Abstract
Double-stranded RNA (dsRNA)-mediated gene silencing, or RNA interference (RNAi), is an emerging biotechnology that has been a breakthrough tool for crop protection. Exogenous dsRNA triggers the RNAi pathway, silences genes, disrupts protein function, and can cause insect mortality. However, effective delivery of the dsRNA is problematic, particularly in systems with long-lived, endophagous insects such as the emerald ash borer (EAB), Agrilus planipennis, a tree-killing nonnative invader that attacks ash, Fraxinus spp. Larvae feed on cambial tissue causing rapid tree death. EAB is susceptible to RNAi, but we lack a practical means of delivery. Here we evaluated delivery of dsRNA to green, F. pennsylvanica, and tropical ash, F. uhdei, through root and/or petiole absorption, and also demonstrated dsRNA absorption through the EAB egg chorion. We labeled exogenous dsRNA using a fluorescing label and then used confocal microscopy and RT-qPCR to evaluate its distribution in plant and insect tissues. Labeled dsRNAs are detectable in root, stem, and leaf tissues 48-h postapplication. In excised ash branches, labeled dsRNA is detectable in the inner bark and in recovered EAB neonates 8-day postapplication. Eggs and larvae emerging from treated eggs also presented fluorescing dsRNA under confocal imaging. Adult EAB-fed tropical ash leaves treated with in vitro synthesized EAB-specific dsSHI through petiole absorption experience a significant knockdown of the shi gene and a significant mortality. Our findings provide a proof of concept that delivery of dsRNAs through topical or systemic application methods is a feasible means of suppressing EAB, providing hope for future tree protection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Key message
-
RNAi through exogenous dsRNA application is an innovative and promising tool for insect suppression.
-
We evaluated labeled dsRNA uptake through ash plants and insect tissue.
-
Confocal microscopy confirmed the presence of dsRNA within plant material, EAB eggs, and insect tissue post dsRNA application.
-
Gene expression in adult EAB fed on ash leaves exposed to dsRNA solution was significantly reduced, as was beetle survival.
-
Root absorption and topical application targeting EAB eggs represent feasible dsRNA delivery methods that should be further evaluated.
Introduction
The RNA interference (RNAi) pathway is a cellular regulatory mechanism that limits the expression of target genes, resulting in gene silencing. It is an ancient antiviral process that appeared before the divergence of plants and animals (Sharp 2001) and has been described in almost all eukaryotic organisms, including insects (Agrawal et al. 2003). The pathway is triggered by the introduction of double-stranded RNA (dsRNA), which is processed by the enzyme dicer into small interference RNAs (siRNAs). The siRNA is loaded into a protein complex named RISC (RNA-induced silencing complex) that guides the siRNA to its complementary messenger RNA (mRNA) sequence. The targeted mRNA is cleaved, preventing the translation of the gene, thus causing its silencing (Agrawal et al. 2003; Meister and Tuschl 2004). The RNAi mechanism involves silencing specific 19–21 base pair sequences (siRNAs) of target genes in a given species or closely related species (Bachman et al. 2013). As a pest management strategy, this characteristic makes the mechanism more specific than conventional pesticides, minimizing the risk to nontarget and beneficial organisms (Baum and Roberts 2014; Bachman et al. 2016). RNAi is emerging as a next-generation pest control strategy and as a tool to manage various agricultural insect pests and pathogens (Mamta and Rajam 2017).
Although RNAi efficiency varies among insects, coleopterans are sensitive to dsRNAs either by feeding or by topical application (Baum et al. 2007; Baum and Roberts 2014; Niu et al. 2019). Beyond agriculture pests, the use of RNAi technology has been evaluated for wood boring forest pests; Asian longhorned beetle, Anoplophora glabripennis (Rodrigues et al. 2017a), southern pine beetle, Dendroctonus frontalis (Kyre et al. 2019), and emerald ash borer (EAB), Agrilus planipennis Fairmaire (Coleoptera: Buprestidae) (Rodrigues et al. 2017b) have all shown susceptibility. Previous studies have demonstrated that the RNAi mechanism is functional in EAB (Zhao et al. 2015). Both larvae and adults are sensitive to ingested and microinjected dsRNA, and suitable target genes for RNAi in EAB have been identified (Rodrigues et al. 2017b, 2018). Thus, this technology shows promise as a potential method for EAB management.
Native to northeastern Asia, EAB is an exotic phloem-feeding beetle that was accidentally introduced into North America through infested solid wood packing material (Cappaert et al. 2005; Poland and McCullough 2006). North American ash, Fraxinus spp., are highly susceptible to EAB, and since its introduction into North America EAB has killed millions of ash throughout its invaded range, it is one of the costliest invaders in US history (Aukema et al. 2011).
Adult EAB oviposit on stems and large branches of ash. Neonate larvae bore through the outer bark to feed on phloem, excavating serpentine galleries and destroying the cambial tissue. Larval feeding disrupts translocation of water and nutrients (Haack et al. 2002), and infested ash die 2–4 years after showing the first signs of infestation (Herms and McCullough 2014). Management of EAB has been a continual challenge (Aukema et al. 2011; Duan et al. 2018; GAO 2006), and development of RNAi technology provides new hope (Rodrigues et al. 2018).
Beyond demonstrating dsRNA efficacy and specificity are the challenges associated with delivery. Both plants and insects contain RNases that could impede or prevent effective delivery of dsRNAs to target sites (Yu et al. 2013); understanding these processes and overcoming any impediments is key to moving RNAi technology forward. Here we evaluate dsRNA distribution through plant and insect tissues. We exposed ash plant tissues and insect tissues to fluorescently labeled dsRNA and then used confocal laser scanning microscopy to image fluorescence in both plant and insect material (Table 1). We evaluated labeled dsRNA uptake by plant tissue, uptake by EAB neonate larvae hatching from dsRNA-treated eggs, and EAB neonate larvae hatched from eggs attached to dsRNA-treated twigs and feeding on treated twigs. We also measured adult EAB survival and gene expression after exposing beetles to dsRNA delivered by leaf petiole absorption.
Methods
In vitro synthesis of dsRNA
Total RNA was extracted from neonate EAB larvae using TRIzol reagent (ThermoFisher, USA). The quality and quantity of the RNA were checked by electrophoresis and spectrophotometer (Nanodrop Technologies, Wilmington, DE, USA). cDNA was synthesized from 1 µg of total RNA using a M-MLV reverse transcriptase kit (ThermoFisher, USA).
The PCR templates for in vitro synthesis of dsRNA were generated using gene-specific primers containing T7 polymerase promoter sequences. PCR conditions were 4 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 60 °C, and 45 s at 72 °C, finishing with an extension step at 72 °C for 10 min. The PCR template was purified using a PCR purification kit (Qiagen Inc., Valencia, CA, USA). After PCR purification, dsRNA synthesis was performed using the MEGAscript RNAi Kit (Ambion Inc., Foster City, CA, USA) following the manufacturer’s instructions. The reaction was incubated for 14 h at 37 °C, followed by 15 min of DNase treatment. The dsRNA was precipitated by adding 0.1 × volume of sodium acetate (3 M, pH 5.2) and 2.5 × the volume of 100% ethanol and kept at − 20 °C for at least 2 h followed by centrifugation at 4 °C and 14,000g for 30 min. The dsRNA pellet was then rinsed with 750 μl of 75% ethanol and centrifuged again at 4 °C for 15 min. The ethanol was removed and the dsRNA was diluted in ultrapure distilled water. The quality of the dsRNA was checked by electrophoresis and quantified using a spectrophotometer (NanoDrop Technologies, Wilmington, DE USA). To attain the desired concentration for each assay, dsRNA samples were vacuum concentrated using Concentrator plus (Eppendorf, Hauppauge, NY, USA).
Labeling dsRNA
To evaluate dsRNA uptake through plant and insect tissue, dsRNAs were fluorescently labeled using the Silencer siRNA Cy™3 labeling kit (ThermoFisher, USA) with minor modifications to the manufacturer’s protocol. dsRNAs were labeled by adding Cy3 labeling reagent and incubating for 3 h at 37 °C. Labeled dsRNA was precipitated with 0.1 volume of NaCl and 2.5 volumes of 100% ethanol followed by incubation at − 20 °C for 30 min. Precipitated labeled dsRNA was recovered by centrifugation at 4 °C and 14,000g for 20 min, and the pellet was further washed with 70% ethanol. The recovered pellet was air-dried and re-suspended in nuclease-free saline. The concentration of labeled dsRNA was determined using a spectrophotometer (NanoDrop Technologies, Wilmington, DE USA).
dsRNA absorption by plant tissue
Ash seedlings
Green ash, F. pennsylvanica, seedlings ~ 5 cm in length (N = 2) were immersed in a 5-ml beaker containing labeled dsSHI (Rodrigues et al. 2017b) (70 µg/ml) in 2 ml RNase and DNase-free water (Table 1a), covered with parafilm, and incubated at room temperature (~ 21 °C) for 48 h to allow capillary action to draw the solution to the top of the plants. Samples were washed with deionized water, and total RNA was extracted from root, stem, and leaf tissue, and cDNA was synthesized from 1 µg of total RNA using a M-MLV reverse transcriptase kit (ThermoFisher, USA). The cDNA was subjected to PCR validation using primers specific to dsRNA (Rodrigues et al. 2017b). PCR conditions were 4 min at 94 °C, followed by 35 cycles of 30 s at 94 °C, 30 s at 60 °C, and 45 s at 72 °C, finishing with an extension step at 72 °C for 10 min.
Ash branches
Greenhouse-grown tropical ash, F. uhdei, were used to evaluate dsRNA uptake (Table 1b). Branches (~ 1 cm diameter, N = 5) were excised and immediately placed into 5-ml plastic test tubes containing 1 ml of a labeled dsGFP solution, covered with parafilm, and maintained at room temperature (~ 21 °C). The dsRNA concentration was 20 μg/ml resulting in a final exposure of 20 μg of dsRNA per branch. After the initial 1 ml of labeled dsGFP was taken up by the plant tissue, 500 µL of water was added to the tubes every other day.
dsRNA uptake by insect tissue
Recovered EAB neonate larvae
Ingestion of dsRNA-treated ash tissue by developing neonate larvae (Table 1c) was evaluated using the same greenhouse-grown tropical ash described above. Branches were artificially infested with laboratory-reared EAB eggs (Olson and Rieske 2019) prior to branch excision. Briefly, pieces of coffee filter (Kroger, Cincinnati OH) with laboratory-reared EAB eggs laid on them were attached to branches and secured using 1.5-cm-wide strips of parafilm (Bernis NA, Neenah, WI). Additional eggs (N = 5–8) were kept under the same conditions, but in Petri dishes with moistened filter paper, to help identify the hatching time. When the eggs in the Petri dish started hatching, the branches were excised and treated with dsRNA as described above. There were five branches per treatment (labeled dsGFP and water), each one artificially infested with six EAB eggs. After 8 days the treated branches were carefully debarked using a sharp knife, and EAB egg hatch, larval development, and phloem consumption were evaluated (Olson and Rieske 2019); confocal microscopy was used to evaluate labeled dsRNA uptake.
EAB egg chorion
To evaluate the uptake of dsRNA through the egg chorion (Table 1d), ~ 15 laboratory-reared EAB eggs were placed in Petri dishes (60 × 9 mm) with moistened filter paper and soaked with 1 μl of labeled dsGFP at a concentration of 1 μg/μl. Water was used as the negative control. The dsRNA treatment was repeated 3 times over 48 h, during which the eggs were kept at 23 °C and 75% relative humidity in a growth chamber. After 3 days the eggs were evaluated under confocal microscopy to assess labeled dsRNA uptake; confocal microscopy was used to evaluate labeled dsRNA uptake.
Confocal laser scanning microscopy
All tissues were imaged using an Olympus FV1200 confocal laser scanning microscopy (Center Valley, PA, USA). The fluorescence of Cy3 in the tissues was detected using a 559-nm laser with a bandpass filter of 570/100 nm. ImageJ software was used for image analysis. Fluorescently labeled samples and control samples, treated either with unlabeled dsRNA or water, were imaged simultaneously using the same parameters to ensure minimal background fluorescence in the negative controls and the positive signals for direct comparison.
Ash seedlings
After 48-h exposure to labeled dsRNA, plants were sectioned into stem (1 mm cross section), root (simply separated from the aerial part), and leaf (1 × 1 cm square) tissues using a razor blade and imaged using confocal microscopy under the conditions described above.
Ash branches and recovered EAB neonate larvae
After 8 days of exposure to labeled dsRNA, branches were carefully debarked, EAB neonate larvae were recovered, and the branches were sectioned. Bark, ash stems, and EAB larvae recovered from treated twigs were imaged using the instrumentation and methods described above.
EAB egg chorion and neonate larvae
After 48-h exposure to labeled dsRNA, EAB eggs were imaged using the same method described above. Similarly, eggs that had been treated with the labeled dsRNA were allowed to hatch and the neonate larvae were imaged after 48 h.
Adult insect assays
Leaves of greenhouse-grown tropical ash containing 3–5 leaflets were excised with petioles intact and immersed in 2 ml of water containing 70 µg/ml of EAB-specific dsSHI for 2 h to allow uptake; dsGFP was used as the control (Table 1e). Laboratory-reared adult EAB (N = 5–10) was then placed on dsRNA-treated leaves and allowed to feed for 10 days (Fig. 1). Assays were held in ventilated plastic cups (500 ml) and maintained in a growth chamber at 25 °C and 16:8 L:D. Each treatment was replicated three times, and beetle survival was recorded daily. Mortality (%) was calculated, and the mean value of the experimental replicates was analyzed using a one-way ANOVA, followed by Tukey’s post hoc t test. In addition, in a fourth replicate, adult EABs (N = 5–10) were collected at 24 h, 72 h, and 120 h following exposure to dsRNA-treated leaves, and total RNA was isolated for gene expression analysis.
Quantitative real-time PCR analysis
The levels of transcript expressed by eggs, neonate larvae, and adults after exposure to dsRNA were measured by quantitative real-time PCR (RT-qPCR). Following total RNA extraction, cDNA was synthesized using M-MLV reverse transcriptase (Life Technologies, Carlsbad, CA, USA) and used as a template for gene expression studies. The expression analyses were conducted using SYBR Green PCR Master Mix. The PCR mixture contained 1 μl of cDNA, 0.2 μl of each primer (10 mM), 5 μl of the SYBR green PCR master mix, and 3.6 μl of ddH2O, totaling 10 μl. Real-time quantitative RT-PCR (RT-qPCR) was performed using the StepOnePlus Real-Time PCR system (Life Technologies, Carlsbad, CA, USA) under the following conditions: one cycle of 20 s at 95 °C, followed by 40 cycles of denaturation at 95 °C (3 s), annealing, and extension for 30 s at 60 °C, ending with generation of a melting curve to confirm a single peak and rule out nonspecific product and primer dimer formations. The reference gene used was TEF1α (Rajarapu et al. 2012; see Supplemental Materials), and the 2-ΔΔCt method was used to calculate the relative expression of the target gene compared to the control (Livak and Schmittgen 2001). A two-tailed t test was used for statistical analysis to compare the means of a single variable.
Results
Uptake of dsRNA into plant tissue
Ash seedlings
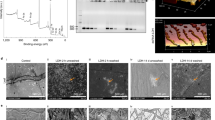
Examination of root and stem sections, and the leaf surface, by confocal imaging revealed the presence of labeled dsRNA in all tissues evaluated, including stem (Fig. 2a, b), root (Fig. 2c, d), and leaf tissue (Fig. 2e), indicating the transport of dsRNA through the vascular tissue from its point of initial application. Molecular validation of dsRNA transport was done by PCR, as described above, after 48 h of treatment. Total RNA was isolated from sampled plant tissues and amplified using gene-specific oligonucleotides. The RT-PCR of cDNA of the shi gene (Fig. 3a) from root (Lane 1), leaf (Lane 2), and stem (Lane 3) tissues following electrophoresis on 1% agarose gel alongside a DNA ladder (Lane M) indicates that dsRNA introduced exogenously was amplified in root and stem tissues, but not in leaf tissue. The dsRNA of shi synthesized from a commercial kit (Fig. 3b) presents the same amplicon size (483 bp) as that recovered from plant tissues exposed to dsSHI, confirming its identity.
Confocal microscope images of green ash stem, root, and leaf tissue after exposure to Cy3 labeled dsSHI. a Stem cross section showing deposition of Cy3 labeled dsSHI on the xylem tissues and b the negative control showing minimal autofluorescence. c Green ash root after 48-h exposure to labeled dsSHI reveals the presence of labeled dsRNA, whereas d the unlabeled dsSHI negative control does not. e Adaxial leaf surface after 48-h exposure to labeled dsSHI reveals the presence of labeled dsRNA in midrib, veins, and the nearby areas. Confocal imaging clearly demonstrates transport of labeled dsRNA from roots to the plant extremities within 48 h
RT-PCR products on a 1% agarose gel of a cDNA of EAB-specific shi from green ash root (lane 1), leaf, showing no amplification (lane 2), and stem (lane 3) tissues adjacent to a DNA ladder (lane M), and b dsRNA of EAB-specific shi synthesized from a commercial kit, which presents the same amplicon size (400–500 bp) as the cDNA recovered from dsRNA-exposed plant tissues
Ash branches
Confocal microscopy of branch sections after 8 days of treatment confirms that labeled dsRNA distributes throughout branch vascular tissues (Fig. 4a) and is visible beneath the inner bark (Fig. 5a, b), whereas untreated controls (Figs. 4b and 5c) fluoresce only slightly. Confocal imaging and RT-qPCR demonstrate that the EAB-specific dsRNA was transported distally through the plant vascular tissues.
Branches of tropical ash were a debarked to evaluate the presence of labeled dsRNA on the inner bark. Inner bark of branch exposed to the labeled dsRNA b clearly shows the fluorescing dsRNA in tissues; the tissue that was consumed by feeding EAB larva lack fluorescence. c Inner bark of branch exposed to water (negative control) showing minimal fluorescence
Uptake of dsRNA through egg chorion
Confocal imaging of EAB eggs following dsRNA treatment confirms absorption of labeled dsRNA through the chorion (Fig. 6a). A single-plane confocal image of the developing EAB embryo inside the egg showed positive fluorescence, suggesting absorption in the developing EAB embryo (Fig. 6b) 48 h after exposure to the labeled dsGFP, whereas untreated control eggs demonstrate minimal fluorescence (Fig. 6c).
EAB eggs exposed to labeled Cy3-dsGFP (a) retain the fluorescing dsRNA solution, which is visible in the developing EAB embryo (b), whereas minimal fluorescence is evident for the negative control (c). Panel b is a single-plane confocal image that shows a section of the developing EAB embryo inside the egg
Uptake of dsRNA into insect tissue
Recovered EAB neonate larvae
EAB neonate larvae were recovered from F. udhei branches exposed to labeled dsRNA, and confocal imaging of the insect tissue (Fig. 7a) confirms the uptake of labeled dsRNA by larvae that fed on treated ash branches exposed to labeled dsRNA.
Single-plane confocal imaging of larvae recovered from ash branches exposed to labeled Cy3-dsRNA (a) detects intense fluorescence in the alimentary canal, confirming the intake of dsRNA. Larvae emerged from eggs exposed to labeled dsRNA (b) also shows dsRNA in the alimentary canal. Larvae recovered from ash branches exposed to water (c) do not fluoresce under confocal microscopy
EAB egg chorion
EAB neonate larvae that emerged from eggs exposed to labeled dsRNA were evaluated, and the labeled dsRNA was also present within the larvae (Fig. 7b, c). Overall confocal imaging confirmed that EAB neonate larvae uptake the dsRNA either by the exposure of eggs or ash branches to the labeled dsRNA treatments.
Adult insect assays
Adult beetles ingesting ash leaves treated with EAB-specific dsSHI experienced 61% mortality by day 10 (Fig. 8), which was significantly greater than beetles ingesting the dsGFP control (~ 15% mortality). Gene expression studies demonstrated no change in gene expression 24 h and 72 h after beetles were fed ash seedlings treated with EAB-specific dsSHI (Fig. 9a, b), but there was a significant depletion of shi transcript levels after 120 h (Fig. 9c).
Discussion
Manipulation of the RNAi pathway has emerged as a useful tool in functional gene regulation and has potential to play a significant role in pest management programs (Burand and Hunter 2013). However, deployment of dsRNA as a pest management strategy will require practical and efficient means of delivery. One of the critical steps in developing a dsRNA delivery methodology is demonstrating uptake of dsRNAs by plant cells and by target insect tissues. This study demonstrates the feasibility of dsRNA delivery through topical application and plant tissue absorption targeting a tree-killing insect pest of global importance, the emerald ash borer.
Microinjection and droplet feeding for dsRNA delivery to EAB larvae and adults induces an effective RNAi response (Rodrigues et al. 2017b, 2018), but lacks practical application. Hunter et al. (2012) first described an initial proof of concept for dsRNA delivery in plants without transformation, and subsequently, additional nontransgenic methods of delivery have been reported. Topical applications of dsRNAs have proven effective in some woody plant systems; a hemipteran psyllid and a xylem feeding leafhopper are able to uptake dsRNAs by feeding on citrus or grapevines treated with topically applied dsRNAs (Hunter et al. 2012; Andrade and Hunter 2016). Plant-mediated dsRNA delivery to control brown marmorated stink bug has proven effective (Ghosh et al. 2017), and soil drench, foliar spray, and trunk injections have been used on citrus (Ghosh et al. 2018).
In this study we demonstrate that adult EAB-fed tropical ash leaves treated with in vitro synthesized EAB-specific dsSHI through petiole absorption experience a significant knockdown of the shi gene, and significant beetle mortality, supporting our assertion that the EAB dsRNA was absorbed by the petiole and delivered to insects that fed on these leaves. Dalakouras et al. (2018) demonstrated that dsRNA delivered by petiole absorption penetrates and disseminates through the plant tissues. The plant vascular system, particularly the xylem, is a RNase-free environment (Doering-Saad et al. 2002; Taning et al. 2016), so any dsRNA accessing this tissue should remain stable and available for target pest consumption for long periods. Our confocal images of green ash stem sections clearly show deposition of labeled dsRNA in the xylem. Further confocal imaging also confirmed the presence of labeled dsRNA in the inner bark of tropical ash and in the alimentary canal of neonate larvae recovered from those treated branches, thus demonstrating the utility of the plant vascular system for effective dsRNA delivery. There is mounting evidence suggesting that exogenous dsRNAs remain intact and are not degraded by woody plants. Dalakouras et al. (2018) showed that hairpin RNA (hpRNA) and small interfering RNA (siRNA) are both transported exclusively in woody plant xylem and do not reach the plant cell cytoplasm, thus remaining unprocessed by the plant RNAi machinery. These studies support our findings and suggest that in our study, the dsRNA reached the target EAB and was not degraded by the plant.
Additionally, root absorption of dsRNA was investigated in ash seedlings and PCR was used to recover dsRNA from roots, stem, and leaves (Fig. 3). Here we recovered the dsRNA from roots and stem but not from the leaves. Majidiani et al. (2019) evaluated uptake of dsRNAs thought root absorption, and even long dsRNA molecules (430 bp length) could be uptaken and trigger gene silencing in the target insect feeding on the leaf. Our inability to recover dsRNAs in leaf tissue through PCR is somewhat puzzling; potential explanations for the lack of detection may be that the concentration of the dsRNA was too low (70 ng/ml), or that our timing was not optimal. Dalakouras et al. (2018) showed an accumulation of hpRNA in leaf tissue over 10 days when delivered via petiole absorption; they recovered hpRNA in leaf tissue after 1 day, but then recovered hpRNA in even higher concentrations after 3 and 10 days, suggesting that it was being concentrated in leaf tissue by the plant over time. In our assay we collected samples after only 48 h. A later sampling interval may have allowed adequate time for the dsRNA to disseminate through the plant and concentrate in leaf tissue, which then may have been recovered and detected by our PCR. Further investigations using RNAseq would also clarify whether the dsRNA is being processed by plant RNAi machinery.
Our confocal images of green ash stem sections clearly show deposition of labeled dsRNA in the xylem. Further confocal imaging also confirmed the presence of labeled dsRNA in the inner bark of tropical ash and in the alimentary canal of neonate larvae recovered from those treated branches, thus demonstrating the utility of the plant vascular system for effective dsRNA delivery.
In insect cells, the RNAi pathway is initiated by cleavage of dsRNA by the RNase III enzyme dicer into 21-nucleotide siRNAs which are loaded onto the RISC complex (Joga et al. 2016). Developing plants that express insect-specific dsRNAs that trigger the RNAi pathway may be challenging, since the introduced dsRNA may be processed by dicer-like enzymes (DCLs) into 21-nucleotide siRNAs within the plant that will then be loaded onto the plant RISC complex, and plant-processed siRNAs may be less efficient in triggering gene silencing than long unprocessed dsRNA molecules. Following dicer cleavage, dsRNAs greater than 200 nucleotides in length result in many siRNAs that may contribute to RNAi response in some insect species (Andrade and Hunter 2016). Thus, to be a suitable strategy to trigger RNAi in insects, dsRNA delivered to plants must remain unprocessed by plant enzymes. Our confocal microscopy images demonstrate deposition of EAB-specific dsSHI in the xylem of green ash, which is devoid of DCLs, providing visual evidence that plant-based delivery is a viable approach for delivering dsRNA molecules to induce RNAi in EAB.
After confirming delivery of dsRNA by plant tissue, we investigated how dsRNA might penetrate the egg chorion and be uptaken by EAB embryos following a topical application. Insect eggs are multilayered and structurally complex, having evolved to protect developing embryos against biotic and abiotic threats. But insect eggs also present aeropyles for respiration, micropyles for fertilization, and pillars for oxygen movement in the inner eggshell; these structures represent potential penetration sites where compounds can penetrate the chorion and reach the embryo (Campbell et al. 2016). Our confocal imagery showed the labeled dsRNA within the egg, within the embryo, and also in the intestinal tract of EAB neonate larvae emerged from treated eggs, suggesting that topical application may be developed as a viable option for dsRNA delivery targeting EAB eggs.
The RNAi pathway can be successfully induced in EAB using these novel absorption-mediated delivery methods. However, advancement of these methods poses challenges that must be addressed to make them practical for field applications. Of primary importance is production of dsRNAs, which, for research purposes, are reliant on commercial kits. However, the price of in vitro synthesized dsRNA is far too high for commercial application. Since large-scale production of dsRNA is needed, a cost-effective production method will be required for mass production of dsRNA (Palli 2014). dsRNA production is under development in many biotech companies, which should bring this technology to large-scale, low-cost, ready-to-use RNAi products. Additional studies must also evaluate uptake and transport, accumulation in plant tissues, integrity, and stability of pest-specific dsRNAs in plant tissues. Carriers for dsRNA delivery have been under intense investigation, and nanoparticle-mediated RNAi shows promising results (Das et al. 2015). Oral delivery of dsRNAs to mosquitoes using polymeric nanoparticles, such as chitosan or carbon quantum dot, increases the stability and enhances cellular uptake of dsRNAs, thereby enhancing the efficacy of the RNAi pathway (Das et al. 2015; Zhang et al. 2010). This approach may be applicable to increase the transport, integrity, and stability of dsRNAs in our system.
Development of RNAi-based biopesticides will be accelerated once mass production of dsRNAs becomes practical, efficient delivery systems are developed, and application under field conditions becomes feasible (Kola et al. 2015). Here, we demonstrate that a gene-specific dsRNA can be delivered by both petiole absorption and oral feeding to induce RNAi in adult EAB. Our initial results on the use of an RNAi-based nontransgenic method for controlling EAB are promising.
Author contributions
LKR, TBR, and SLR conceived the experiments; FP, TBR, and SLR conducted the experiments; TK, FP, TBR, and SLR performed confocal imaging; FP processed and edited the images; FP, TBR, SLR, and LKR analyzed the results; FP, TBR, SLR, and LKR prepared the manuscript. All authors provided input and approved the manuscript.
References
Andrade EC, Hunter WB (2016) RNA interference–natural gene-based technology for highly specific pest control (HiSPeC). RNA INTERFERENCE
Agrawal N, Dasaradhi PVN, Mohmmed A, Malhotra P, Bhatnagar RK, Mukherjee SK (2003) RNA interference: biology, mechanism, and applications. Microbiol Mol Biol Rev 67:657–685
Aukema JE, Leung B, Kovacs K, Chivers C, Britton KO, Englin J et al (2011) Economic impacts of non-native forest insects in the continental United States. PLoS ONE 6:e24587. https://doi.org/10.1371/journal.pone.0024587
Bachman PM, Bolognesi R, Moar WJ, Mueller GM, Paradise MS, Ramaseshadri P, Tan J, Uffman JP, Warren J, Wiggins BE, Levine SL (2013) Characterization of the spectrum of insecticidal activity of a double-stranded RNA with targeted activity against western corn Rootworm (Diabrotica virgifera virgifera LeConte). Transgenic Res 22:1207–1222. https://doi.org/10.1007/s11248-013-9716-5
Bachman PM, Huizinga KM, Jensen PD, Mueller G, Tan J, Uffman JP, Levine SL (2016) Ecological risk assessment for DvSnf7 RNA: a plant-incorporated protectant with targeted activity against western corn rootworm. Regul Toxicol Pharmacol 81:77–88
Baum JA, Roberts JK (2014) Progress towards RNAi-mediated insect pest management. Adv Insect Physiol 47:249–295
Baum JA et al (2007) Control of coleopteran insect pests through RNA interference. Nat Biotechnol 25:1322–1326
Burand JP, Hunter WB (2013) RNAi: future in insect management. J Invertebr Pathol 112(Suppl. 1):S68–S74
Cappaert D, McCullough DG, Poland TM, Siegert NW (2005) Emerald ash borer in North America: a research and regulatory challenge. Am Entomol 51:152–163
Campbell BE, Pereira RM, Koehler PG (2016) Chapter 5: complications with controlling insect eggs. In: Trdan S (ed) Insecticide resistance. InTech, Rijeka. https://doi.org/10.5772/61848
Das S, Debnath N, Cui Y, Unrine J, Palli S (2015) Chitosan, carbon quantum dot, and silica nanoparticle mediated dsRNA delivery for gene silencing in Aedes aegypti: a comparative analysis. ACS Appl Mater Interfaces. https://doi.org/10.1021/acsami.5b05232
Dalakouras A, Jarausch W, Buchholz G, Bassler A, Braun M, Manthey T, Krczal G, Wassenegger M (2018) Delivery of hairpin RNAs and small RNAs into woody and herbaceous plants by trunk injection and petiole absorption. Front Plant Sci 9:1253. https://doi.org/10.3389/fpls.2018.01253
Doering-Saad C, Newbury HJ, Bale JS, Pritchard J (2002) Use of aphid stylectomy and RT-PCR for the detection of transporter mRNAs in sieve elements. J Exp Bot 53:631–637. https://doi.org/10.1093/jexbot/53.369.631
Duan J, Bauer LS, van Driesche RG, Gould JR (2018) Progress and challenges of protecting North American ash trees from the emerald ash borer using biological control. Forests. https://doi.org/10.3390/f9030142
Ghosh SK, Hunter WB, Park AL, Gundersen-Rindal DE (2017) Double-stranded RNA delivery system for plant-sap-feeding hemipteran insects. PLoS ONE 2(12):e017186. https://doi.org/10.1371/journal.pone.0171861
Ghosh SK, Hunter WB, Park AL, Gundersen-Rindal DE (2018) Double-stranded RNA oral delivery methods to induce RNA interference in phloem and plant-sap-feeding hemipteran insects. J Vis Exp 135:e57390. https://doi.org/10.3791/57390
Government Accounting Office (GAO) (2006) Invasive forest pests: Lessons learned from three recent infestations may aid in managing future efforts. Rep. US GAO, GAO-06-353. http://www.gao.gov/assets/250/249776.pdf. Accessed 29 July 2019
Haack RA, Jendek E, Liu HP, Marchant KR, Petrice TR, Poland TM, Ye H (2002) The emerald ash borer: a new exotic pest in North America. Newsl Mich Entomol Soc 47(3–4):1–5
Herms DA, McCullough DG (2014) Emerald ash borer invasion of North America: history, biology, ecology, impacts, and management. Annu Rev Entomol 59:13–30
Hunter WB, Glick E, Paldi N, Bextine BR (2012) Advances in RNA interference: dsRNA treatment in trees and grapevines for insect pest suppression. Southwest Entomol 37:85–87
Joga MR, Zotti MJ, Smagghe G, Christiaens O (2016) RNAi efficiency, systemic properties, and novel delivery methods for pest insect control: what we know so far. Front Physiol 7:553. https://doi.org/10.3389/fphys.2016.00553
Kola VSR, Renuka P, Madhav MS, Mangrauthia SK (2015) Key enzymes and proteins of crop insects as candidate for RNAi based gene silencing. Front Physiol 6:119. https://doi.org/10.3389/fphys.2015.00119
Kyre BR, Rodrigues TB, Rieske LK (2019) RNA interference and validation of reference genes for gene expression analyses using qPCR in southern pine beetle, Dendroctonus frontalis. Sci Rep 9:5640. https://doi.org/10.1038/s41598-019-42072-6
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2-ΔΔCt method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Majidiani S, PourAbad RF, Laudani F et al (2019) RNAi in Tuta absoluta management: effects of injection and root delivery of dsRNAs. J Pest Sci 92:1409–1419. https://doi.org/10.1007/s10340-019-01097-6
Mamta B, Rajam MV (2017) RNAi technology: a new platform for crop pest control. Physiol Mol Biol Plants. 23:487–501. https://doi.org/10.1007/s12298-017-0443-x
Meister G, Tuschl T (2004) Mechanisms of gene silencing by double-stranded RNA. Nature 431:343–349. https://doi.org/10.1038/nature02873
Niu J, Yang W, Tian Y, Fan J, Ye C, Shang F, Ding B, Zhang J, An X, Yang L, Chang T, Christiaens O, Smagghe G, Wang J (2019) Topical dsRNA delivery induces gene silencing and mortality in the pea aphid. Pest Manag Sci 75:2873–2881. https://doi.org/10.1002/ps.5457
Olson DG, Rieske LK (2019) Host range expansion may provide enemy free space for the highly invasive emerald ash borer. Biol Invasions 21:625–635. https://doi.org/10.1007/s10530-018-1853-6
Palli SR (2014) RNA interference in Colorado potato beetle: steps toward development of dsRNA as a commercial insecticide. Curr Opin Insect Sci. 6:1–8. https://doi.org/10.1016/j.cois.2014.09.011
Poland TM, McCullough DG (2006) Emerald ash borer: invasion of the urban forest and the threat to North America’s ash resource. J For 104:118–124
Rajarapu SP, Mamidala P, Mittapalli O (2012) Validation of reference genes for gene expression studies in the emerald ash borer (Agrilus planipennis). Insect Sci 19:41–46. https://doi.org/10.1111/j.1744-7917.2011.01447.x
Rodrigues TB, Dhandapani RK, Duan JJ, Palli SR (2017a) RNA interference in the Asian longhorned beetle: identification of key RNAi genes and reference genes for RT-qPCR. Sci Rep 7:8913. https://doi.org/10.1038/s41598-017-08813-1
Rodrigues TB, Rieske LK, Duan JJ, Mogilicherla K, Palli SR (2017b) Development of RNAi method for screening candidate genes to control emerald ash borer, Agrilus planipennis. Sci Rep 7:7379. https://doi.org/10.1038/s41598-017-07605-x
Rodrigues TB, Duan JJ, Palli SR, Rieske LK (2018) Identification of highly effective target genes for RNAi-mediated control of emerald ash borer, Agrilus planipennis. Sci Rep 8:5020. https://doi.org/10.1038/s41598-018-23216-6
Sharp PA (2001) RNA interference. Genes Dev 15:485–490
Taning CNT, Andrade EC, Hunter WB, Christiaens O, Smagghe G (2016) Asian citrus psyllid RNAi pathway RNAi evidence. Sci Rep 6:38082. https://doi.org/10.1038/srep38082
Yu N, Christiaens O, Liu J, Niu J, Cappelle K, Caccia S, Smagghe G (2013) Delivery of dsRNA for RNAi in insects: an overview and future directions. Insect Sci 20:4–14. https://doi.org/10.1111/j.1744-7917.2012.01534.x
Zhang X, Zhang J, Zhu KY (2010) Chitosan/doublestranded RNA nanoparticle-mediated RNA interference to silence chitin synthase genes through larval feeding in the African malaria mosquito (Anopheles gambiae). Insect Mol Biol 19:683–693
Zhao C, Gonzales MAA, Poland TM, Mittapalli O (2015) Core RNAi machinery and gene knockdown in the emerald ash borer (Agrilus planipennis). J Insect Physiol 72:70–78
Acknowledgements
This publication is from the Kentucky Agricultural Experiment Station and is published with the approval of the director. This work was supported by the University of Kentucky, the USDA Forest Service Forest Health Research and Education Center, and the Kentucky Agricultural Experiment Station under McIntire-Stennis 2351197000.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Smagghe.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pampolini, F., Rodrigues, T.B., Leelesh, R.S. et al. Confocal microscopy provides visual evidence and confirms the feasibility of dsRNA delivery to emerald ash borer through plant tissues. J Pest Sci 93, 1143–1153 (2020). https://doi.org/10.1007/s10340-020-01230-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10340-020-01230-w