Abstract
In this paper, each of the two following proteins, the angiotensin-converting enzyme 2 (ACE2) and the Main protease (Main pro) of the Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) were grafted for the first time on homemade neutravidin poly(GMA-co-EDMA) capillary columns for the research of their ligands. The effect of the column diameter on the quantity of immobilized biotinylated protein was studied. For a capillary length of 40 mm, when its internal diameter varied from 75 to 25 μm, the grafted quantity of ACE2 decreased by 85% (from 1.50 to 0.24 μg). Among all the studied ligands, a particular vigilance has been given for dexamethasone, a widely used molecule today for adult patients hospitalized with SARS-CoV-2. Competition experiments were performed with SARS-CoV-2 Receptor Binding Domain used as reference molecule with the ACE2 affinity column to assess the orthosteric binding site of dexamethasone (Dex) on ACE2. This ligand was then immobilized on Multiwall Carbon Nanotubes (Dex/MWCNT). By comparison of the normalized breakthrough curves measured for Dex and Dex/MWCNT on both the ACE2 and Main pro affinity columns, it was showed for the first time that nanovectorisation of Dex with MWCNT enhanced and stabilized its binding to both ACE2 and Main pro. This last result reinforced the use of Dex and the interest of MWCNT for boosting immune health against COVID 19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bio-affinity chromatography was employed to study biological interactions, such as protein–protein, drug–protein, enzyme–inhibitor and receptor–ligand interactions [1,2,3,4]. In the past few years, miniaturized affinity LC systems [5, 6] were developed. Nano LC columns are generally 75 μm in diameter, because the particle sizes for packed Nano columns were between 3 and 5 μm [1].
The alternative to the particle packed columns were Poly glycidyl methacrylate-co-ethylene dimethacrylate) (Poly (GMA-co-EDMA)) monoliths columns [7,8,9,10,11]. So as to decrease strongly the time required for the immobilization procedure of the receptor in the capillary column, neutravidin (or streptavidin) [12,13,14,15] has been first grafted on the organic monolith surface. This neutravidin (or streptavidin) poly(GMA-co-EDMA) monolithic LC capillary column can be stored for at least 3 months at 4 °C and used just before the grafting of the biotinylated receptor which took only a few minutes. A such immobilization process with Nano LC capillary columns has thus been used for human serum albumin (HSA) [12], arginase [13], SARS-CoV-2 Receptor Binding Domain (RBD) [14] and acetylcholinesterase (AChE) [15] for the purpose of the research of their potential ligands.
In this work, homemade nano-organic neutravidin monolithic LC capillary columns were used for the first time for the grafting of (1) the angiotensin-converting enzyme 2 (ACE2) which was the host functional receptor for the SARS CoV2 causing COVID19 [16] and (2) the SARS-CoV-2-3CL-Mpro proteinase which played a critical role in viral protein maturation by cleaning proproteins after their translation into the host cell cytosol [17].
Angiotensin-converting enzyme 2 (ACE2) consists of 805 amino acids and is a type I membrane protein expressed in lung heart kidney and intestine [18]. The full length ACE2 consists of an N terminal peptidase domain (PD) (residues Ser19–Asp615) and a C terminal collectrin-like domain (CLD) that ends with a transmembrane alpha helix and an approximately 40-residue intracellular segment [18].
In the case of SARS-CoV-2, the glycoprotein (S protein) on the virion surface exploits ACE2 for host infection [19]. It was demonstrated that the ectodomain of SARS-CoV-2-S protein binds to the PD of ACE2 with a dissociation constant Kd = 15 nM [20].
The 3CL Mpro proteinase is composed of Domain I (residues 8–101), Domain II (residues 102–184) and Domain III (residues 201–303). Domains I and II are composed of antiparallel beta barrel structures and are the catalytic structures. Domain III is composed of five alpha helices and plays an essential role in the protease function [21].
Investigations proved that low doses of dexamethasone could reduce the mortality in patients with severe COVID-19. The binding mechanism of the dexamethasone ligand to these two receptors has been particularly scrutinized. Carbon nanotubes were promising agents to increase the binding of some ligands to their molecular targets [22]. Thus, the difference on the binding to ACE2 or Mainpro of Dexametasone and its nanovector (i.e., Dex/MWCNT) was studied in this paper for the first time.
Materials and Methods
Equipment
The ThermoScientific Ultimate 3000 RSLCNano system with a UV detection was used.
The poly(GMA-co-EDMA) monoliths were prepared as explained in our previous work [23]. This monolith had a specific area of 4 m2 g−1 and a permeability K = 6.10–14 m2. The medium size for macropores was approximatively 2.0 μm and the skeleton size was approximatively 1.0 μm. The grafting process of neutravidin onto the surface of these monoliths was previously described by the use of the Schiff base method [12,13,14,15]. These neutravidin poly(GMA-co-EDMA) monolith capillary columns (75 μm or 25 μm i.d.) were stable at least 3 months at 4 °C and can be used just before the fast immobilization process of the biotinylated receptor (ACE2 or Main pro).
Reagents
Dexamethasone, chloroquine, hydroxychloroquine, isorhamnetin, quercetin, phenylalanine, chlorogenic acid, piceatannol, kaempferol, rutin and Polyethylene Glycol—4000 (PEG 4000) were obtained from Sigma Aldrich. The recombinant angiotensin-converting enzyme 2 (ACE2), the SARS-CoV-2-Main protease (Main pro) and the SARS-CoV-2-Receptor Binding Domain (RBD) were purchased from Acrobiosystem. Short length multiwall carbon nanotubes (median length 600 nm) were obtained from Nanothinx (Greece). All the other chemical products were of analytical grade and all the buffer solutions were filtered through a 0.45 µm membrane filter and degassed before their use for HPLC.
The receptor (ACE2 or Main pro) was biotinylated with a molar ratio RBD/biotin = 2 with the EZ-link NHS-Biotin Reagent (Thermoscientific) following the manufacturer's protocol.
Preparation of Dex/MWCNT Nanovector
The Dex/MWCNT nanovector was prepared as explained in a previous work [24]. MWCNTs were first suspended in a 3:1 mixture of concentrated sulfuric and nitric acids (96% and 69%, respectively) for 4 h. The resulting oxidized nano MWCNT (oMWCNTs) were collected through a 0.2 μm PTFE membrane, and washed with water until reaching neutral pH and dried under vacuum for 2 h. Then, 6 mg of oMWCNTs and 6 mg of PEG 4000 were mixed and dispersed in 100 mL of deionized water. The mixture was sonicated in a water bath and stirred at room temperature for 60 min. 100 mL of Dex at 30 μM was thus added in this solution for 2 h. The Dex/MWCNT particles were collected through a 0.45 μm Millipore filter rinsed with water and dried under vacuum for 4 h. The mass fraction of Dex (0.096 (m/m)) in the Dex/MWCNT nanovector was inferred from the remaining free Dex contained in the solution and calculated using UV absorbance of Dex at 238 nm (EDex = 0.039 L mg−1 cm−1).
Frontal Analysis Experiments
Different concentrations of potential ligands (L) were prepared in the 50 mM PBS pH 7.4 and were flushed at a fixed flow-rate through our prepared miniaturized affinity capillary columns (ACE2 or Main pro affinity column). All the experiments were repeated three times.
Sigmoidal breakthrough curves were obtained and studied by the use of the frontal analysis equation [25]:
In this equation, Q was the number of moles of ligands (L) needed to reach the midpoint of the sigmoidal curve, Kd was the dissociation constant of the ligand to its studied receptor and mL was the total moles of bound receptors on the affinity column.
The dead time t0 of the affinity column was determined by the injection of DMSO in the chromatographic system.
For some experiments the breakthrough curves were given in the Normalized format.
In this format the y-axis described the ratio of the UV absorbance of the ligand at the column outlet to that of the UV absorbance of the ligand at the column inlet (Abs. Normalized) with values between 0 and 1. In the breakthrough curve, the stable plateau formed at saturation time tp was characterized by an Abs. Normalized = 1.
The x-axis described the ratio of the saturation tp of the plateau to the dead time of the affinity column (Normalized.Time; tP/t0).
When the ligand displayed no interactions with the affinity column, the Normalized breakthrough time tP/t0 → 1 [26].
Biotinylated-Receptor Immobilization on the Neutravidin Poly(GMA-co-EDMA) Monolithic LC Capillary Columns
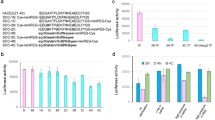
For the receptor ACE2, a biotinylated—ACE2 solution with a concentration of 10 μM flushed through our neutravidin column (40 mm length, 75 μm internal diameter) in a 50 mM PBS pH 7.4 at 300 nL min−1 and 280 nm produced a sigmoidal curve with a breakthrough time equal to tp = 6.36 min due to the strong and rapid noncovalent biotin–neutravidin interaction (Kd = 10–15 M). The column dead time t0 at the same chromatographic conditions measured by the breakthrough time of DMSO was 0.45 min. The corresponding reduced breakthrough time (tr = tp − t0) was thus equal to 5.91 min. These values corresponded to a total mole of bound receptors on the ACE2 column equal to mACE2 = 300.10–9*5.91*10*10–6 = 17.7 × 10–12 mol = 17.7 pmol. For a set of 6 columns, around 18.00 pmol of RBD were thus immobilized (18.00 ± 0.31 pmol, n = 6) which corresponded to 1.50 μg of ACE2 (MW 85 kDa) for a column length of 40 mm with a 75 μm i.d.
The same procedure was used for the immobilisation of biotinylated-ACE2 onto the neutravidin column with the same length (40 mm) but with an internal diameter of 25 μm. The concentration of the flushed biotinylated-ACE2 solution was 15 μM in the 50 mM PBS pH 7.4 and the flow rate was fixed at 30 nL min−1. The corresponding reduced breakthrough time was 6.24 min. These values corresponded now to a mACE2 value equal to 30.10–9*6.24*15*10–6 = 2.80 × 10–12 mol = 2.80 pmol.
For a set of 6 columns, around 2.80 pmol of ACE2 were thus immobilized (2.80 ± 0.09 pmol, n = 6) which corresponded to 0.24 μg of ACE2 (MW 85 kDa) for a column length of 40 mm with a 25 μm i.d.
The use of a column with an internal diameter of 25 μm allowed to decrease strongly the quantity of grafted protein on the chromatographic support (86%). Therefore, the Main pro receptor was grafted onto the neutravidin poly(GMA-co-EDMA) monolith inside a capillary column with a 25 μm internal diameter. The concentration of the flushed biotinylated-main pro solution was 10 μM in the 50 mM PBS pH 7.4 and the flow rate was fixed at 30 nL min−1. The corresponding reduced breakthrough time was 5.33 min. These values corresponded now to a mMain pro value equal to 30.10–9*5.33*10*10–6 = 1.60 × 10–12 mol = 1.60 pmol. The quantity of immobilized Main pro was (1.60 ± 0.05 pmol, n = 6).
Results and Discussion
Research of Ligand Targeting ACE2
Typical breakthrough curves were obtained for a series of potential ligands of ACE2 (including dexamethasone) by the use of the two capillary columns with 25 μm and 75 μm internal diameter.
A solution of the ligand at a given concentration (in the range 2–50 μM) was flushed in a 50 mM PBS pH 7.4 through the miniaturized column. The column temperature was 25 °C and the detection wavelength was 254 nm for all the studied ligands (except for dexamethasone λ = 238 nm). The column length was 40 mm. The flow rate was equal to 200 nL min−1 and 30 nL min−1 for a column diameter, respectively, equal to 75 μm and 25 μm.
An example is given in Fig. 1A. The curves 1/Q versus 1/[L] were drawn for each ligand and were linear (r2 > 0.997). For example, the linear equation obtained for quercetin with the column of 75 μm i.d. was 1/Q = 2.876 105/[quercetin] + 5.639 1010 (r2 = 0.9981). From their slopes and intercepts, the corresponding values of mL and Kd for each ligand were calculated (Table. 1). For each ligand, the Kd values obtained on the two columns were similar and chloroquine and dexamethasone described similar Kd values as those obtained by cell chromatography (Kd = 1.30 μM) for chloroquine [27] or by surface plasmon resonance (Kd = 9.0 μM) [28] for dexamethasone. These results confirmed the excellent efficiency of our immobilization protocol of the ACE2 receptor onto the organic monolith.
Normalized breakthrough curve of quercetin solution at four concentration on the ACE2 affinity column (A) and the corresponding Normalized breakthrough curve on the control column (with no ACE2 immobilized) (B). The column diameter and length were 75 μm and 40 mm, respectively. Mobile phase: 50 mM PBS pH 7.4. Flow-rate: 200 nL/min. Column temperature: 25 °C. Detection wavelength: 254 nm
As well, the normalized breakthrough times obtained on our 75 μm and 25 μm neutravidin poly(GMA-co-EDMA) monolithic LC capillary columns (i.e., with no immobilized ACE2) was the same upon ligand concentration with a tplateau/to → 1 (1.20) (Fig. 1B) showing that non-specific interaction during the binding process remained negligible [29].
For fragment-based drug discovery (FBDD), competition experiments are very useful to confirm the receptor binding site of a fragment. So as to demonstrate that our homemade miniaturized columns can be used for a such purpose, normalized breakthrough curves were compared in the absence and presence of the competition agent in the percolated solution inside the miniaturized capillary column (25 um, id 40 mm length).
This was highlighted for dexamethasone in the presence of the competitive agent SARS-CoV-2 RBD which bind to ACE2 with an equilibrium dissociation constant in the nanomolar range [30]. A solution of dexamethasone (5 μM) was flushed in a 50 mM PBS pH 7.4 through successively (1) the neutravidin column (i.e., control column with no ACE2), (2) the ACE2 column in the presence of 5 μM of the SARS-CoV-2 RBD competition agent and (3) the ACE2 column without competition agent. The flow-rate was 30 nL min−1, the column temperature was 25 °C and the detection wavelength was 238 nm. The corresponding normalized breakthrough curves are given in Fig. 2.
Normalized breakthrough curves of a dexamethasone solution (5 μM) on the: A Neutravidin column (i.e., control column with no ACE2). B ACE2 affinity column with the SARS-CoV-2 RBD competition agent of high affinity. C ACE2 affinity column without competition agent. The column diameter and length were 25 μm and 40 mm, respectively. Mobile phase: 50 mM PBS pH 7.4. Flow-rate: 30 nL/min. Column temperature: 25 °C. Detection wavelength: 238 nm
The normalized breakthrough time of Dex decreased strongly when SARS-CoV-2 RBD was added in the percolated solution and close to the value obtained on the Neutravidin column (control column with no ACE2) (tp/to → 1). This result confirmed that dexamethasone bound on the orthosteric site of ACE2 with specific interaction.
Research of Ligand Targeting the Main Pro of SARS-CoV-2
The inhibition of viral proteases can reduce the assembly of mature viral particles [13] and can thus reduce the COVID19 infection. Works remained to be done on developing rapid analytical method in searching for drugs inhibiting the Main protease (Main pro) of SARS-CoV-2. For this purpose, normalized breakthrough times were compared for two extreme drug concentrations. Indeed, if a molecule was a ligand of Main pro, at a very high concentration of this molecule, the total binding sites onto the Main pro capillary column are thus occupied and the breakthrough time of this molecule measured on the Main pro immobilized column was obviously close to the one measured on the Neutravidin control column (with no immobilized Main pro). This made it possible to avoid changing the column during experiments (time saving) and potential uncertainties of the measures on the control column. This strategy was used for the first time for the study of the binding of 5 potential ligands to Main pro.
A solution of the drug at a low concentration (10 μM) and a high concentration (1400 μM) was thus successively flushed through our Main pro column (40 mm length, 25 μm internal diameter) in a 50 mM PBS pH 7.4 at 30 nL min−1, at a column temperature equal to 25 °C with a detection wavelength equal to 254 nm.
Phenylalanine described identical breakthrough times at a low (5 μM) and a very high concentration (1400 μM) and were thus non-ligands for Main pro (Fig. 3). On the other hand, significant differences between breakthrough times were observed at these two extreme concentrations for four molecules (Fig. 3). When the normalized breakthrough time increased the affinity of the ligand to the receptor increased. These compounds have thus an affinity for Main pro in the sequence chlorogenic acid < piceatannol < kaempferol < rutin (Fig. 3). This result was in good agreement with a previous result demonstrating that rutin was a ligand of Main Pro [31].
Normalized breakthrough curve of 5 compounds at a low and a high-level concentration in PBS on the homemade Main pro column. The column diameter and length were 25 μm and 40 mm, respectively. Mobile phase: 50 mM PBS pH 7.4. Flow rate: 30 nL/min. Column temperature: 25 °C. Detection wavelength: 254 nm
Role of Carbon Nanotube on the Dexametasone/Receptor Binding
Carbon nanotubes have been recognized as promising nanocarriers for an extended range of molecules [22]. For the moment, the corticosteroid dexamethasone is a widely used molecule for adult patients hospitalized with COVID-19 [32] due to its binding to both ACE2 and Main pro receptors of SARS CoV-2.
A solution of dexamethasone (5 μM) and a solution of Dex/MWCNT (5 μM) were successively flushed at a flow rate of 30 nL min−1 and 25 °C in a 50 mM PBS pH 7.4 through (1) the Neutravidin column (i.e., control column with no ACE2), (2) the ACE2 column, and (3) the Main Pro column. The column diameter and length were, respectively, equal to 25 μm and 40 mm and the detection wavelength was 238 nm. The corresponding normalized breakthrough curves are given in Fig. 4.
Normalized breakthrough curves of Dex and Dex/MWCNT solutions (5 μM) on the A Neutravidin column (control column), B ACE2 affinity column and C main Pro affinity column. The column diameter and length were 25 μm and 40 mm, respectively. Mobile phase: 50 mM PBS pH 7.4. Flow-rate: 30 nL/min. Column temperature: 25 °C. Detection wavelength: 238 nm
Dex and Dex/MWCNT described identical breakthrough times on the neutravidin poly(GMA-co-EDMA) capillary column with (t/t0 → 1) (Fig. 4A) showing very weak non-specific interactions with the chromatographic support on absence of ACE2 and Main pro.
In opposition, the normalized breakthrough curves for Dex and MWCNT/Dex obtained on both the ACE2 column (Fig. 4B) and on the Main pro column (Fig. 4C) described normalized breakthrough times values well above 1 and always the highest for the nanovector.
The shape of the breakthrough curve was less steep on the Main pro column than on the ACE2 column. This can be explained by the fact that linear chromatographic conditions were favored on the Main Pro column [33]. The equivalent in zonal chromatography would be a more symmetrical peak obtained on the Main Pro column. In addition, these results suggested that the Kd dissociation constant values for Dex and its nanovector for Main pro became appreciable relative to the concentration of the ligand at the column inlet (i.e., Kd > > 5 μM) [33].
These results demonstrated that the dexamethasone ligand and its nanovector were binders for both ACE2 and Main Pro with specific interactions. As well, it was showed that the nanovectorisation of dexamethasone with MWCNT enhanced and stabilized its recognition mechanism with ACE2 and Main pro and may reinforced its efficiency in clinical trial against COVID19.
Conclusions
Here, we have showed for the first time, that our homemade neutravidin poly(GMA-co-EDMA) capillary columns, can be used for distinguish ligand or non-ligands of the ACE2 and the Main pro of the Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2). Competition experiments were carried out to confirm the binding of dexamethasone to the orthosteric site of ACE2 by the use of SARS-CoV-2 RBD reference molecule. As well, and for the first time, by comparison of the normalized breakthrough curves of dexamethasone and its nanovector obtained on the ACE2 and Main pro affinity columns, it was demonstrated that the nanovector stabilized and reinforced the dexamethasone binding to ACE2 and Main pro. This nanovector could be thus a useful tool to increase the therapeutic efficiency of dexamethasone against COVID19.
References
Reichelt S (ed) (2015) Affinity chromatography methods in molecular biology. Springer, New York
Bi C, Beeram S, Li Z, Zheng X, Hage DS (2015) Kinetic analysis of drug protein interactions by affinity chromatography. Drug Discov Today Technol 17:16–21
Roque ACA, Lowe CR (2008) Affinity chromatography: history, perspectives, limitations and prospects. Method Mol Biol 421:1–21
Hage DS (2017) Analysis of biological interactions by affinity chromatography: clinical and pharmaceutical applications. Clin Chem 63:1083–1093
Wilson SR, Olsen C, Lundanes E (2019) Nano liquid chromatography columns. Analyst 144:7090–7104
Aydoga C, Rigano F, Krcmova LK, Chung DS, Macka M, Mondello L (2020) Miniaturized LC in molecular omics. Anal Chem 92(17):11485–11497
Courtois J, Szumski M, Bystrom B, Iwasiewicz A, Shchukarev A, Irgum K (2006) A study of surface modification and anchoring techniques used in the preparation of monolithic microcolumns in fused silica capillaries. J Sep Sci 29(1):14–24
Bing Y, Hongbo Z, Hailin C, Guihuan C, Tao X, Yangchun L (2017) Recent development and application of monolithic columns. Rev Adv Mater Sci 48:58–67
Frantisek S (2010) Porous polymer monoliths amazing wide variety of techniques enabling their preparation. J Chromatogr A 1217(6):902–924
Candish E, Khodabandeh A, Gaborieau A, Rodemann T, Shellie RA, Gooley AA (2017) Poly(ethylene glycol) functionalization of monolithic poly (divinyl benzene) for improved miniaturized solid phase extraction of protein-rich samples. Anal Bioanal Chem 409(8):2189–2199
Darrio Arrua R, Talebi M, Causon TJ, Hilder EF (2012) Review of recent advances in the preparation of organic polymer monoliths for liquid chromatography of large molecules. Anal Chim Acta 738:1–12
Andre C, Guillaume YC (2020) An organic monolithic capillary column functionalized with human serum albumin and its application for the nano-chromatography study of its binding with universal cancer peptides and its impact on immunogenicity. J Liq Chromatogr Relat Technol 43(17–18):1–7
Andre C, Guillaume YC (2021) Development of a new arginase capillary column for the fast screening of arginase inhibitors and evaluation of their binding affinity. J Chromatogr B 1175(9):122751
Guillaume YC, Andre C (2022) Immobilization of the SARS-CoV-2-receptor binding domain onto methacrylate-based monolith for nano LC at 30 nL min−1 and application for research of its ligands. Anal Method 14:156–164
Andre C, Guillaume YC (2022) Development of nano bio LC columns for the search of acethylcholinesterase molecular targets. J Sep Sci. https://doi.org/10.1002/jssc.202200047
Lan JL, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang SX, Wang Q, Zhang L, Wang X (2020) Structure of the SARS-CoV-2 spike receptor binding domain bound to the ACE2 receptor. Nature 581:215–220
Anirudhan V, Lee H, Cheng H, Cooper L, Rong L (2021) Targeting SARS-CoV-2 viral proteases as a therapeutic strategy to treat COVID-19. J Med Virol 93(5):2722–2734
Perlot T, Penninger JM (2013) ACE2-from the renin-angiotensin system to gut microbiota and malnutrition. Microbes Infect 15(6):866–873
Zhang Q, Xiang R, Huo S, Zhou Y, Jiang S, Wang Q, Yu F (2021) Molecular mechanism of interaction between SARS-CoV-2 and host cells and interventional therapy. Sign Transduct Target Ther 6(1):233. https://doi.org/10.1038/s41392-021-00653-w
Yan R, Zhang Y, Li Y, Xia L, Guo Y, Zhou Q (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367(6485):1444–1448
Kneller DW, Phillips G, O’Neill HM, Jedrzejczak R, Stols L, Langan P, Joachimiak A, Coates L, Kovalevsky A (2020) Structural plasticity of SARS CoV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography. Nat Commun 11(1):3202. https://doi.org/10.1038/s41467-020-16954-7
Anzar N, Hasan R, Tyagi M, Ydav N, Narang J (2020) Carbon nanotube—a review on synthesis, properties and plethora of applications in the field of biomedical science. Sens Int 1:100003. https://doi.org/10.1016/j.sintl.2020.100003
Andre C, Guillaume YC (2021) Development of nano affinity columns for the study of ligand (including SARS-CoV-2 related proteins) binding to heparan sulfate proteoglycans. Anal Methods 13:3050–3058
Haghigi M, Khoshfetrat (2016) Carbon nanotubes as an appropriate dexamethasone carrier in the presence of PEG. International Congress of Nanoscience and Nanotechnology. Research gate.net/publication/334451172.
He X, Sui Y, Wang S (2018) Stepwise frontal affinity chromatography model for drug and protein interaction. Anal Bioanal Chem 410:5807–5815
Renaud JP, Chung C, Danielson UH, Egner U, Hennig M, Hubbard RE, Nar H (2016) Biophysics in drug discovery: impact, challenges and opportunities. Nat Rev Drug Discov 15:679–698
Fu J, Jia Q, Zhou H, Zhang L, Wang S, Liang P, Lv Y, Han, (2021) Cell membrane chromatography for the analysis of the interaction between chloroquine and hydroxychloroquine with ACE2 receptor. J Chromatogr B 1162:122469
Zhang Y, Hu S, Wang J, Xue Z, Wang C, Wang N (2021) Dexamethasone inhibits SARS-CoV-2 spike pseudotype virus viropexis by binding to ACE2. Virology 554:83–88
Ohlson S, Duong-Thi MD (2018) Fragment screening for drug leads by weak affinity chromatography (WAK-MS). Methods 146:26–38
Yang J, Petitjean SJL, Alsteens D (2020) Molecular interaction and inhibition of SARS-CoV-2 binding to the ACE2 receptor. Nat Commun 11(4541):1–10. https://doi.org/10.1038/s41467-020-18319-6
Xu Z, Yang L, Zhang X, Zhang Q, Yang Z, Liu Y, Wei S, Liu W (2020) Discovery of potential flavonoid inhibitors against COVID-19 3CL proteinase based on virtual screening strategy. Front Mol Biosci 7(5556481):1–8. https://doi.org/10.3389/fmolb.2020.556481
The RECOVERY collaborative group (2021) Dexamethasone in hospitalized patients with Covid 19. N Engl J Med 384:693–704. https://doi.org/10.1056/NEJMoa2021436
Schriemer DC (2004) Biosensor alternative—frontal affinity chromatography. Anal Chem 76(23):440A-448A
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Yves Claude Guillaume has no conflict of interest regarding the publication of this article. Claire André has no conflict of interest regarding the publication of this article.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Guillaume, Y.C., André, C. ACE2 and SARS-CoV-2-Main Protease Capillary Columns for Affinity Chromatography: Testimony of the Binding of Dexamethasone and its Carbon Nanotube Nanovector. Chromatographia 85, 773–781 (2022). https://doi.org/10.1007/s10337-022-04181-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-022-04181-9