–396
Barner H (1973) Procurement of Douglas-fir seed for provenance research. In: Proceedings of IUFRO working party on Douglas-Fir provenance, Göttingen, West Germany, pp 82–89
Bastien J, Sanchez L, Michaud D (2013) Douglas-Fir (Pseudotsuga menziesii (Mirb.) Franco). In: Pâques L (ed) Forest tree breeding in Europe: current state-of-the-art. Springer, Dodrecht, pp 325–373
Bauer W, Braun W, Braunger M, Dörr T, Ebinger T, Göckel C, Karopka M, Mann P, Morell M, Schmid R, Thumm H, Wieners M (2009) Pflanzgut und Pflanzung. ForstBWPraxis, Ministerium für Ernährung und Ländlichen Raum, Stuttgart
Behm A, Bußler H, Gmelin-Bösselmann S, Kirchner C, Luckas M, Rank K, Reichert A, Salder K, Schirmer R (2007) Forstliches Vermehrungsgut Für Bayern: Gesetzliche Bestimmungen, Verwaltungsvorschriften, Herkunftsempfehlungen, Merkblätter und Hinweise für die Praxis, 4th edn. Bayerische Forstverwaltung, Teisendorf
Blake JI, Teeter LD, South DB (1989) Analysis of the economic benefits from increasing uniformity in Douglas-fir nursery stock. In: Mason W, Deans JD, Thompson S (eds) Producing uniform conifer planting stock. Oxford University Press, Oxford, pp 252–261
BLE (2016) Forstvermehrungsgutrecht: Empfehlungen des gemeinsamen Gutachterausschusses (gGA) der Länder für dessen Umsetzung. Bundesanstalt für Landwirtschaft und Ernährung
Boyle JR (1999) Introduction to planted forests: contributions to the quest for sustainable societies. In: Boyle JR, Winjum JK, Kavanagh K, Jensen EC (eds) Plantet forests: contributions to the quest for sustainable societies. Kluwer Academic Publishers, Dodrecht
Chakraborty D, Wang T, Andre K, Konnert M, Lexer MJ, Matulla C, Weißenbacher L, Schüler S (2016) Adapting Douglas-fir forestry in Central Europe: evaluation, application, and uncertainty analysis of a genetically based model. Eur J For Res 135:919–936. doi:10.1007/s10342-016-0984-5
Eckenwalder JE (2009) Conifers of the world. Timber Press, Oregon
Eilmann B, de Vries SMG, den Ouden J, Mohren GMJ, Sauren P, Sass-Klaassen U (2013) Origin matters! difference in drought tolerance and productivity of coastal Douglas-fir (Pseudotsuga menziesii (Mirb.)) provenances. For Ecol Manag 302:133–143. doi:10.1016/j.foreco.2013.03.031
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree [Argania spinosa (L.) Skeels] endemic to Morocco. Theor Appl Genet 92:832–839. doi:10.1007/BF00221895
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578. doi:10.1111/j.1471-8286.2007.01758.x
Forstvermehrungsgutgesetz (FoVG) (2002) Forstvermehrungsgutgesetz of 22 May 2002 BGBl. I S. 1658
Fussi B, Konnert M, Dounavi A (2013) Identification of varieties and gene flow in Douglas fir exemplified in artificially established stands in Germany. Ann For Res 56:249
Gould PJ, Harrington CA, Clair JBS (2012) Growth phenology of coast Douglas-fir seed sources planted in diverse environments. Tree Physiol 32:1482–1496. doi:10.1093/treephys/tps106
Gugger PF, Sugita S, Cavender-Bares J (2010) Phylogeography of Douglas-fir based on mitochondrial and chloroplast DNA sequences: testing hypotheses from the fossil record. Mol Ecol 19:1877–1897
Haase DL (2007) Morphological and physiological evaluations of seedling quality. In: Riley LE, Dumroese RK, Landis TD (eds) National proceedings: forest and conservation nursery associations-2006. USDA Forest Service, Fort Collings, pp 3–8
Hauser L, Seamons TR, Dauer M, Naish KA, Quinn TP (2006) An empirical verification of population assignment methods by marking and parentage data: hatchery and wild steelhead (Oncorhynchus mykiss) in Forks Creek, Washington, USA. Mol Ecol 15:3157–3173. doi:10.1111/j.1365-294X.2006.03017.x
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. doi:10.1093/bioinformatics/btm233
Janßen A, Rau HM (2008) Leistungssteigerung durch klassische forstliche Züchtung. Arch f Forstwes u Landsch ökol 42:127–133
Kleinschmit J, Bastien JC (1992) IUFRO’s role in Douglas-fir (Pseudotsuga menziesii (Mirb.) Franco) tree improvement. Silvae Genet 41:161–173
Kleinschmit J, Svolta J, Weisgerber H, Jestedt M, Dimpfelmeier R, Ruetz W, Dieterich H (1979) Ergebnisse aus dem internationalen Douglasien-Herkunftsversuch von 1970 in der Bundesrepublik Deutschland. Silvae Genet 28:226–244
Klumpp RT (1999) Untersuchungen zur Genökologie der Douglasie (Pseudotsuga menziesii [Mirb.] FRANCO). Dissertation, University Göttingen
Köble R, Seufert G (2001) Novel maps for forest tree species in Europe. In: Proceedings of the 8th European symposium on the physic-chemical behavior of air pollutants: “a changing atmosphere!” Torino (ITA)
Konnert M, Fussi B (2012) Natürliche und künstliche Verjüngung der Douglasie in Bayern aus genetischer Sicht. Schweiz Z Forstwes 163:79–87. doi:10.3188/szf.2012.0079
Konnert M, Ruetz W (2006) Genetic aspects of artificial regeneration of Douglas-fir (Pseudotsuga menziesii) in Bavaria. Eur J For Res 125:261–270. doi:10.1007/s10342-006-0116-8
Konnert M, Ruetz W (2011) Besuch von Erntebeständen im Ursprungsland der Douglasie. AFZ-DerWald 5:9–11
Konnert M, Fady B, An Gömöry D, A’hara S, Wolter F, Ducci F, Koskela J, Bozzano M, Maaten T, Kowalczyk J (2015) Use and transfer of forest reproductive material in Europe in the context of climate change. European Forest Genetic Resources Programme (EUFORGEN), Bioversity International, Rome, Italy. xvi and, p 75
Landis TD, Dumroese RK, Haase DL (2010) The container tree nursery manual: seedling processing, storage, and outplanting. Agriculture Handbook 674. USDA Forest Service
Li P, Adams WT (1989) Range-wide patterns of allozyme variation in Douglas-fir (Pseudotsuga menziesii). Can J For Res 19:149–161
Müller F, Strohschneider I (2004) Forstliches Vermehrungsgutgesetz—Kommentar und Anwendungshilfe. BFW Bundesamtsblätter, Vienna
Nei M (1972) Genetic distance between populations. Am Soc Nat 106:283–292
Neophytou C, Weisser AM, Landwehr D, Šeho M, Kohnle U, Ensminger I, Wildhagen H (2016) Assessing the relationship between height growth and molecular genetic variation in Douglas-fir (Pseudotsuga menziesii) provenances. Eur J For Res 135:465–481
OECD (2007) OECD scheme for the certification of forest reproductive material moving in international trade OECD forest seed and plant scheme. OECD Forest Seed and Plant Scheme, OECD Trade and Agricultural Directorate, Paris
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354. doi:10.1111/j.1365-294X.1995.tb00227.x
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539. doi:10.1093/jhered/esh074
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ratnam W, Rajora OP, Finkeldey R, Aravanopoulos F, Bouvet JM, Vaillancourt RE, Kanashiro M, Fady B, Tomita M, Vinson C (2014) Genetic effects of forest management practices: global synthesis and perspectives. For Ecol Manag 333:52–65. doi:10.1016/j.foreco.2014.06.008
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rose R, Ketchum JS (2003) Interaction of initial seedling diameter, fertilization and weed control on Douglas-fir growth over the first four years after planting. Ann For Sci 60:625–635. doi:10.1051/forest:2003055
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rousset F (2008) Genepop’007: a complete reimplementation of the Genepop software for Windows and Linux. Mol Ecol Resour 8:103–106
St. Clair JB (2006) Genetic variation in fall cold hardiness in coastal Douglas-fir in western Oregon and Washington. Can J Bot 84:1110–1121. doi:10.1139/b06-084
St. Clair JB, Mandel NL, Vance-Borland KW (2005) Genecology of Douglas Fir in Western Oregon and Washington. Ann Bot 96:1199–1214. doi:10.1093/aob/mci278
Steiner KC (1979) Variation in bud-burst timing among populations of interior Douglas-fir. Silvae Genet 28:76–79
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504. doi:10.1093/bioinformatics/btn478
Takezaki N, Nei M (1996) Genetic distances and reconstruction of phylogenetic trees from microsatellite DNA. Genetics 144:389–399
The Council of the European Union (1999) Council Directive 105/EC of 22 December 1999 on the marketing of forest reproductive material
van Loo M, Hintsteiner W, Pötzelsberger E, Schüler S, Hasenauer H (2015) Intervarietal and intravarietal genetic structure in Douglas-fir: nuclear SSRs bring novel insights into past population demographic processes, phylogeography, and intervarietal hybridization. Ecol Evol 5:1802–1817. doi:10.1002/ece3.1435
Wei XX, Beaulieu J, Damase PK, Vargas-Hernández J, López-Upton J, Jaquish B (2011) Range-wide chloroplast and mitochondrial DNA imprints reveal multiple lineages and complex biogeographic history for Douglas-fir. Tree Genet Genom 7:1025–1040
Weißenbacher L (2008) Herkunftswahl bei Douglasie—der Schlüssel für einen erfolgreichen Anbau. BFW Praxisinformation 16:3–5
Acknowledgements
Open access funding provided by Austrian Science Fund (FWF). The authors thank the LIECO GmbH for providing plant material. Financial support for this study came from (in alphabetic order): the alpS GmbH (Project: B04 AdaptAF B), the Austrian Research Promotion Agency (FFG) and Austrian Science Fund (FWF) (Project ID: P26504). We also thank the two anonymous reviewers and the journal editor for their comments in improving the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Christian Ammer.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Eckhart, T., Walcher, S., Hasenauer, H. et al. Genetic diversity and adaptive traits of European versus American Douglas-fir seedlings. Eur J Forest Res 136, 811–825 (2017). https://doi.org/10.1007/s10342-017-1072-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10342-017-1072-1