Abstract
Asobara japonica (Hymenoptera: Braconidae), Ganaspis brasiliensis and Leptopilina japonica (Hymenoptera: Figitidae) are Asian larval parasitoids of spotted wing drosophila, Drosophila suzukii (Diptera: Drosophilidae). This study evaluated these parasitoids’ capacity to attack and develop from 24 non-target drosophilid species. Results showed that all three parasitoids were able to parasitize host larvae of multiple non-target species in artificial diet; A. japonica developed from 19 tested host species, regardless of the phylogenetic position of the host species, L. japonica developed from 11 tested species; and G. brasiliensis developed from only four of the exposed species. Success rate of parasitism (i.e., the probability that an adult wasp successfully emerged from a parasitized host) by the two figitid parasitoids was low in hosts other than the three species in the melanogaster group (D. melanogaster, D. simulans, and D. suzukii). The failure of the figitids to develop in most of the tested host species appears to correspond with more frequent encapsulation of the parasitoids by the hosts. The results indicate that G. brasiliensis is the most host specific to D. suzukii, L. japonica attacks mainly species in the melanogaster group and A. japonica is a generalist, at least physiologically. Overall, the developmental time of the parasitoids increased with the host’s developmental time. The body size of female A. japonica (as a model species) was positively related to host size, and mature egg load of female wasps increased with female body size. We discuss the use of these parasitoids for classical biological control of D. suzukii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Key Message
-
Asobara japonica, Ganaspis brasiliensis and Leptopilina japonica are parasitoids of Drosophila suzukii.
-
We tested physiological abilities of these parasitoids on D. suzukii and 24 non-target drosophilid species.
-
Asobara japonica showed a broad host range.
-
Leptopilina japonica broadly attacked about half of the non-target species.
-
Ganaspis brasiliensis parasitized D. suzukii and three closely related species.
Introduction
Classical biological control is a useful strategy for invasive pest species when effective natural enemies are lacking in the invaded range (Hajek et al. 2016; Van Driesche et al. 2010). Assessing a natural enemies’ host range is a critical step in developing classical biological control programs (Hoddle et al. 2020), as insect parasitoid species may attack phylogenetically related hosts that share physiological traits or ecological niches (Desneux et al. 2012). Non-target tests of candidate agents have become increasingly important as part of a shifting paradigm in classical biological control from potential benefits to risks of releasing selected natural enemies (Heimpel and Cock 2018). Natural enemies that exhibit host species plasticity are rarely purposely released, even though this may aid in their establishment, as would natural enemies that exhibit plasticity to environmental tolerances. Here, the host range of three solitary endoparasitoids, Asobara japonica Belokobylskij (Hym.: Braconidae), Leptopilina japonica Novković & Kimura and Ganaspis brasiliensis (Ihering) (both Hym.: Figitidae), were evaluated as a part of a North American classical biological control program for the invasive Drosophila suzukii Matsumura (Dip.: Drosophilidae) (Daane et al. 2016; Giorgini et al. 2019).
Drosophila suzukii is native to East Asia but has invaded and widely established in regions of the Americas, Europe and North Africa (Asplen et al. 2015; Boughdad et al. 2021; Ferronato et al. 2019). Drosophila suzukii infests ripening fruits and is able to develop on many soft-skinned and damaged fruit crops as well as wild host plants (Kenis et al. 2016; Lee et al. 2015; Poyet et al. 2015; Stewart et al. 2014). Multiple insecticide applications are often needed to prevent economic losses to crops (Farnsworth et al. 2017; Mermer et al. 2020; Van Timmeren and Isaacs 2013), increasing management costs. Moreover, crop systems are challenged by repeated reinvasions of adult flies from non-crop or untreated-crop habitats that provide refuge for D. suzukii populations throughout the year (Hennig and Mazzi 2018; Santoiemma et al. 2019; Tonina et al. 2018; Wang et al. 2019b). Improved suppression by natural enemies could reduce fly populations in reservoir habitats before adult flies move into crop habitats, thereby reducing pest pressure (Lee et al. 2019).
Numerous parasitoid species, mostly belonging to Braconidae (Asobara), Figitidae (Ganaspis and Leptopilina), Diapriidae (Trichopria) and Pteromalidae (Pachycrepoideus), attack frugivorous drosophilids, with most species reported from flies in fallen and decaying fruits (Carton et al. 1986). In contrast, surveys of naturally occurring parasitoids attacking D. suzukii in North America or Europe have shown only low levels of parasitism, typically from pupal parasitoids (Trichopria drosophilae Perkins and Pachycrepoideus vindemiae (Rondani)) and little or no larval parasitism (reviewed by Lee et al. 2019). Larval parasitoids commonly found attacking D. melanogaster Meigen and other drosophilids in rotting fruits (e.g., Asobara tabida Nees, Leptopilina heterotoma (Thomson), and Leptopilina boulardi Barbotin) rarely develop from D. suzukii due to the fly’s immune resistance (Kacsoh and Schlenke 2012). It is likely that improved natural regulation of D. suzukii could be achieved through the introduction of more host-specific parasitoids that are not as susceptible to the fly’s immune response. Explorations for co-evolved parasitoids in South Korea, China and Japan discovered several larval parasitoids of D. suzukii (Daane et al. 2016; Giorgini et al. 2019; Girod et al. 2018a). Among them, A. japonica, G. brasiliensis and L. japonica were the dominant parasitoids, with G. brasiliensis and L. japonica predominantly or exclusively reared from D. suzukii and reaching parasitism rates > 70%.
Basic biological traits and potential competitive interactions of these three parasitoids have been evaluated in quarantine facilities in Switzerland (Girod et al. 2018b, c) and California (Biondi et al. 2017, 2021; Hougardy et al. 2019; Wang et al. 2018, 2019a, 2020, 2021). Non-target impacts were first reported on five European drosophilid species for populations of G. brasiliensis from Kunming, China and Tokyo, Japan; populations of L. japonica from Kunming and Beijing, China; and a population of A. japonica from Tokyo, Japan (Girod et al. 2018b). Prior to this work, there was evidence from field collections that A. japonica is a generalist parasitoid of drosophilids (Ideo et al. 2008; Nomano et al. 2015), whereas L. japonica appears to be an oligophagous species, observed to parasitize D. suzukii and several other Drosophila species from the melanogaster group (Kimura and Suwito 2015; Mitsui and Kimura 2010; Novkovic et al. 2011). Ganaspis brasiliensis’ host range is more difficult to describe as some populations appear to specialize on D. suzukii (Girod et al. 2018a, b; Matsuura et al. 2018), whereas other populations have broader host ranges (Buffington and Forshage 2016; Giorgini et al. 2019; Kasuya et al. 2013). The identification of G. brasiliensis populations or the existence of cryptic species will be discussed later and is the focus of several ongoing studies in Europe and North America. Reported herein is the non-target evaluation of these three larval parasitoids against 24 drosophilid species found in North America. Due to the phylogenetic and ecological diversity in the Drosophila species tested, we used no-choice assays to assess host acceptance—i.e., can the parasitoid locate the host—and host suitability—i.e., can the parasitoid attack and develop from the host. Any studies on parasitoid host searching and acceptance conducted under the constraints of the quarantine are necessarily simplistic compared with the natural environment; still, this work is critical to the selection of candidate parasitoids by understanding their possible non-target risks.
Materials and methods
Studies were conducted under controlled conditions (23 ± 2 °C, 14L: 10D (and natural light), 40–60% RH) at the University of California’s Quarantine Facility (Berkeley, CA, USA). Colonies of the target host (D. suzukii) and three larval parasitoid species (A. japonica, G. brasiliensis, and L. japonica) were maintained using methods similar to previously reported studies (Biondi et al. 2017; Wang et al. 2019a, 2020). Briefly, the D. suzukii colony was initiated from field collections of infested cherries in Parlier, CA, USA. Adult flies were held in Bug Dorm cages (BioQuip Products Inc., Rancho Dominguez, CA, USA) with 10% honey-water solution provided as food. Fly larvae were reared on a standard cornmeal and yeast diet in Petri dishes (1.5 cm high, 14.0 cm diameter) that had been exposed to adult flies for 24 h. For colony maintenance and experiments, flies that had developed into 1–2 d old larvae were used as all three parasitoids prefer younger host larvae (Wang et al. 2018, 2021).
Parasitoid colonies were initiated from field collections of D. suzukii in South Korea on wild Rubus spp. in 2014; details of collection sites, locations and the parasitoid species composition and numbers collected are presented in Daane et al. (2016). Adult parasitoids were held in clean (no diet) drosophila vials supplied with a streak of 50% honey-water until use for either colony maintenance or experiments. Drosophila suzukii was used as the rearing host for 4–5 generations prior to all tests. For parasitoid rearing, about 20 D. suzukii larvae were transferred to drosophila vials filled with 2 cm of artificial diet and then exposed to one 3–6 d old female parasitoid for 2–3 d. The larvae were then held for the emergence of adult flies (unparasitized flies emerged in 4–7 d) or parasitoids (emerged after 17–23 d, depending on the species and gender). Emerged wasps were collected daily and held in clean drosophila vials provided with 50% honey-water. The two figitid species were kept with males from adult emergence and were assumed to be mated, whereas virgin female A. japonica were used as this species has a strongly female-biased sex ratio and the tested strain was primarily parthenogenetic (Daane et al. 2016). Details on some aspects of the biology of these parasitoids, reared on D. suzukii, have been presented previously (Biondi et al. 2021; Girod et al. 2018a, c; Hougardy et al. 2019; Wang et al. 2018, 2019a, 2021).
Selection of non-target Drosophila species
There are over 2,000 described Drosophila species worldwide, consisting of two subfamilies Steganinae and Drosophilinae. Steganinae is a relatively small subfamily, whereas the Drosophilinae consists of many more genera, including Drosophila that has over 1500 described species. Most Drosophila species are non-pest insects, breeding in decaying substrates, and only a few, including D. suzukii, are considered invasive pests in North America. Moreover, only 13 Drosophila are listed as endangered species (US Fish and Wildlife Services) and these are picture-wing or pomace Drosophila in the Hawaiian Islands, and no Drosophila species are listed as threatened or endangered in the continental USA. For non-target studies, we selected 24 species from North America that represented diversity in their geographical origins, phylogenetic relationships (2 subfamilies, 7 genera, 9 subgenera, 22 species groups) and ecological niches (fruits, mushrooms, wood, flowers, sap). Table S1 shows their taxonomic positions, rearing diet, origins and feeding habits. With the exception of D. suzukii, all fly species were purchased from the University of California’s San Diego Drosophila Stock Center, and were originally collected from 11 different US states, except for a Japanese species (Scaptomyza elmoi Takada, selected as a close relative of the endangered Hawaiian Drosophilae) and a Samoan species (Samoaia leonensis Wheeler and Kambysellis). Fly species were reared for 3–8 generations in three different artificial mediums using similar methods as described above for D. suzukii. We note here that the earlier work by Girod et al. (2018b) selected five common European drosophilid species based on their phylogenetic relatedness and sympatry with D. suzukii; these five species were also used in our study. In addition, we focused on species found in North America, as required for the North American Plant Protection Organization (NAPPO) guidelines (NAPPO 2015). We thus included S. elmoi as a representative of Hawaiian drosophilids; more rare or endangered Hawaiian species were not tested because they are often difficult to obtain and/or rear, and importation to the UC Berkeley quarantine would have involved additional permits as they are non-native to North America.
The phylogenetic relationship of the tested drosophilid species was constructed based on available COI gene sequences from the NCBI database, with the exception of S. elmoi and Gitona americana that did not have available gene sequences. For these two species, a DNA extraction was performed using DNeasy Blood & Tissue Kit (Qiagen). The target CO1 gene was amplified using polymerase chain reaction (PCR), using primer pair LepF1-short/LepR1-short (LepF1-short: 5′-ATTCAACCAATCATAAAGATAT-3′ and LepR1-short: 5′-TAAACTTCTGGATGTCCAAAAA-3′). For each sample, a 1.0 µL sample DNA was mixed with 5.0 µL 5X Green GoTaq Reaction Buffer (ProMega), 0.2 µM dNTP, 0.2 µM each primer, and 0.2 µL Taq DNA Polymerase (New England BioLabs) in a total reaction volume of 25 µL. The thermal profile used was 94 °C for 1 min, 5 cycles of 94 °C for 1.5 min, 45 °C for 1.5 min, and 72 °C for 1.5 min, 35 cycles of 94 °C for 1 min, 50 °C for 1.5 min, and 72 °C for 1 min, and a final extension of 72 °C for 5 min. Following target CO1 amplification, samples were treated with ExoSAP (0.5 µL Exonuclease I, 0.5 µL Shrimp Alkaline Phosphatase, 1.0 µL 10X Exonuclease Reaction Buffer, and 5.0 µL PCR product), and run at 37 °C for 15 min, then at 80 °C for 15 min. Samples were then sequenced using an ABI 3730xl DNA Sequencer (Thermo Fisher Scientific).
Host acceptance and offspring survival
Test procedures for each host and parasitoid species combination were similar and were conducted to ascertain the innate or physiological potential of parasitoids to parasitize and survive in each host species. No-choice tests were used because innate parasitism could be masked in choice assays by a preference for higher-ranked hosts and by relative differences in host quality. For each test, twenty 2–d-old fly larvae were transferred to vials with artificial diet using a wet and soft brush. The vials were then streaked with 50% honey-water as food for the adult wasps, and a female parasitoid was placed in each vial for a 24 h period. The exposed larvae were then held for the emergence of adult flies (4–7 d) or parasitoids (17–23 d), which were recorded daily. Larvae of some Drosophila species are able to defend themselves from parasitoid eggs placed inside their bodies by surrounding the egg with blood cells that eventually melanize and form a black capsule surrounding the egg, which results in the death of the immature parasitoid by asphyxiation and is visible in developed adult flies (Chabert et al. 2012; Wang et al. 2016). Therefore, we also examined all emerged flies for the presence of a black capsule inside the fly’s abdomen. There were 30 replicates for each host species (e.g., 600 exposed hosts for each species), and 20–25 replicates of unexposed fly larvae that served as a control to estimate the natural mortality under experimental conditions. Control mortality varied from 11.1 to 47.2% among host species (F24,491 = 6.2, P < 0.001), with the three species in the melanogaster group (D. melanogaster, D. simulans Sturtevant, and D. suzukii) having the lowest natural mortality (Fig. S1).
Effect of host species on the body size and mature egg load of female Asobara japonica
Asobara japonica developed from a wider variety of host species than the other two parasitoids (see results) and, for this reason, we used it as a model to measure the effect of host species size on the body size and potential fecundity (mature egg load) of female parasitoids. A subsample of 20–25 female wasps that emerged from each of 16 different host species were measured for their hind tibia and ovipositor lengths to the nearest 0.001 mm with an ocular micrometer under a microscope and dissected for their mature egg load when the females were 5–6 d old (i.e., when mature egg load would have been highest; Wang et al. 2018, 2021). Emerged female wasps were held in vials (8 cm high, 2 cm diameter) with a streak of 20% honey water and were killed in the freezer immediately before dissection. The length l and width w of 25 individual pupae of each host species were measured, and the volume (V) of the prolate ellipsoid puparium was calculated as a proxy measurement of host body size using the formula V = 4/3π × (l/2) × (w/2)2 (Otto and Mackauer 1998).
Data analysis
Results are presented as mean ± SE. As described in Biondi et al. (2021) for assessing the impact of fly larvae in fruit versus artificial diet, the impact of host species on the development of parasitoid offspring was compared using “Successful Rate of Parasitism" (SP) to determine the probability that a parasitized host would give rise to an adult wasp. SP is calculated as ep/ (ef − efp), where ep = number of emerged parasitoids, ef = the average number of emerged flies in the absence of parasitoids (control), and efp = number of emerged flies in the presence of parasitoids. In instances when ep > (ef − efp), we set SP = 1. The proportion of hosts that were successfully parasitized was calculated using the “Degree of Parasitism” (DP), which is similar to Abbott’s or Schneider–Orelli formula used to correct treatment mortality and is calculated as (ef − efp)/ef. When ef < efp, we set ef − efp = 0. “Encapsulation rate” (ER) was calculated as ER = bc/(ef − efp + bc), where bc = the number of emerged flies bearing black capsules. The effects of host and parasitoid species, and their interaction on DP, SP or ER were analyzed using a Generalized Linear Model (GLM) with a binomial distribution and a logit link function; effects on the number of emerged wasps were analyzed using GLM with a Poisson distribution and a logit identity link function.
To determine if parasitoid performance parameters were correlated with fly phylogeny, Moran’s I, a statistical measure of spatial autocorrelation, was calculated using the phylosignal function in the phylosignal package in R version 4.0.2 (R Core Team 2020). The values for Moran’s I range from − 1 to + 1, with more positive values indicating that the tested trait is more similar between closely related taxa than expected by chance; it has the advantage of being insensitive to tree size and balance, and does not assume an evolutionary model (Münkemüller et al. 2012).
For host body size and development time, data were excluded from host species with < 5 emerged parasitoids and the remaining data were analyzed using one-way ANOVA. We were able to analyze relationships between mean development time of A. japonica and L. japonica for multiple host species, and between adult female A. japonica offspring body size and mature egg load and host size using linear regression and one-way ANOVA. Prior to analyses, proportion data (control host mortality) were arcsine square-root transformed to homogenize variances and all data were checked for the normality of residuals and homoscedasticity using Shapiro’s and Bartlett’s tests, respectively. If a significant difference was detected, pairwise comparisons were performed using Tukey HSD tests. Analyses were performed using JMP®, Pro 14 (SAS Institute Inc., Cary, NC, 1989–2019).
Results
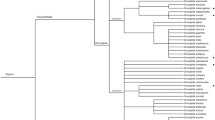
All three parasitoid species were able to parasitize D. suzukii under quarantine conditions, but the number of parasitized non-target host species varied greatly (Fig. 1). Asobara japonica developed from 19 out of 24 tested non-target species, including 16 of the 19 tested Drosophila spp. as well as species in the genera Chymomyza, Hirtodrosophila, Samoaia, Scaptodrosophila, and Scaptomyza. As mentioned, S. elmoi represented endangered Hawaiian species. Leptopilina japonica developed from 9 non-target species; except for the tested Hirtodrosophila sp., all were Drosophila spp. and most produced < 0.5 adult offspring per female per 24 h exposure period, compared with 3.2 and 3.7 from D. melanogaster and D. suzukii, respectively. For example, only 1, 2 and 3 L. japonica individuals (out of 600 exposed larvae for each species) developed from D. robusta Surtervant, D. pseudoobscura (Frolova and Astaurov) and D. subobscura Collin, respectively. Ganaspis brasiliensis developed from only three non-target species and two of these were D. melanogaster and D. simulans, both closely related to D. suzukii; only five G. brasiliensis individuals (of 600 exposed) emerged from the other non-target D. persimilis. All non-target species attacked by G. brasiliensis were also attacked by L. japonica (Fig. 1). Overall, more offspring were produced by A. japonica than by G. brasiliensis or L. japonica on D. suzukii and on the non-target hosts from which all three parasitoids emerged.
Mean (± SE) number of offspring that emerged as adult wasps after (a) A. japonica, (b) L. japonica and (c) G. brasiliensis adult females were exposed to drosophilid host larvae (exposed in diet) for a 24 h period, with host species arranged in a phylogenetic tree constructed based on COI gene sequences. Values were analyzed using a two-way analysis of variance on the effects of parasitoid species, host species and their interaction (see Table 1)
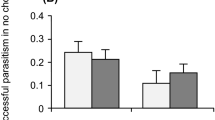
Both host and parasitoid species affected the parasitoids’ performance measurements—degree of parasitism (DP), success rate of parasitism (SP), encapsulation rate (ER) and the number of parasitoid offspring that emerged (Table 1, Fig. 2). For SP, DP and ER, the maximum rate is ‘1’ for each performance measure, but with the stacked bars shown in Fig. 2, the highest value would be ‘2’ (e.g., for 100% SP and DP, must have 0% ER, conversely 100% ER and DP must have 0% SP). Overall, DP and SP were higher for A. japonica than for L. japonica or G. brasiliensis (Fig. 2). Encapsulation of parasitoids occurred in only 5 host species for A. japonica, but in 13 host species for L. japonica and 18 host species for G. brasiliensis (Fig. 2, Fig. S2). None of the tested parasitoid performance parameters (DP, SP, ER, number of offspring per day per female) were associated with fly phylogeny for A. japonica, whereas significant phylogenetic signals were detected for SP and number of offspring for L. japonica and G. brasiliensis, and also for ER for G. brasiliensis (Table 2).
Performance of (a) A. japonica, (b) L. japonica and (c) G. brasiliensis on various host species based on the Degree of Parasitism (DP), Success Rate of Parasitism (SP), and Encapsulation Rate (ER), with host species arranged in a phylogenetic tree constructed based on COI gene sequences; the four host species successfully parasitized by G. brasiliensis are highlighted by the gray bar. Values are means and were subjected to a two-way analysis of variance on the effects of parasitoid species, host species and their interaction (see Table 1)
Preimaginal periods varied largely among various drosophilid species, ranging from 14.6 days in D. simulans to 34.8 days in C. amoena (Loew) (F24,5463 = 838.6, P < 0.001) (Fig. 3a) and the developmental time of both A. japonica (Fig. 3b) and L. japonica (Fig. 3c) increased with the host developmental time. Based on the developmental time on D. suzukii, A. japonica (17.5 ± 0.22, n = 108) developed faster than L. japonica (19.8 ± 0.34, n = 130) and L. japonica developed faster than G. brasiliensis (22.4 ± 0.22, n = 84) (F2,319 = 64.5, P < 0.001).
(a) Immature developmental time of the tested drosophilid host species (values are mean ± SE and bars bearing different letters are significantly different, Tukey’s HSD, P < 0.05) and the corresponding relationship of mean developmental time for (b) A. japonica (y = 0.291x + 12.365, r2 = 0.486) and (c) L. japonica (y = 0.423x + 12.842, r2 = 0.847) on those hosts that were attacked and were successfully parasitized (open circles are for hosts with < 5 offspring and were not included in the regression analyses)
The body size of host species varied among the 16 measured Drosophila species, in terms of pupal length (F15,379 = 984.1, P < 0.001), pupal width (F15,379 = 169.6, P < 0.001) and pupal volume (F15,379 = 377.4, P < 0.001) (Table 3). Adult A. japonica females that emerged from larger host species had larger body sizes (hind tibia: F15,379 = 62.9, P < 0.001; ovipositor length F15,379 = 45.7, P < 0.001) and higher mature egg loads (F15,379 = 20.2, P < 0.001) (Table 3). Ovipositor length was positively related to hind tibia length in female A. japonica (ovipositor = 0.320 + 0.520 × hind tibia; F15,379 = 45.7, P < 0.001, R2 = 0.635, P < 0.001). Overall, mean body size of A. japonica females increased with mean host size (y = 0.928x + 1.311, R2 = 0.568, P < 0.001) and mean female mature egg load increased with the mean body size of female wasps (y = 62.15x – 33.63, R2 = 0.695, P < 0.001).
Discussion
This study showed clear differences in the physiological host ranges of A. japonica, G. brasiliensis and L. japonica populations collected from South Korea. Results indicate that G. brasiliensis was the most host specific; L. japonica could develop on a wider host range; and A. japonica developed from most of the 25 tested drosophilid species regardless of their phylogenetic position. All three parasitoids were able to parasitize various host larvae although success levels were generally higher for A. japonica than for G. brasiliensis or L. japonica as measured by offspring production and degree of parasitism and successful parasitism.
These data are critical to define the physiological host range and to determine suitability for use in a classical biological control program, although this may not translate directly into ecological suitability and performance in a pest’s invaded range, as the ecological host range may further limit a parasitoid’s host range in the field. Therefore, ascertaining a parasitoids’ host range and specificity in no-choice, small cage studies, such as those conducted within the confines of a quarantine, is a conservative approach to measuring risk posed by imported natural enemies (van Lenteren et al. 2006). As reviewed in Hoddle et al. (2020), no-choice experiments may overestimate the number of species attacked by a natural enemy species (i.e., fundamental or physiological host range) as the natural enemy is confined with hosts that it may not normally encounter or that it would ignore or abandon if found in nature. The physiological host range is often much larger than the ‘ecological’ host range of the arthropod natural enemy under evaluation (Heimpel and Mills 2017). Quarantine studies are then, to some extent, a relative measure of the ecological risk of the tested natural enemy species. For example, A. japonica is likely an effective D. suzukii parasitoid but also appears from our Quarantine tests to be a greater non-target threat. This is backed up by field surveys (e.g., ecological host range) that show A. japonica attacks not only Drosophila spp. but also species in the genera or subgenera Chymomyza, Dorsilopha, Scaptodrosophila and Sophophora (Ideo et al. 2008; Mitsui and Kimura 2010; Mitsui et al. 2007).
In contrast, the two figitids had narrower physiological host ranges, especially the more host-specific populations of G. brasiliensis (G1 and G3) (Daane et al. 2016; Giorgini et al. 2019; Girod et al. 2018b; Matsuura et al. 2018; Nomano et al. 2017; Seehausen et al. 2021). Moreover, host phylogeny strongly influenced the success rate of parasitism in both figitids. We found that the South Korean L. japonica readily developed from the three melanogaster species group members (D. melanogaster, D. simulans and D. suzukii), and only a few individuals emerged from another 7 species. Girod et al. (2018b) reported similar results for the two Chinese L. japonica populations; both successfully parasitized D. melanogaster and D. subobscura, with only one progeny emerging from D. immigrans. In Japan, L. japonica was collected almost entirely from cherry fruit infested by D. suzukii (Matsuura et al. 2018), while in China L. japonica was collected from fresh fruits infested by D. pulchrella Takamori and Watabe, D. subobscura, and D. suzukii, (which also possess a serrated ovipositor that allows it to penetrate the intact skin of fruits) (Giorgini et al. 2019), while in South Korea it was collected from fresh fruits infested by D. suzukii (Daane et al. 2016). Kimura and Novkovic (2015) found that the L. japonica population was a highly efficient parasitoid of some Drosophila species from its original locality, but was less successful on species from other localities. However, the host range of this parasitoid appears to be largely limited to the melanogaster species group. The G. brasiliensis population we tested had an even narrower physiological host range that included only the closely related D. melanogaster, D. simulans, and D. suzukii, with a small fraction attacked of the more distantly related D. persimilis, suggesting that little to no non-target impact in the field would occur.
There is some plasticity in these parasitoids’ responses to different host species. For example, larval development time varied considerably among the 25 tested drosophilid species. Both A. japonica and L. japonica seemed to adjust their larval development rate to that of their hosts (Fig. 3). These results agree with Kohyama et al. (2017) who showed that A. japonica and Leptopilina ryukyuensis Novkovic & Kimura retarded their own development before host pupariation. The retardation of development time is likely advantageous in koinobiont parasitoids because excessive development of parasitoid larvae before host pupariation would have negative effects on host development and eventually on parasitoid fitness (Harvey and Strand 2002). Asobara japonica also displayed plasticity in female body size, which increased with host size. Body size is often positively correlated with many other fitness components such as adult longevity, fecundity, foraging efficiency and mating ability (e.g., Wang and Messing 2004). As expected, mature egg load also increased with female body size. Such developmental plasticity may contribute to the persistence of A. japonica under variable conditions.
A critical aspect of this work is the acknowledgement that host specificity may vary among geographical populations of G. brasiliensis to the extent that populations studied may represent cryptic species. We report herein that a South Korean G. brasiliensis population parasitized D. melanogaster, D. simulans, and D. suzukii and, to a lesser extent, D. persimilis. Girod et al. (2018b) report a Japanese population of G. cf. brasiliensis was strictly specific to D. suzukii,, whereas a population from China parasitized D. melanogaster, D. suzukii and, to a lesser extent, D. subobscura. Giorgini et al. (2019) tested a G. brasiliensis population from China on 9 of the 25 host species tested in this study and found similar host relationships as reported herein. A first attempt to understand G. brasiliensis population differences and host relationships was made by Nomano et al. (2017), who grouped G. brasiliensis into five lineages (G1-G5) based on molecular analyses of the cytochrome oxidase subunit 1 (CO1) gene and three nuclear DNA regions (ITS1, ITS2 and RpL37). However, Buffington and Forshage (2016) found different lineages were morphologically indistinguishable. Moreover, a recent phylogenomic reconstruction on a representative group of the G1-G5 lineages using a large (ca. 1300 loci) ultraconserved element (UCE) data set showed differences among G. brasiliensis populations, but COI-assignments were intermingled and not monophyletic (Buffington et al. unpubl. data). Still, to reference different studies we will use the G1-G5 designations and our discussion will focus on only the G1 and G3 lineages. Our South Korean populations reported herein were a mixture of G1 and G3 lineages, based on CO1 analyses of voucher specimens. Populations of G. brasiliensis collected from Yunnan, China, also found a mixture G1 and G3 specimens, and genetic distance, based on the COI gene, was large enough (5–7%) to suggest reproductive isolation (Giorgini et al. 2019). Moreover, G1 was the dominant lineage (> 65%) reared from D. suzukii in both the South Korean and Chinese collections processed at the Berkeley quarantine, and the lineages were sympatric and co-existed on the same host plants inhabited by D. suzukii (in South Korea) and by D. suzukii and/or D. pulchrella or D. subpulchrella Takamori and Watabe (in China). Other researchers reported that G1 populations collected from D. suzukii-infested cherries in Japan specialize on D. suzukii (Girod et al. 2018b; Matsuura et al. 2018; Nomano et al. 2017). Seehausen et al. (2021) suggest G1 and G3 are, in fact, cryptic species that can be differentiated by acid-soluble insect protein spectra, some host-searching behaviors, and incompatible crossing. Therefore, based on our molecular examination of voucher specimens from the current study, and the discussion above, we suggest the populations studied herein were a combination of G1 and G3 with the possibility that G1 attacked only D. melanogaster and D. suzukii (a G1 voucher specimen was reared from D. melanogaster) and that G3 attacked D. melanogaster, D. simulans, and D. suzukii, and a small number of D. persimilis.
We also note that the figitids attacked more host species than they were able to successfully develop from, as indicated by the encapsulation rate (Fig. 2). In fact, there was a moderately high encapsulation rate recorded for both G. brasiliensis or L. japonica attacking D. melanogaster and D. suzukii, but rarely in A. japonica. We further suggest that although the G3 lineage tested in the current study can readily attack D. melanogaster and D. simulans, we would not expect it to significantly impact these non-target species if it were released. First, both G. brasiliensis G1 and G3 populations from South Korea and China were found mainly on various fresh fruits, not in over-ripe or rotted fruit that other drosophilid species such as D. melanogaster and D. simulans favor; larvae of these species are attacked by a number of braconid and other figitid parasitoids that do not attack D. suzukii, further suggesting that the parasitoid communities attacking drosophilids are partitioned by microhabitat. In previous experiments we showed that G. brasiliensis was outcompeted by L. japonica and believe that competition may have driven G. brasiliensis to specialize on D. suzukii and other drosophilids (e.g., D. pulchrella) that attack ripening fruit (Wang et al. 2019a). Drosophila melanogaster and D. simulans are two common cosmopolitan species that occur in a wide range of habitats, and they likely overlap with D. suzukii when fresh fruits become unavailable to D. suzukii (e.g., Wang et al. 2016). Ganaspis brasiliensis would likely encounter competition in these alternative fly hosts from the diverse complex of larval parasitoids attacking D. melanogaster and D. simulans on rotting fruit, including A. tabida, L. heterotoma and L. boulardi (Carton et al. 1986; Chabert et al. 2012; Kacsoh and Schlenke 2012; Miller et al. 2015). Ganaspis brasiliensis might attack D. melanogaster or D. simulans when ripe fruit is no longer available and D. suzukii is rare. This could help persistence and increase population sizes and/or enhance their activity in crop systems and may eventually lead to an increased impact on D. suzukii populations. It is also possible that any introduced D. suzukii parasitoids would prefer to attack D. suzukii for a “competition-free space”, to avoid competition with other larval parasitoids on D. melanogaster and D. simulans (Messing and Wang 2009).
We conclude that G. brasiliensis should be considered for release from quarantine for the classical biological control of D. suzukii in North America. This conclusion builds upon an earlier study in Europe (Girod et al. 2018b) of G. brasiliensis (referred to as G. cf. brasiliensis) populations from Yunnan, China, and Tokyo, Japan; L. japonica populations from Yunnan and Beijing, China; and an A. japonica population from Tokyo, Japan. They used D. busckii Coquillett, D. hydei Sturtevant, D. immigrans Sturtevant, D. melanogaster and D. subobscura as non-target species (these fly species were also tested in this study). Girod et al. (2018b) similarly found that G. brasiliensis was the most host specific, followed by L. japonica, and then A. japonica, the latter of which developed from all five non-target species—although there was a lower success rate on D. immigrans (1.2%). Similarly, we found A. japonica attacked these hosts, but that no A. japonica emerged from D. immigrans. This difference may have resulted from population differences (South Korean vs. Japan) as A. japonica was reported to parasitize D. immigrans in field collections in Japan (Ideo et al. 2008), but no evidence for such host-parasitoid field association in South Korea is available. We therefore suggest that A. japonica should not be considered for release, even though it is an effective parasitoid of D. suzukii it presents too great of a non-target risk based on its physiological host range.
Author contributions
AB, XG and KMD conceived and designed experiments; KMD obtained funding; AB and XG collected data; XG, BNH and AB analyzed data; all authors contributed critically to the submitted draft.
Availability of data and material
Data available from the Dryad Digital Repository (to be provided on acceptance of manuscript).
Code availability
Not applicable.
References
Asplen MK, Anfora G, Biondi A, Choi DS, Chu D, Daane KM, Gibert P, Gutierrez AP, Hoelmer KA, Hutchison WD, Isaacs R, Jiang ZL, Karpati Z, Kimura MT, Pascual M, Philips CR, Plantamp C, Ponti L, Vetek G, Vogt H, Walton VM, Yu Y, Zappala L, Desneux N (2015) Invasion biology of spotted wing drosophila (Drosophila suzukii): a global perspective and future priorities. J Pest Sci 88:469–494. https://doi.org/10.1007/s10340-015-0681-z
Biondi A, Wang XG, Miller JC, Miller B, Shearer PW, Zappala L, Siscaro G, Walton VW, Hoelmer KA, Daane KM (2017) Innate olfactory responses of Asobara japonica toward fruits infested by the invasive spotted wing drosophila. J Insect Behav 30:495–506. https://doi.org/10.1007/s10905-017-9636-y
Biondi A, Wang XG, Daane KM (2021) Host preference of three Asian larval parasitoids to closely related Drosophila species: implications for biological control of Drosophila suzukii. J Pest Sci 94:273–283. https://doi.org/10.1007/s10340-020-01272-0
Boughdad A, Haddi K, El Bouazzati A, Nassiri A, Tahiri A, El Anbri C, Eddaya T, Zaid A, Biondi A (2021) First record of the invasive spotted wing Drosophila infesting berry crops in Africa. J Pest Sci 94:261–271. https://doi.org/10.1007/s10340-020-01280-0
Buffington ML, Forshage M (2016) Redescription of Ganaspis brasiliensis (Ihering, 1905), new combination (Hymenoptera: Figitidae), a natural enemy of the invasive Drosophila suzukii (Matsumura, 1931) (Diptera: Drosophilidae). Proc Entomol Soc Wash 118:1–13. https://doi.org/10.4289/0013-8797.118.1.1
Carton Y, Boulétreau B, van Alphen JJM, van Lenteren JC (1986) The Drosophila parasitic wasps. In: Ashburner A, Carson HL, Thompson JN (eds) The genetics and biology of Drosophila, vol 3e. Academic Press, London, pp 347–394
Chabert S, Allemand R, Poyet M, Eslin P, Gibert P (2012) Ability of European parasitoids (Hymenoptera) to control a new invasive Asiatic pest, Drosophila suzukii. Biol Control 63:40–47. https://doi.org/10.1016/j.biocontrol.2012.05.005
Daane KM, Wang XG, Biondi A, Miller B, Miller JC, Riedl H, Shearer PW, Guerrieri E, Giorgini M, Buffington M, van Achterberg K, Song Y, Kang T, Yi H, Jung C, Lee DW, Chung BK, Hoelmer KA, Walton VM (2016) First exploration of parasitoids of Drosophila suzukii in South Korea as potential classical biological agents. J Pest Sci 89:823–835. https://doi.org/10.1007/s10340-016-0740-0
Desneux N, Blahnik R, Delebecque CJ, Heimpel GE (2012) Host phylogeny and specialisation in parasitoids. Ecol Lett 15:453–460. https://doi.org/10.1111/j.1461-0248.2012.01754.x
Farnsworth D, Hamby KA, Bolda M, Goodhue RE, Williams JC, Zalom FG (2017) Economic analysis of revenue losses and control costs associated with the spotted wing drosophila, Drosophila suzukii (Matsumura), in the California raspberry industry. Pest Manag Sci 73:1083–1090. https://doi.org/10.1002/ps.4497
Ferronato P, Woch AL, Soares PL, Bernardi D, Botton M, Andreazza F, Oliveira EE, Correa AS (2019) A phylogeographic approach to the Drosophila suzukii (Diptera: Drosophilidae) invasion in Brazil. J Econ Entomol 112:425–433. https://doi.org/10.1093/jee/toy321
Giorgini M, Wang XG, Wang Y, Chen FS, Hougardy E, Zhang HM, Chen ZQ, Chen HY, Liu CX, Cascone P, Formisano G, Carvalho GA, Biondi A, Buffington M, Daane KM, Hoelmer KA, Guerrieri E (2019) Exploration for native parasitoids of Drosophila suzukii in China reveals a diversity of parasitoid species and narrow host range of the dominant parasitoid. J Pest Sci 92:509–522. https://doi.org/10.1007/s10340-018-01068-3
Girod P, Borowiec N, Buffington M, Chen GH, Fang Y, Kimura MT, Peris-Felipo FJ, Ris N, Wu H, Xiao C, Zhang JP, Aebi A, Haye T, Kenis M (2018a) The parasitoid complex of D. suzukii and other fruit feeding Drosophila species in Asia. Sci Rep 8:e11839. https://doi.org/10.1038/s41598-018-29555-8
Girod P, Lierhmann O, Urvois T, Turlings TCJ, Kenis M, Haye T (2018b) Host specificity of Asian parasitoids for potential classical biological control of Drosophila suzukii. J Pest Sci 91:1241–1250. https://doi.org/10.1007/s10340-018-1003-z
Girod P, Rossignaud L, Haye T, Turlings TCJ, Kenis M (2018c) Development of Asian parasitoids in larvae of Drosophila suzukii feeding on blueberry and artificial diet. J Appl Entomol 142:483–494. https://doi.org/10.1111/jen.12496
Hajek AE, Hurley BP, Kenis M, Garnas JR, Bush SJ, Wingfield MJ, van Lenteren JC, Cock MJW (2016) Exotic biological control agents: a solution or contribution to arthropod invasions? Biol Invasions 18:953–969. https://doi.org/10.1007/s10530-016-1075-8
Harvey JA, Strand MR (2002) The developmental strategies of endoparasitoid wasps vary with host feeding ecology. Ecology 83:2439–2451. https://doi.org/10.2307/3071805
Heimpel GE, Cock MJW (2018) Shifting paradigms in the history of classical biological control. Biocontrol 63:27–37. https://doi.org/10.1007/s10526-017-9841-9
Heimpel GE, Mills NJ (2017) Biological control: ecology and applications. Cambridge University Press, Cambridge
Hennig EI, Mazzi D (2018) Spotted wing drosophila in sweet cherry orchards in relation to forest characteristics, by catch, and resource availability. Insects. https://doi.org/10.3390/insects9030118
Hoddle MS, Lake EC, Minteer CR, Daane KM (2020) Chapter2, importation biological control. In: Mason PG, Dennis N (eds) Biological control: a global initiative. CSIRO Publishing, Canberra
Hougardy E, Hogg BN, Wang XG, Daane KM (2019) Comparison of thermal performances of two Asian larval parasitoids of Drosophila suzukii. Biol Control. https://doi.org/10.1016/j.biocontrol.2019.104000
Ideo S, Watada M, Mitsui H, Kimura MT (2008) Host range of Asobara japonica (Hymenoptera: Braconidae), a larval parasitoid of drosophilid flies. Entomol Sci 11:1–6. https://doi.org/10.1111/j.1479-8298.2007.00244.x
Kacsoh BZ, Schlenke TA (2012) High hemocyte load is associated with increased resistance against parasitoids in Drosophila suzukii, a relative of D. melanogaster. PLoS ONE 7:e34721. https://doi.org/10.1371/journal.pone.0034721
Kasuya N, Mitsui H, Ideo S, Watada M, Kimura MT (2013) Ecological, morphological and molecular studies on Ganaspis individuals (Hymenoptera: Figitidae) attacking Drosophila suzukii (Diptera: Drosophilidae). Appl Entomol Zool 48:87–92. https://doi.org/10.1007/s13355-012-0156-0
Kenis M, Tonina L, Eschen R, van der Sluis B, Sancassani M, Mori N, Haye T, Helsen H (2016) Non-crop plants used as hosts by Drosophila suzukii in Europe. J Pest Sci 89:735–748. https://doi.org/10.1007/s10340-016-0755-6
Kimura MT, Novkovic B (2015) Local adaptation and ecological fitting in host use of the Drosophila parasitoid Leptopilina japonica. Ecol Res 30:499–505. https://doi.org/10.1007/s11284-015-1244-8
Kimura MT, Suwito A (2015) Altitudinal patterns of abundances and parasitism in frugivorous drosophilids in west Java, Indonesia. J Nat Hist 49:1627–1639. https://doi.org/10.1080/00222933.2015.1005709
Kohyama TI, Onizawa K, Kimura MT (2017) Growth rate adjustment of two Drosophila parasitoids in response to the developmental stage of hosts. Ecol Entomol 42:785–792. https://doi.org/10.1111/een.12444
Lee JC, Dreves AJ, Cave AM, Kawai S, Isaacs R, Miller JC, Van Timmeren S, Bruck DJ (2015) Infestation of wild and ornamental noncrop fruits by Drosophila suzukii (Diptera: Drosophilidae). Ann Entomol Soc Am 108:117–129. https://doi.org/10.1093/aesa/sau014
Lee JC, Wang XG, Daane KM, Hoelmer KA, Isaacs R, Sial AA, Walton VM (2019) Biological control of spotted-wing drosophila (Diptera: Drosophilidae): current and pending tactics. J Integr Pest Manag 10:13. https://doi.org/10.1093/jipm/pmz012
Matsuura A, Mitsui H, Kimura MT (2018) A preliminary study on distributions and oviposition sites of Drosophila suzukii (Diptera: Drosophilidae) and its parasitoids on wild cherry tree in Tokyo, central Japan. Appl Entomol Zool 53:47–53. https://doi.org/10.1007/s13355-017-0527-7
Mermer S, Pfab F, Tait G, Isaacs R, Fanning PD, Van Timmeren S, Loeb GM, Hesler SP, Sial AA, Hunter JH, Kaur Bal H, Drummond F, Ballman E, Collins J, Xue L, Jiang D, Walton VM (2020) Timing and order of different insecticide classes drive control of Drosophila suzukii; a modeling approach. J Pest Sci. https://doi.org/10.1007/s10340-020-01292-w
Messing RH, Wang X-G (2009) Competitor-free space mediates non-target impact of an introduced biological control agent. Ecol Entomol 34:107–113. https://doi.org/10.1111/j.1365-2311.2008.01046.x
Miller B, Anfora G, Buffington M, Daane KM, Dalton DT, Hoelmer KM, Stacconi MVR, Grassi A, Ioriatti C, Loni A, Miller JC, Ouantar M, Wang XG, Wiman NG, Walton VM (2015) Seasonal occurrence of resident parasitoids associated with Drosophila suzukii in two small fruit production regions of Italy and the USA. Bull Insectol 68:255–263
Mitsui H, Kimura MT (2010) Distribution, abundance and host association of two parasitoid species attacking frugivorous drosophilid larvae in central Japan. Eur J Entomol 107:535–540
Mitsui H, Van Achterberg K, Nordlander G, Kimura MT (2007) Geographical distributions and host associations of larval parasitoids of frugivorous Drosophilidae in Japan. J Nat Hist 41:1731–1738. https://doi.org/10.1080/00222930701504797
Münkemüller T, Lavergne S, Bzeznik B, Dray S, Jombart T, Schiffers K, Thuiller W (2012) How to measure and test phylogenetic signal. Methods Ecol Evol 3:743–756. https://doi.org/10.1111/j.2041-210X.2012.00196.x
NAPPO (2015) The Secretariat of the North American Plant Protection Organization, Ontario, Canada
Nomano FY, Mitsui H, Kimura MT (2015) Capacity of Japanese Asobara species (Hymenoptera; Braconidae) to parasitize a fruit pest Drosophila suzukii (Diptera; Drosophilidae). J Appl Entomol 139:105–113. https://doi.org/10.1111/jen.12141
Nomano FY, Kasuya N, Matsuura A, Suwito A, Mitsui H, Buffington ML, Kimura MT (2017) Genetic differentiation of Ganaspis brasiliensis (Hymenoptera: Figitidae) from East and Southeast Asia. Appl Entomol Zool 52:429–437. https://doi.org/10.1007/s13355-017-0493-0
Novkovic B, Mitsui H, Suwito A, Kimura MT (2011) Taxonomy and phylogeny of Leptopilina species (Hymenoptera: Cynipoidea: Figitidae) attacking frugivorous drosophilid flies in Japan, with description of three new species. Entomol Sci 14:333–346. https://doi.org/10.1111/j.1479-8298.2011.00459.x
Otto M, Mackauer M (1998) The developmental strategy of an idiobiont ectoparasitoid, Dendrocerus carpenteri: influence of variations in host quality on offspring growth and fitness. Oecologia 117:353–364. https://doi.org/10.1007/s004420050668
Poyet M, Le Roux V, Gibert P, Meirland A, Prevost G, Eslin P, Chabrerie O (2015) The wide potential trophic niche of the Asiatic fruit fly Drosophila suzukii: the key of its invasion success in temperate Europe? PLoS ONE 10:e0142785. https://doi.org/10.1371/journal.pone.0142785
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing. Vienna, Austria. https://www.R-project.org/
Santoiemma G, Trivellato F, Caloi V, Mori N, Marini L (2019) Habitat preference of Drosophila suzukii across heterogeneous landscapes. J Pest Sci 92:485–494. https://doi.org/10.1007/s10340-018-1052-3
Seehausen ML, Ris N, Driss L, Racca A, Girod P, Warot S, Borowiec N, Toševski I, Kenis M (2021) Evidence for a cryptic parasitoid species reveals its suitability as a biological control agent. Sci Rep 10:19096. https://doi.org/10.1038/s41598-020-76180-5
Stewart TJ, Wang XG, Molinar A, Daane KM (2014) Factors limiting peach as a potential host for Drosophila suzukii (Diptera: Drosophilidae). J Econ Entomol 107:1771–1779. https://doi.org/10.1603/ec14197
Tonina L, Mori N, Sancassani M, Dall’Ara P, Marini L (2018) Spillover of Drosophila suzukii between noncrop and crop areas: implications for pest management. Agric For Entomol 20:575–581. https://doi.org/10.1111/afe.12290
Van Driesche RG, Carruthers RI, Center T, Hoddle MS, Hough-Goldstein J, Morin L, Smith L, Wagner DL, Blossey B, Brancatini V, Casagrande R, Causton CE, Coetzee JA, Cuda J, Ding J, Fowler SV, Frank JH, Fuester R, Goolsby J, Grodowitz M, Heard TA, Hill MP, Hoffmann JH, Huber J, Julien M, Kairo MTK, Kenis M, Mason P, Medal J, Messing R, Miller R, Moore A, Neuenschwander P, Newman R, Norambuena H, Palmer WA, Pemberton R, Panduro AP, Pratt PD, Rayamajhi M, Salom S, Sands D, Schooler S, Schwarzlander M, Sheppard A, Shaw R, Tipping PW, van Klinken RD (2010) Classical biological control for the protection of natural ecosystems. Biol Control 54:S2–S33. https://doi.org/10.1016/j.biocontrol.2010.03.003
van Lenteren JC, Bale J, Bigler E, Hokkanen HMT, Loomans AM (2006) Assessing risks of releasing exotic biological control agents of arthropod pests. Annu Rev Entomol 51:609–634. https://doi.org/10.1146/annurev.ento.51.110104.151129
Van Timmeren S, Isaacs R (2013) Control of spotted wing drosophila, Drosophila suzukii, by specific insecticides and by conventional and organic crop protection programs. Crop Prot 54:126–133. https://doi.org/10.1016/j.cropro.2013.08.003
Wang XG, Messing RH (2004) Fitness consequences of body-size-dependent host species selection in a generalist ectoparasitoid. Behav Ecol Sociobiol 56:513–522. https://doi.org/10.1007/s00265-004-0829-y
Wang XG, Stewart TJ, Biondi A, Chavez BA, Ingels C, Caprile J, Grant JA, Walton VM, Daane KM (2016) Population dynamics and ecology of Drosophila suzukii in Central California. J Pest Sci 89:701–712. https://doi.org/10.1007/s10340-016-0747-6
Wang XG, Nance AH, Jones JML, Hoelmer KA, Daane KM (2018) Aspects of the biology and reproductive strategy of two Asian larval parasitoids evaluated for classical biological control of Drosophila suzukii. Biol Control 121:58–65. https://doi.org/10.1016/j.biocontrol.2018.02.010
Wang XG, Hogg BN, Hougardy E, Nance AH, Daane KM (2019a) Potential competitive outcomes among three solitary larval endoparasitoids as candidate agents for classical biological control of Drosophila suzukii. Biol Control 130:18–26
Wang XG, Kaçar G, Daane KM (2019b) Temporal dynamics of host use by Drosophila suzukii in California’s San Joaquin Valley: implications for area-wide pest management. Insects. https://doi.org/10.3390/insects10070206
Wang XG, Biondi A, Daane KM (2020) Functional responses of three candidate Asian larval parasitoids evaluated for classical biological control of Drosophila suzukii (Diptera: Drosophilidae). J Econ Entomol 113:73–80. https://doi.org/10.1093/jee/toz265
Wang X, Biondi A, Nance AH, Zappala L, Siscaro G, Hoelmer KA, Daane KM (2021) Assessment of Asobara japonica as a potential biological control agent for the spotted wing drosophila, Drosophila suzukii. Entomol Gen 41:1–12. https://doi.org/10.1127/entomologia/2020/1100
Acknowledgements
We thank John Hutchins, Dominique Shield, Ariana Bautista, Marilyn Chan, Chesa Cox, Christy Kim and Hiromi Murata (University of California, Berkeley) for assistance with work in quarantine; Kei-Li Ooi (University of California, Berkeley) for the construction of host phylogeny tree; Betsey Miller, Jeffrey Miller, Peter Shearer and Vaughn Walton (Oregon State University) and Yoohan Song (Gyeongsang University, Jinju, South Korea) for helping with parasitoid collections in South Korea; Matthew Buffington (USDA-ARS, Systematic Entomology Laboratory, Washington D.C., USA) and Emilio Guerrieri and Massimo Giorgini (Institute for Sustainable Plant Protection, CNR, Portici, Italy) for help with parasitoid identification; and Kim Hoelmer (USDA-ARS, Beneficial Insects Introduction Research Unit, Newark, DE, USA) for programmatic support. Funding was provided by the USDA APHIS (Farm bill, fund 14-8130-0463) and the National Institute of Food and Agriculture, and USDA Specialty Crops Research Initiative under Agreements 2015-51181-24252 and 2020-51181-32140. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture (USDA). USDA is an equal opportunity provider and employer.
Funding
Funds were provided by the USDA APHIS (Farm bill, fund 14-8130-0463), the National Institute of Food and Agriculture, USDA Specialty Crops Research Initiative Agreements Nos. 2015-51181-24252 and 2020-51181-32140.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Blas Lavandero.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Daane, K.M., Wang, X., Hogg, B.N. et al. Potential host ranges of three Asian larval parasitoids of Drosophila suzukii. J Pest Sci 94, 1171–1182 (2021). https://doi.org/10.1007/s10340-021-01368-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10340-021-01368-1