Abstract
Lipids represent a very diverse group of compounds with a high variety of physicochemical properties determining their functional activities. For omics-wide identification of lipid species from complex biological samples, several crucial analytical steps including extraction, chromatographic separation and mass spectrometry analysis need to be carefully considered and validated. Here we review applications of three main chromatography techniques—reversed phase, normal phase and hydrophilic interaction liquid chromatography—for analysis of complex natural lipidomes aiming to uncover the diversity of lipid species. To correlate lipid separation with their physicochemical properties, lipid chemical space was reconstructed and used to explain principles underlying different chromatographic techniques. Furthermore, examples of available methods for analysis of complex natural lipidomes characterized by high diversity and dynamic range of lipid concentrations are illustrated for adipose tissue lipidomics.
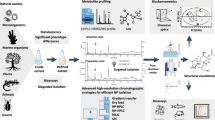
Graphical Abstract






Similar content being viewed by others
Abbreviations
- 1,2-DAG:
-
1,2-Diacylglycerol
- 1,3-DAG:
-
1,3-Diacylglycerol
- 1-LPC:
-
1-Lysophosphatidylcholine
- 1-LPG:
-
1-Lysophosphatidylglycerol
- 1-MAG:
-
1-Monoacylglycerol
- 2-LPC:
-
2-Lysophosphatidylcholine
- 2-LPE:
-
2-Lysophosphatidylethanolamine
- 2-LPG:
-
2-Lysophosphatidylglycerol
- Ag-HPLC:
-
Argentation/silver ion HPLC
- AmFm:
-
Ammonium formate
- AmOAc:
-
Ammonium acetate
- ASA:
-
Accessible surface area
- ASG:
-
Acylated steryl glycosides
- AT:
-
Adipose tissue
- BEH:
-
Ethylene bridged hybrid
- C18:
-
Octadecyl derivatized silica
- C1P:
-
Ceramide-1-phosphate
- C30:
-
Triacontanyl derivatized silica
- C4:
-
Butyl derivatized silica
- C8:
-
Octyl derivatized silica
- CE:
-
Cholesteryl esters
- Cer:
-
Ceramide
- Cer-NOH:
-
Non-hydroxy fatty acid ceramides
- Cer-OH:
-
Hydroxy fatty acid ceramide
- CHCl3:
-
Chloroform
- Chol:
-
Free cholesterol
- CL:
-
Cardiolipin
- CMNH:
-
Charge modulated hydroxyethyl amide HILIC
- CN:
-
Cyanopropyl
- CS:
-
Cholesteryl sulfate
- cyHex:
-
Cyclohexane
- DAG:
-
Diacylglycerol
- DCM:
-
Dichloromethane
- DDA:
-
Data-dependent acquisition
- DGCC:
-
Diacylglycerol-carboxymethylcholine
- DGDG:
-
Digalactosyldiacylglycerols
- DGTS:
-
Diacylglycerol-trimethylhomoserin
- dhCer1P:
-
Dihydro-ceramide-1-phosphate
- Dimet-SPH:
-
Dimethyl-sphingosine
- Diol:
-
Dihydroxy-propyl
- ECN:
-
Equivalent carbon number
- ESI:
-
Electrospray ionization
- EtOAc:
-
Ethyl acetate
- EtOH:
-
Ethanol
- FA:
-
Formic acid
- FAME:
-
Fatty acid methyl esters
- FFA:
-
Free fatty acid
- FPP:
-
Fully porous particles
- GalCer:
-
Galactosylceramide
- GalCer-NOH:
-
Non-hydroxy galactosylceramide
- GlcCer:
-
Glucosylceramide
- GM1:
-
Ganglioside
- H2O:
-
Water
- H3PO4 :
-
Phosphoric acid
- Hept:
-
Heptane
- Hex:
-
Hexane
- HexCer:
-
Hexosylceramide
- HILIC:
-
Hydrophilic interaction chromatography
- HOAc:
-
Acetic acid
- i-Hex:
-
Iso-hexane
- i-Oct:
-
Iso-octane
- IPA:
-
Isopropanol
- IPC:
-
Inositol-phosphoceramide
- LAA:
-
Lipoamino acid
- LacCer:
-
Lactosylceramide
- LC–MS:
-
Liquid chromatography–mass spectrometry
- LPA:
-
Lysophosphatidic acid
- LPA:
-
Lysophosphatidic acid
- LPC:
-
Lyso-phosphatidylcholine
- LPE:
-
Lysophosphatidylethanolamine
- LPG:
-
Lysophosphatidylglycerol
- LPI:
-
Lysophosphatidylinositol
- MAG:
-
Monoacylglycerol
- MB:
-
Mobile phase
- MD:
-
Molecular descriptor
- MeCN:
-
Acetonitrile
- MeDAG:
-
Monoalkyl diacylglycerol
- MeOH:
-
Methanol
- MGDG:
-
Monogalatosyldiacylglycerols
- MS:
-
Mass spectrometry
- MTBE:
-
Methyl-tert-butyl-ether
- NAPE:
-
N-acylphosphatidylethanolamine
- NH3 :
-
Ammonia
- NMM:
-
N-methylmorpholine
- NPC:
-
Normal phase chromatography
- PA:
-
Phosphatidic acid
- PC:
-
Phosphatidylcholine
- PC1:
-
Principal component 1
- PC2:
-
Principal component 2
- PCA:
-
Principal component analysis
- PE:
-
Phosphatidylethanolamine
- PG:
-
Phosphatidylglycerol
- Phyto-SPH:
-
Phytosphingosine
- PI:
-
Phosphatidylinositol
- P-PC:
-
Plasmalogen-phosphatidylcholine
- P-PE:
-
Plasmalogen-phosphtadiylethanolamine
- PS:
-
Phosphatidylserine
- PVA:
-
Polyvinyl alcohol
- RP:
-
Reversed phase
- RPC:
-
Reversed phase chromatography
- S1P:
-
Sphingosine-1-phosphate
- Sa1P:
-
Sphinganine-1-phosphate
- SCP:
-
Solid core particles
- SFC:
-
Supercritical fluid chromatography
- SG:
-
Steryl glycosides
- Si:
-
Silica
- Si-H:
-
Silica hydride
- SM:
-
Sphingomyelin
- SP:
-
Stationary phase
- SPA:
-
Sphinganine
- SPH:
-
Sphingosine
- SPH:
-
Sphingosine
- SQ:
-
Squalene
- SQDG:
-
Sulfoqunovosyl diacylglycerol
- ST:
-
Sulfatide
- TAG:
-
Triacylglycerols
- TEA:
-
Trimethylamine
- TFA:
-
Trifluoro acetic acid
- THF:
-
Tetrahydrofurane
- Trimet-SPH:
-
Trimethyl-sphingosine
- VAT:
-
Visceral adipose tissue
- WAT:
-
White adipose tissue
References
Fahy E et al (2009) Update of the LIPID MAPS comprehensive classification system for lipids. J Lipid Res 50:S9–S14
van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9:112–124
Karantonis HC, Nomikos T, Demopoulos CA (2009) Triacylglycerol metabolism. Curr Drug Targets 10:302–319
Ahmadian M, Duncan RE, Jaworski K, Sarkadi-Nagy E, Sul HS (2007) Triacylglycerol metabolism in adipose tissue. Future Lipidol 2:229–237
Wymann MP, Schneiter R (2008) Lipid signalling in disease. Nat Rev Mol Cell Biol 9:162–176
Masoodi M, Kuda O, Rossmeisl M, Flachs P, Kopecky J (2015) Lipid signaling in adipose tissue: connecting inflammation and metabolism. Biochim Biophys Acta Mol Cell Biol Lipids 1851:503–518
Yetukuri L, Ekroos K, Vidal-Puig A, Oresic M (2008) Informatics and computational strategies for the study of lipids. Mol Biosyst 4:121–127
Řezanka T, Kolouchová I, Gharwalová L, Palyzová A, Sigler K (2018) Lipidomic analysis: from archaea to mammals. Lipids 53:5–25
Rampler E et al (2018) A novel lipidomics workflow for improved human plasma identification and quantification using RPLC-MSn methods and isotope dilution strategies. Anal Chem 90:6494–6501
Gao X et al (2012) A reversed-phase capillary ultra-performance liquid chromatography–mass spectrometry (UPLC–MS) method for comprehensive top-down/bottom-up lipid profiling. Anal Bioanal Chem 402:2923–2933
Peng B et al (2018) Identification of key lipids critical for platelet activation by comprehensive analysis of the platelet lipidome. Blood. https://doi.org/10.1182/blood-2017-12-822890
Carvalho M et al (2012) Effects of diet and development on the Drosophila lipidome. Mol Syst Biol 8:600
Hummel J et al (2011) Ultra performance liquid chromatography and high resolution mass spectrometry for the analysis of plant lipids. Front Plant Sci 2:54
Schwudke D, Liebisch G, Herzog R, Schmitz G, Shevchenko A (2007) Shotgun lipidomics by tandem mass spectrometry under data-dependent acquisition control. Methods Enzymol 433:175–191
Schwudke D et al (2006) Lipid profiling by multiple precursor and neutral loss scanning driven by the data-dependent acquisition. Anal Chem 78:585–595
Tumanov S, Kamphorst JJ (2017) Recent advances in expanding the coverage of the lipidome. Curr Opin Biotechnol 43:127–133
Cajka T, Fiehn O (2014) Comprehensive analysis of lipids in biological systems by liquid chromatography–mass spectrometry. Trends Anal Chem 61:192–206
Instant JChem 18.8.0. (2018) Instant JChem was used for the prediction of properties from lipid structures. http://www.chemaxon.com. Accessed 1 Aug 2018
Dorsey JG, Cooper WT (1994) Retention mechanisms of bonded-phase liquid chromatography. Anal Chem 66:857A–867A
Vailaya A (2005) Fundamentals of reversed phase chromatography: thermodynamic and exothermodynamic treatment. J Liq Chromatogr Relat Technol 28:965–1054
Layne J (2002) Characterization and comparison of the chromatographic performance of conventional, polar-embedded, and polar-endcapped reversed-phase liquid chromatography stationary phases. J Chromatogr A 957:149–164
Ovčačíková M, Lísa M, Cífková E, Holčapek M (2016) Retention behavior of lipids in reversed-phase ultrahigh-performance liquid chromatography-electrospray ionization mass spectrometry. J Chromatogr A 1450:76–85
Háková E et al (2015) Localization of double bonds in triacylglycerols using high-performance liquid chromatography/atmospheric pressure chemical ionization ion-trap mass spectrometry. Anal Bioanal Chem 407:5175–5188
Nakanishi H, Iida Y, Shimizu T, Taguchi R (2010) Separation and quantification of sn-1 and sn-2 fatty acid positional isomers in phosphatidylcholine by RPLC-ESIMS/MS. J Biochem 147:245–256
Witting M, Maier TV, Garvis S, Schmitt-Kopplin P (2014) Optimizing a ultrahigh pressure liquid chromatography-time of flight-mass spectrometry approach using a novel sub-2 µm core-shell particle for in depth lipidomic profiling of Caenorhabditis elegans. J Chromatogr A 1359:91–99
Bird SS, Marur VR, Stavrovskaya IG, Kristal BS (2012) Separation of cis–trans phospholipid isomers using reversed phase LC with high resolution MS detection. Anal Chem 84:5509–5517
Narváez-rivas M, Zhang Q (2016) Comprehensive untargeted lipidomic analysis using core–shell C30 particle column and high field orbitrap mass spectrometer. J Chromatogr A 1440:123–134
Norman HA, St John JB (1986) Separation and quantitation of molecular species of plant phosphatidylcholine by high-performance liquid chromatography with flame ionization detection. J Lipid Res 27:1104–1107
Lee C, Fisher SK, Agranoff BW, Hajra AK (1991) Quantitative analysis of molecular species of diacylglycerol and phosphatidate formed upon muscarinic receptor activation of human SK-N-SH neuroblastoma cells. J Biol Chem 266:22837–22846
Ousley AH, Morell P (1992) Individual molecular species of phosphatidylcholine and phosphatidylethanolamine in myelin turn over at different rates. J Biol Chem 267:10362–10369
Rainville PD, Stumpf CL, Shockcor JP, Plumb RS, Nicholson JK (2007) Novel application of reversed-phase UPLC-oaTOF-MS for lipid analysis in complex biological mixtures: a new tool for lipidomics. J Proteome Res. 6(2):552–558. https://doi.org/10.1021/pr060611b
Ogiso H, Suzuki T, Taguchi R (2008) Development of a reverse-phase liquid chromatography electrospray ionization mass spectrometry method for lipidomics, improving detection of phosphatidic acid and phosphatidylserine. Anal Biochem 375:124–131
Hu C et al (2008) RPLC-ion-trap-FTMS method for lipid profiling of plasma: method validation and application to p53 mutant mouse model. J Proteome Res 7:4982–4991
Le Grandois J et al (2009) Investigation of natural phosphatidylcholine sources: separation and identification by liquid chromatography–electrospray ionization–tandem mass spectrometry (LC–ESI–MS2) of molecular species. J Agric Food Chem 57:6014–6020
Vale P (2010) Profiles of fatty acids and 7-O-acyl okadaic acid esters in bivalves: can bacteria be involved in acyl esterification of okadaic acid? Comp Biochem Physiol C Toxicol Pharmacol 151:18–24
Sandra K, dos Santos Pereira A, Vanhoenacker G, David F, Sandra P (2010) Comprehensive blood plasma lipidomics by liquid chromatography/quadrupole time-of-flight mass spectrometry. J Chromatogr A 1217:4087–4099
Bird SS, Marur VR, Sniatynski MJ, Greenberg HK, Kristal BS (2011) Lipidomics profiling by high-resolution LC–MS and high-energy collisional dissociation fragmentation: focus on characterization of mitochondrial cardiolipins and monolysocardiolipins. Anal Chem 83:940–949
Bird SS, Marur VR, Sniatynski MJ, Greenberg HK, Kristal BS (2011) Serum lipidomics profiling using LC–MS and high-energy collisional dissociation fragmentation: focus on triglyceride detection and characterization. Anal Chem 83:6648–6657
Gruber F, Bicker W, Oskolkova OV, Tschachler E, Bochkov VN (2012) A simplified procedure for semi-targeted lipidomic analysis of oxidized phosphatidylcholines induced by UVA irradiation. J Lipid Res 53:1232–1242
Yamada T et al (2013) Development of a lipid profiling system using reverse-phase liquid chromatography coupled to high-resolution mass spectrometry with rapid polarity switching and an automated lipid identification software. J Chromatogr A 1292:211–218
Bird SS, Marur VR, Stavrovskaya IG, Kristal BS (2013) Qualitative characterization of the rat liver mitochondrial lipidome using LC–MS profiling and high energy collisional dissociation (HCD) all ion fragmentation. Metabolomics 9:67–83
Gallart-Ayala H et al (2013) Versatile lipid profiling by liquid chromatography-high resolution mass spectrometry using all ion fragmentation and polarity switching. Preliminary application for serum samples phenotyping related to canine mammary cancer. Anal Chim Acta 796:75–83
Wang J-R et al (2014) Improved sphingolipidomic approach based on ultra-high performance liquid chromatography and multiple mass spectrometries with application to cellular neurotoxicity. Anal Chem 86:5688–5696
Martano C, Mugoni V, Dal Bello F, Santoro MM, Medana C (2015) Rapid high performance liquid chromatography–high resolution mass spectrometry methodology for multiple prenol lipids analysis in zebrafish embryos. J Chromatogr A 1412:59–66
Zou J et al (2015) Evaluation of the change in sphingolipids in the human multiple myeloma cell line U266 and gastric cancer cell line MGC-803 treated with arsenic trioxide. J Chromatogr B Analyt Technol Biomed Life Sci 1004:98–107
Mi J-N, Wang J-R, Jiang Z-H (2016) Quantitative profiling of sphingolipids in wild Cordyceps and its mycelia by using UHPLC-MS. Sci Rep 6:20870
Triebl A, Trötzmüller M, Hartler J, Stojakovic T, Köfeler HC (2017) Lipidomics by ultrahigh performance liquid chromatography-high resolution mass spectrometry and its application to complex biological samples. J Chromatogr B Analyt Technol Biomed Life Sci 1053:72–80
Ballschmiter K, Wößner M (1998) Recent developments in adsorption liquid chromatography (NP-HPLC). Fresenius J Anal Chem 361:743–755
Christie WW, Urwin RA (1995) Separation of lipid classes from plant tissues by high performance liquid chromatography on chemically bonded stationary phases. J High Resolut Chromatogr 18:97–100
Olsson P, Holmbäck J, Herslöf B (2012) Separation of lipid classes by HPLC on a cyanopropyl column. Lipids 47:93–99
Nordbäck J, Lundberg E, Christie WW (1998) Separation of lipid classes from marine particulate material by HPLC on a polyvinyl alcohol-bonded stationary phase using dual-channel evaporative light-scattering detection. Mar Chem 60:165–175
Deschamps FS, Chaminade P, Ferrier D, Baillet A (2001) Assessment of the retention properties of poly(vinyl alcohol) stationary phase for lipid class profiling in liquid chromatography. J Chromatogr A 928:127–137
Sommer U, Herscovitz H, Welty FK, Costello CE (2006) LC–MS-based method for the qualitative and quantitative analysis of complex lipid mixtures. J Lipid Res 47:804–814
Sokol E et al (2013) Profiling of lipid species by normal-phase liquid chromatography, nanoelectrospray ionization, and ion trap-orbitrap mass spectrometry. Anal Biochem 443:88–96
Mclaren DG et al (2011) An ultraperformance liquid chromatography method for the normal-phase separation of lipids. Anal Biochem 414:266–272
Abreu S, Solgadi A, Chaminade P (2017) Optimization of normal phase chromatographic conditions for lipid analysis and comparison of associated detection techniques. J Chromatogr A 1514:54–71
Arnoldsson KC, Kaufmann P (1994) Lipid class analysis by normal phase high performance liquid chromatography, development and optimization using multivariate methods. Chromatographia 38:317–324
Prache N, Abreu S, Sassiat P, Thiébaut D, Chaminade P (2016) Alternative solvents for improving the greenness of normal phase liquid chromatography of lipid classes. J Chromatogr A 1464:55–63
Holčapek M et al (2015) Determination of nonpolar and polar lipid classes in human plasma, erythrocytes and plasma lipoprotein fractions using ultrahigh-performance liquid chromatography–mass spectrometry. J Chromatogr A 1377:85–91
Hutchins PM, Barkley RM, Murphy RC (2008) Separation of cellular nonpolar neutral lipids by normal-phase chromatography and analysis by electrospray ionization mass spectrometry. J Lipid Res 49:804–813
Donot F et al (2016) Rapid analysis and quantification of major neutral lipid species and free fatty acids by HPLC-ELSD from microalgae. Eur J Lipid Sci Technol 118:1550–1556
Gardner MS et al (2017) Simultaneous quantification of free cholesterol, cholesteryl esters, and triglycerides without ester hydrolysis by UHPLC separation and in-source collision induced dissociation coupled MS/MS. J Am Soc Mass Spectrom 28:2319–2329
Perona JS, Ruiz-Gutierrez V (2004) Quantification of major lipid classes in human triacylglycerol-rich lipoproteins by high-performance liquid chromatography with evaporative light-scattering detection. J Sep Sci 27:653–659
Jiang P, Lucy CA (2016) Coupling normal phase liquid chromatography with electrospray ionization mass spectrometry: strategies and applications. Anal Methods 8:6478–6488
Tavares KM et al (2016) Free tocopherols as chemical markers for Arabica coffee adulteration with maize and coffee by-products. Food Control 70:318–324
Nguyen HP, Schug KA (2008) The advantages of ESI-MS detection in conjunction with HILIC mode separations: fundamentals and applications. J Sep Sci 31:1465–1480
Méndez A, Bosch E, Rosés M, Neue UD (2003) Comparison of the acidity of residual silanol groups in several liquid chromatography columns. J Chromatogr A 986:33–44
McCalley DV, Neue UD (2008) Estimation of the extent of the water-rich layer associated with the silica surface in hydrophilic interaction chromatography. J Chromatogr A 1192:225–229
Grumbach ES, Diehl DM, Neue UD (2008) The application of novel 1.7 µm ethylene bridged hybrid particles for hydrophilic interaction chromatography. J Sep Sci 31:1511–1518
Wernisch S, Pennathur S (2016) Evaluation of coverage, retention patterns, and selectivity of seven liquid chromatographic methods for metabolomics. Anal Bioanal Chem 408:6079–6091
Tang D-Q, Zou L, Yin X-X, Ong C (2016) N. HILIC-MS for metabolomics: an attractive and complementary approach to RPLC-MS. Mass Spectrom Rev 35:574–600
Guo Y, Gaiki S (2011) Retention and selectivity of stationary phases for hydrophilic interaction chromatography. J Chromatogr A 1218:5920–5938
Cífková E, Hájek R, Lísa M, HolĿapek M (2016) Hydrophilic interaction liquid chromatography–mass spectrometry of (lyso)phosphatidic acids, (lyso)phosphatidylserines and other lipid classes. J Chromatogr A 1439:65–73
Vosse C, Wienken C, Cadenas C, Hayen H (2018) Separation and identification of phospholipids by hydrophilic interaction liquid chromatography coupled to tandem high resolution mass spectrometry with focus on isomeric phosphatidylglycerol and bis(monoacylglycero)phosphate. J Chromatogr A 1565:105–113
Buszewski B, Noga S (2012) Hydrophilic interaction liquid chromatography (HILIC)—a powerful separation technique. Anal Bioanal Chem 402:231–247
Zhu C et al (2012) An efficient hydrophilic interaction liquid chromatography separation of 7 phospholipid classes based on a diol column. J Chromatogr A 1220:26–34
Hines KM, Herron J, Xu L (2017) Assessment of altered lipid homeostasis by HILIC-ion mobility-mass spectrometry-based lipidomics. J Lipid Res 58:809–819
Lísa M, Cífková E, Holčapek M (2011) Lipidomic profiling of biological tissues using off-line two-dimensional high-performance liquid chromatography–mass spectrometry. J Chromatogr A 1218:5146–5156
Anesi A, Guella G (2015) A fast liquid chromatography–mass spectrometry methodology for membrane lipid profiling through hydrophilic interaction liquid chromatography. J Chromatogr A 1384:44–52
Pang L-Q, Liang Q-L, Wang Y-M, Ping L, Luo G-A (2008) Simultaneous determination and quantification of seven major phospholipid classes in human blood using normal-phase liquid chromatography coupled with electrospray mass spectrometry and the application in diabetes nephropathy. J Chromatogr B 869:118–125
Zheng L et al (2010) Profiling of lipids in leishmania donovani using hydrophilic interaction chromatography in combination with fourier transform mass spectrometry. Rapid Commun Mass Spectrom 24:2074–2082
Reis A et al (2013) A comparison of five lipid extraction solvent systems for lipidomic studies of human LDL. J Lipid Res 54:1812–1824
Scherer M, Schmitz G, Liebisch G, Schmitz G (2009) High-throughput analysis of sphingosine 1-phosphate, sphinganine 1-phosphate, and lysophosphatidic acid in plasma samples by liquid chromatography-tandem mass spectrometry. Clin Chem 55:1218–1222
Cífková E et al (2015) Lipidomic differentiation between human kidney tumors and surrounding normal tissues using HILIC–HPLC/ESI–MS and multivariate data analysis. J Chromatogr B Analyt Technol Biomed Life Sci 1000:14–21
Cascone A, Eerola S, Ritieni A, Rizzo A (2006) Development of analytical procedures to study changes in the composition of meat phospholipids caused by induced oxidation. J Chromatogr A 1120:211–220
Wang S et al (2013) A novel stop-flow two-dimensional liquid chromatography–mass spectrometry method for lipid analysis. J Chromatogr A 1321:65–72
Fei F, Bowdish DME, McCarry BE (2014) Comprehensive and simultaneous coverage of lipid and polar metabolites for endogenous cellular metabolomics using HILIC–TOF–MS. Anal Bioanal Chem 406:3723–3733
Scherer M, Leuthäuser-Jaschinski K, Ecker J, Schmitz G, Liebisch G (2010) A rapid and quantitative LC–MS/MS method to profile sphingolipids. J Lipid Res 51:2001–2011
Fauland A et al (2011) A comprehensive method for lipid profiling by liquid chromatography-ion cyclotron resonance mass spectrometry. J Lipid Res 52:2314–2322
Řezanka T, Siristova L, Matoulková D, Sigler K (2011) Hydrophilic interaction liquid chromatography: ESI–MS/MS of plasmalogen phospholipids from Pectinatus bacterium. Lipids 46:765–780
Lísa M (2017) Silver-ion liquid chromatography–mass spectrometry. Handb Adv Chromatogr Spectrom Tech. https://doi.org/10.1016/B978-0-12-811732-3.00004-2
Nikolova-Damyanova B, Christie WW (1992) Advances in lipid methodology, vol 1 (P. J. Barnes & Associates). The Oily Press, Bridgwater
Nikolova-Damyanova B (2009) Retention of lipids in silver ion high-performance liquid chromatography: facts and assumptions. J Chromatogr A 1216:1815–1824
Lísa M, Velínská H, Holčapek M (2009) Regioisomeric characterization of triacylglycerols using silver-Ion HPLC/MS and randomization synthesis of standards. Anal Chem 81:3903–3910
François I, dos Santos Pereira A, Sandra P (2010) Considerations on comprehensive and off-line supercritical fluid chromatography × reversed-phase liquid chromatography for the analysis of triacylglycerols in fish oil. J Sep Sci 33:1504–1512
Beccaria M et al (2015) Determination of the triacylglycerol fraction in fish oil by comprehensive liquid chromatography techniques with the support of gas chromatography and mass spectrometry data. Anal Bioanal Chem 407:5211–5225
Danne-Rasche N, Coman C, Ahrends R, Nano (2018) LC/NSI MS refines lipidomics by enhancing lipid coverage, measurement sensitivity, and linear dynamic range. Anal Chem. https://doi.org/10.1021/acs.analchem.8b01275
Pettitt TR, Dove SK, Lubben A, Calaminus SDJ, Wakelam, MJO (2006) Analysis of intact phosphoinositides in biological samples. J Lipid Res 47:1588–1596
Knittelfelder OL, Weberhofer BP, Eichmann TO, Kohlwein SD, Rechberger GN (2014) A versatile ultra-high performance LC–MS method for lipid profiling. J Chromatogr B 951–952:119–128
Cífková E, Holčapek M, Lísa M (2013) Nontargeted lipidomic characterization of porcine organs using hydrophilic interaction liquid chromatography and off-line two-dimensional liquid chromatography–electrospray ionization mass spectrometry. Lipids 48:915–928
Hu J et al (2013) Characterization and quantification of triacylglycerols in peanut oil by off-line comprehensive two-dimensional liquid chromatography coupled with atmospheric pressure chemical ionization mass spectrometry. J Sep Sci 36:288–300
Walczak J, Bocian S, Buszewski B (2015) Two-dimensional high performance liquid chromatography–mass spectrometry for phosphatidylcholine analysis in egg yolk. Food Anal Methods 8:661–667
Narváez-Rivas M, Vu N, Chen G-Y, Zhang Q (2017) Off-line mixed-mode liquid chromatography coupled with reversed phase high performance liquid chromatography-high resolution mass spectrometry to improve coverage in lipidomics analysis. Anal Chim Acta 954:140–150
Mondello L et al (2005) Silver-ion reversed-phase comprehensive two-dimensional liquid chromatography combined with mass spectrometric detection in lipidic food analysis. J Chromatogr A 1086:91–98
Dugo P, Kumm T, Crupi ML, Cotroneo A, Mondello L (2006) Comprehensive two-dimensional liquid chromatography combined with mass spectrometric detection in the analyses of triacylglycerols in natural lipidic matrixes. J Chromatogr A 1112:269–275
Holčapek M, Ovčačíková M, Lísa M, Cífková E, Hájek T (2015) Continuous comprehensive two-dimensional liquid chromatography–electrospray ionization mass spectrometry of complex lipidomic samples. Anal Bioanal Chem 407:5033–5043
Ling YS et al (2014) Two-dimensional LC–MS/MS to enhance ceramide and phosphatidylcholine species profiling in mouse liver. Biomed Chromatogr 28:1284–1293
Byrdwell WC (2010) Dual parallel mass spectrometry for lipid and vitamin D analysis. J Chromatogr A 1217:3992–4003
Nie H et al (2010) Lipid profiling of rat peritoneal surface layers by online normal- and reversed-phase 2D LC QToF-MS. J Lipid Res 51:2833–2844
Li M et al (2014) A not-stop-flow online normal-/reversed-phase two-dimensional liquid chromatography–quadrupole time-of-flight mass spectrometry method for comprehensive lipid profiling of human plasma from atherosclerosis patients. J Chromatogr A 1372:110–119
Yang L et al (2017) Lipidomic analysis of plasma in patients with lacunar infarction using normal-phase/reversed-phase two-dimensional liquid chromatography–quadrupole time-of-flight mass spectrometry. Anal Bioanal Chem 409:3211–3222
Yang L et al (2015) Comprehensive lipid profiling of plasma in patients with benign breast tumor and breast cancer reveals novel biomarkers. Anal Bioanal Chem 407:5065–5077
Bang DY, Moon MH (2013) Online two-dimensional capillary strong anion exchange/reversed phase liquid chromatography–tandem mass spectrometry for comprehensive lipid analysis. J Chromatogr A 1310:82–90
Wang S, Zhou L, Wang Z, Shi X, Xu G (2017) Simultaneous metabolomics and lipidomics analysis based on novel heart-cutting two-dimensional liquid chromatography–mass spectrometry. Anal Chim Acta 966:34–40
Casetta B, Vecchione G, Tomaiuolo M, Margaglione M, Grandone E (2009) Setting up a 2D-LC/MS/MS method for the rapid quantitation of the prostanoid metabolites 6-oxo-PGF1α and TXB2 as markers for hemostasis assessment. J Mass Spectrom 44:346–352
Jung U, Choi M-S (2014) Obesity and its metabolic complications: the role of adipokines and the relationship between obesity, inflammation, insulin resistance, dyslipidemia and nonalcoholic fatty liver disease. Int J Mol Sci 15:6184–6223
Pietiläinen KH et al (2011) Association of lipidome remodeling in the adipocyte membrane with acquired obesity in humans. PLoS Biol 9:e1000623
Luo L, Liu M (2016) Adipose tissue in control of metabolism. J Endocrinol 231:R77–R99
Kotronen A et al (2010) Comparison of lipid and fatty acid composition of the liver, subcutaneous and intra-abdominal adipose tissue, and serum. Obesity 18:937–944
Baker RC, Nikitina Y, Subauste AR (2014) Analysis of adipose tissue lipid using mass spectrometry. Methods Enzymol 538:89–105
Sen I et al (2015) Lipid profiles of adipose and muscle tissues in mouse models of juvenile onset of obesity without high fat diet induction: a fourier transform infrared (FT-IR) spectroscopic study. Appl Spectrosc 69:679–688
Roberts LD, West JA, Vidal-Puig A, Griffin JL (2014) Methods for performing lipidomics in white adipose tissue. Methods Enzymol 538:211–231
Al-Sulaiti H et al (2018) Triglyceride profiling in adipose tissues from obese insulin sensitive, insulin resistant and type 2 diabetes mellitus individuals. J Transl Med 16:175
Guan M et al (2017) Comprehensive qualification and quantification of triacylglycerols with specific fatty acid chain composition in horse adipose tissue, human plasma and liver tissue. Talanta 172:206–214
Medina-Gomez G et al (2007) PPAR gamma 2 prevents lipotoxicity by controlling adipose tissue expandability and peripheral lipid metabolism. PLoS Genet 3:e64
Kolak M et al (2007) Adipose tissue inflammation and increased ceramide content characterize subjects with high liver fat content independent of obesity. Diabetes 56:1960–1968
Mattila I, Seppänen-Laakso T, Suortti T, Orešič M (2008) Application of lipidomics and metabolomics to the study of adipose tissue. Methods Mol Biol (Clifton NJ) 456:123–130
Prieur X et al (2011) Differential lipid partitioning between adipocytes and tissue macrophages modulates macrophage lipotoxicity and M2/M1 polarization in obese mice. Diabetes 60:797–809
Hoene M et al (2014) The lipid profile of brown adipose tissue is sex-specific in mice. Biochim Biophys Acta Mol Cell Biol Lipids 1841:1563–1570
Haensig M et al (2014) Improved mitral valve performance after transapical aortic valve implantation. Ann Thorac Surg 97:1247–1253 (discussion 1253–4)
Yore MM et al (2014) Discovery of a class of endogenous mammalian lipids with anti-diabetic and anti-inflammatory effects. Cell 159:318–332
Aicheler F et al (2015) Retention time prediction improves identification in nontargeted lipidomics approaches. Anal Chem 87:7698–7704
Mcilroy GD et al (2016) Fenretinide mediated retinoic acid receptor signalling and inhibition of ceramide biosynthesis regulates adipogenesis, lipid accumulation, mitochondrial function and nutrient stress signalling in adipocytes and adipose tissue. Biochem Pharmacol 100:86–97
Carroll JB et al (2015) HdhQ111 mice exhibit tissue specific metabolite profiles that include striatal lipid accumulation. PLoS One 10:e0134465
Funding
Financial support from the German Federal Ministry of Education and Research (BMBF) within the framework of the e:Med research and funding concept for SysMedOS project (to MF) and EU H2020 funded project MASSTRPLAN (Grant number 675132; to MF) are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Published in Chromatographia’s 50th Anniversary Commemorative Issue.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lange, M., Ni, Z., Criscuolo, A. et al. Liquid Chromatography Techniques in Lipidomics Research. Chromatographia 82, 77–100 (2019). https://doi.org/10.1007/s10337-018-3656-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-018-3656-4