Abstract
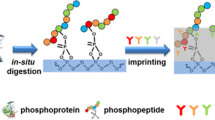
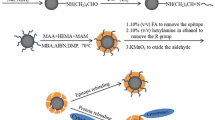
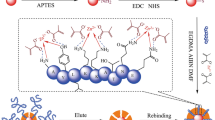
Phosphorylation is one of the most common post-translational modifications of proteins. Recognition of phosphorylated peptides with high selectivity is an important prerequisite for the structural identification of protein phosphorylation. By the application of molecular imprinting technology, a kind of tyrosine-phosphorylated peptide imprinted particles was prepared by the combination of reversible addition-fragmentation chain transfer (RAFT) polymerization and “epitope” strategy that applied in tyrosine-phosphorylated peptides recognition. Phenylphosphonic acid was used as the dummy template of the phosphorylated angiotensin II, which was one of the natural tyrosine-phosphorylated peptides. After the template modified by hydrogen bond with ureidopropyl group on the surface of silica, the surface imprinted particles with controlled and imprinted shell were synthesized by radical polymerization with RAFT strategy. The imprinted particles were obtained after the peptide removed and dithioester group destructed under alkaline condition. The binding capacity of phenylphosphonic acid reached 0.198 mg g−1 with imprinting factor (IF) as 2.70, while the binding capacity of phosphorylated angiotensin II reached 0.792 mg g−1 with IF as 1.96, which were obviously higher than with IF of that without RAFT strategy. Furthermore, phosphorylated angiotensin II could be selectively recognized by the imprinted particles even in presence of angiotensin II without phosphorylated. The performance of the phosphopeptide recognition remained 92% after five cycles of adsorption and desorption. All these results demonstrated that the tyrosine-phosphorylated imprinted particles prepared by combing RAFT polymerization and “epitope” strategy are promising to achieve the phosphopeptide recognition with higher recognition ability, selectivity and reusability.











Similar content being viewed by others
Data Availability
No datasets were generated or analysed during the current study.
References
Doyle HA, Mamula MJ (2001) Post-translational protein modifications in antigen recognition and autoimmunity. Trends Immunol 22:443–449. https://doi.org/10.1016/s1471-4906(01)01976-7
Demmers JAA, van der Wal L, Bezstarosti K (2020) Detection of protein ubiquitination sites by peptide enrichment and mass spectrometry. J Vis Exp. https://doi.org/10.3791/59079
Hauser A, Penkert M, Hackenberger CPR (2017) Chemical approaches to investigate labile peptide and protein phosphorylation. Acc Chem Res 50:1883–1893. https://doi.org/10.1021/acs.accounts.7b00170
Huang BL, Liu Y, Yao HW, Zhao Y (2020) NMR-based investigation into protein phosphorylation. Int J Biol Macromol 145:53–63. https://doi.org/10.1016/j.ijbiomac.2019.12.171
Franciosa G, Martinez-Val A, Olsen JV (2020) Deciphering the human phosphoproteome. Nat Biotechnol 38:285–286. https://doi.org/10.1038/s41587-020-0441-3
Lafite P, Daniellou R (2012) Rare and unusual glycosylation of peptides and proteins. Nat Prod Rep 29:729–738. https://doi.org/10.1039/c2np20030a
Chatterjee J, Rechenmacher F, Kessler H (2013) N-methylation of peptides and proteins: an important element for modulating biological functions. Angew Chem Int Ed 52:254–269. https://doi.org/10.1002/anie.201205674
Wang YY, Yao J, Yang PY, Deng C, Fan H (2012) Immobilization of antibodies on magnetic carbonaceous microspheres for selective enrichment of lysine-acetylated proteins and peptides. Chin J Chem 30:2549–2555. https://doi.org/10.1002/cjoc.201200542
Delom F, Chevet E (2006) Phosphoprotein analysis: from proteins to proteomes. Proteome Sci 4:1–15. https://doi.org/10.1186/1477-5956-4-15
Hunter T (2000) Signaling—2000 and beyond. Cell 100:113–127. https://doi.org/10.1016/S0092-8674(00)81688-8
Eyrich B, Sickmann A, Zahedi RP (2011) Catch me if you can: mass spectrometry-based phosphoproteomics and quantification strategies. Proteomics 11:554–570. https://doi.org/10.1002/pmic.201000489
Swaney DL, Villén J (2016) Enrichment of phosphopeptides via immobilized metal affinity chromatography. Cold Spring Harb Protoc 2016:88005. https://doi.org/10.1101/pdb.prot088005
Diez IA, Govender I, Naicker P, Stoychev S, Jordaan J, Jensen ON (2021) Zirconium(IV)-IMAC revisited: improved performance and phosphoproteome coverage by magnetic microparticles for phosphopeptide affinity enrichment. J Proteome Res 20:453–462. https://doi.org/10.1021/acs.jproteome.0c00508
Ruprecht B, Koch H, Domasinska P, Frejno M, Kuster B, Lemeer S (2017) Optimized enrichment of phosphoproteomes by Fe-IMAC column chromatography. Methods Mol Biol 1550:47–60. https://doi.org/10.1007/978-1-4939-6747-6_5
Cernigoj U, Gaspersic J, Fichtenbaum A, Lendero Krajnc N, Vidic J, Mitulovic G, Strancar A (2016) Titanium dioxide nanoparticle coating of polymethacrylate-based chromatographic monoliths for phosphopetides enrichment. Anal Chim Acta 942:146–154. https://doi.org/10.1016/j.aca.2016.08.044
Chen YM, Hoehenwarter W (2019) Rapid and reproducible phosphopeptide enrichment by tandem metal oxide affinity chromatography: application to boron deficiency induced phosphoproteomics. Plant J 98:370–384. https://doi.org/10.1111/tpj.14215
Mirza MR, Rainer M, Guzel Y, Choudhary IM, Bonn GK (2012) A novel strategy for phosphopeptide enrichment using lanthanide phosphate co-precipitation. Anal Bioanal Chem 404:853–862. https://doi.org/10.1007/s00216-012-6215-0
Han GH, Ye ML, Zhou HJ, Jiang X, Feng S, Jiang X, Tian R, Wan D, Zou H, Gu J (2008) Large-scale phosphoproteome analysis of human liver tissue by enrichment and fractionation of phosphopeptides with strong anion exchange chromatography. Proteomics 8:1346–1361. https://doi.org/10.1002/pmic.200700884
Hu YC, Li Y, Gao H, Jiang B, Zhang X, Li X, Wu Q, Liang Z, Zhang L, Zhang Y (2019) Cleavable hydrophobic derivatization strategy for enrichment and identification of phosphorylated lysine peptides. Anal Bioanal Chem 411:4159–4166. https://doi.org/10.1007/s00216-019-01770-w
Bijttebier S, Theunis C, Jahouh F, Martins DR, Verhemeldonck M, Grauwen K, Dillen L, Mercken M (2021) Development of immunoprecipitation-two-dimensional liquid chromatography-mass spectrometry methodology as biomarker read-out to quantify phosphorylated tau in cerebrospinal fluid from Alzheimer disease patients. J Chromatogr A 1651:462299. https://doi.org/10.1016/j.chroma.2021.462299
Hunter T (2015) Discovering the first tyrosine kinase. Proc Natl Acad Sci USA 112:7877–7882. https://doi.org/10.1073/pnas.1508223112
Xu L, Hu YF, Shen F, Li Q, Ren X (2013) Specific recognition of tyrosine-phosphorylated peptides by epitope imprinting of phenylphosphonic acid. J Chromatogr A 1293:85–91. https://doi.org/10.1016/j.chroma.2013.04.013
Yang XQ, Xia Y (2016) Selective enrichment and separation of phosphotyrosine peptides by thermosensitive molecularly imprinted polymers. J Sep Sci 39:419–426. https://doi.org/10.1002/jssc.201501063
Duarte M, Subedi P, Yilmaz E, Marcus K, Laurell T, Ekstrom S (2018) Molecularly imprinted polymers synthesized via template immobilization on fumed silica nanoparticles for the enrichment of phosphopeptides. J Mol Recognit 31:2677. https://doi.org/10.1002/jmr.2677
Rachkov A, Minoura N (2001) Towards molecularly imprinted polymers selective to peptides and proteins. The epitope approach. Biochim et Biophys Acta (BBA)-Protein Struct Mol Enzymol 1544:255–266. https://doi.org/10.1016/s0167-4838(00)00226-0
Yang K, Li S, Liu L, Chen Y, Zhou W, Pei J, Liang Z, Zhang L, Zhang Y (2019) Epitope imprinting technology: progress, applications, and perspectives toward artificial antibodies. Adv Mater 31:e1902048. https://doi.org/10.1002/adma.201902048
Li Q, Shen F, Zhang X, Hu Y, Zhang Q, Xu L, Ren X (2013) One-pot synthesis of phenylphosphonic acid imprinted polymers for tyrosine phosphopeptides recognition in aqueous phase. Anal Chim Acta 795:82–87. https://doi.org/10.1016/j.aca.2013.07.040
Liu QS, Zhang KN, Jin YH, Wang X, Liu Y, Liu H, Xie M (2018) Phosphate-imprinted magnetic nanoparticles using phenylphosphonic acid as a template for excellent recognition of tyrosine phosphopeptides. Talanta 186:346–353. https://doi.org/10.1016/j.talanta.2018.04.025
Zhang G, Jiang L, Zhou J, Hu L, Feng S (2019) Epitope-imprinted mesoporous silica nanoparticles for specific recognition of tyrosine phosphorylation. Chem Commun 55:9927–9930. https://doi.org/10.1039/c9cc03950c
Rouhi M, Incel A, Shinde S (2021) Role of comonomers in the recognition of anionic biomolecules in water: hydrogen-bonded imprinted polymeric receptor. ACS Appl Polymer Mater 3:4904–4912. https://doi.org/10.1021/acsapm.1c00685
Chiefari J, Chong YKB, Ercole F, Julia K (1998) Living free-radical polymerization by reversible addition−fragmentation chain transfer: the RAFT process. Macromolecules 31:5559–5562. https://doi.org/10.1021/ma9804951
Thang SH, Chong BYK, Mayadunne RTA, Moad G, Rizzardo E (1999) A novel synthesis of functional dithioesters, dithiocarbamates, xanthates and trithiocarbonates. Tetrahedron Lett 40:2435–2438. https://doi.org/10.1016/s0040-4039(99)00177-x
Li QR, Yang KG, Liang Y, Jiang B, Liu J, Zhang L, Liang Z, Zhang Y (2014) Surface protein imprinted core-shell particles for high selective lysozyme recognition prepared by reversible addition-fragmentation chain transfer strategy. ACS Appl Mater Interfaces 6:21954–21960. https://doi.org/10.1021/am5072783
Chen FF, Wang JY, Lu RC, Chen H, Xie X (2018) Fast and high-efficiency magnetic surface imprinting based on microwave-accelerated reversible addition fragmentation chain transfer polymerization for the selective extraction of estrogen residues in milk. J Chromatogr A 1562:19–26. https://doi.org/10.1016/j.chroma.2018.05.047
Kusrini E, Yusof NF, Mehamod FS, Suah FBM, Juwono FH, Yatim A, Setiawan EA (2018) The effect of RAFT polymerization on the physical properties of thiamphenicol-imprinted polymer. E3S Web Conf 67:2267–1242. https://doi.org/10.1051/e3sconf/20186703050
Kislenko E, İncel A, Gawlitza K, Sellergren B, Rurack K (2023) Towards molecularly imprinted polymers that respond to and capture phosphorylated tyrosine epitopes using fluorescent bis-urea and bis-imidazolium receptors. J Mater Chem B 11:10873–10882. https://doi.org/10.1039/d3tb01474f
Min Y, Jiang B, Wu C, Xia SM, Zhang XD, Liang Z, Zhang LH, Zhang YK (2014) 1.9 μm superficially porous packing material with radially oriented pores and tailored pore size for ultra-fast separation of small molecules and biomolecules. J Chromatogr A 1356:148–156. https://doi.org/10.1016/j.chroma.2014.06.049
Acknowledgements
This work was supported by National Natural Science Foundation of China (No. 21804099).
Funding
This work was funded by National Natural Science Foundation of China (Grant Number 21804099). Yongjian Wang, Nurimangul Muntiza, Wenbin Zhang, Qinran Li and Hongfeng Zhang declared that the funds were supported by National Natural Science Foundation of China (Grant Number 21804099). Qiliang Deng declared that there was no fund support in the research process. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Yongjian Wang, Nurimangul Muntiza and Wenbin Zhang proceed the main experiment and wrote the main manuscript text. Hongfeng Zhang established chromatographic separation methods and provided support for the use of chromatographic instruments. Dr. Qinran Li revised the whole manuscript and fnished the final draft. Prof. Qiliang Deng put forward some useful suggestions for the revision and improvement of the manuscript. All of the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Muntiza, N., Zhang, W. et al. Tyrosine-Phosphorylated Peptide Imprinted Particles Prepared by Reversible Addition–Fragmentation Chain Transfer Polymerization and “Epitope” Strategy: Selective Recognition of Phosphorylated Angiotensin II. Chromatographia (2024). https://doi.org/10.1007/s10337-024-04340-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10337-024-04340-0