Abstract
The taxonomic affinity of the near-flightless Zapata Rail Cyanolimnas cerverai, a critically endangered and highly localized species endemic to Cuba, has long been debated. Morphological analyses have suggested that this species, which constitutes a monotypic genus, could be related either to the extinct Tahitian Cave Rails (Nesotrochis sp.) or to the South American rail tribe Pardirallini, i.e., the genera Neocrex, Mustelirallus, and Pardirallus. While pronounced phenotypic convergence—and divergence—among rails have repeatedly proven morphology-based phylogenies unreliable, thus far no attempt to sequence DNA from the enigmatic Cyanolimnas has succeeded. In this study, we extracted historic DNA from a museum specimen collected in 1927 and sequenced multiple short fragments that allowed us to assemble a partial sequence of the mitochondrial cytochrome oxidase I gene. Phylogenetic analyses confirm that Cyanolimnas belongs in tribe Pardirallini as sister to genus Neocrex, from which it diverged about 6 million years ago. Their divergence from Mustelirallus was estimated at about 9 million years ago. Based on morphology and our mitochondrial phylogeny, we conclude that it is unjustified to retain the monotypic genus Cyanolimnas and tentatively recommend that C. cerverai and the two Neocrex species are ascribed to genus Mustelirallus.
Zusammenfassung
Die ersten genetischen Daten für das vom Aussterben bedrohte Kubasumpfhuhn Cyanolimnas cerverai und ihre taxonomischen Folgen
Die taxonomische Position des fast flugunfähigen Kubasumpfhuhns Cyanolimnas cerverai, einer vom Aussterben bedrohten und in ihrer Ausbreitung stark eingeschränkten Art, die auf Kuba endemisch ist, wird seit langem diskutiert. Morphologische Analysen haben darauf hingedeutet, dass diese Art, die eine monotypische Gattung darstellt, entweder mit einer Gattung ausgestorbener tahitianischer Rallen (Nesotrochis sp.) oder mit dem südamerikanischen Rallentribus Pardirallini, d.h. den Gattungen Neocrex, Mustelirallus und Pardirallus, verwandt sein könnte. Während ausgeprägte phänotypische Konvergenz und Divergenz bei Rallen wiederholt gezeigt haben, dass auf der Morphologie basierende Stammbäume unzuverlässig sind, war bislang kein Versuch, die DNA der enigmatischen Cyanolimnas zu sequenzieren, von Erfolg gekrönt. In dieser Studie haben wir historische DNA aus einem Museumspräparat aus dem Jahr 1927 extrahiert und multiple kurze Fragmente sequenziert, die uns erlaubten, eine partielle Sequenz des mitochondrialen Cytochromoxidase I-Gens zu ermitteln. Phylogenetische Analysen bestätigen, dass Cyanolimnas in den Tribus Pardirallini als Schwestergruppe der Gattung Neocrex gehört, von der die Abspaltung vor etwa sechs Millionen Jahren erfolgte. Die Abspaltung von Mustelirallus erfolgte vor geschätzt etwa neun Millionen Jahren. Auf der Grundlage der Morphologie und unseres mitochondrialen Stammbaums kommen wir zu dem Schluss, dass es nicht gerechtfertigt ist, die monotypische Gattung Cyanolimnas beizubehalten, und empfehlen, dass C. cerverai und die zwei Neocrex-Arten der Gattung Mustelirallus zugeordnet werden.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The critically endangered Zapata Rail Cyanolimnas cerverai is endemic to Cuba and unquestionably is one of the most poorly known birds in the West Indian region (Kirwan et al. 2019; Kirkconnell et al. 2020; Taylor et al. 2020). Its declining population is apparently threatened by dry-season burning of its marsh habitat, predation by introduced small Indian Mongooses (Herpestes auropunctatus), Black Rats (Rattus rattus) and African Catfish Clarias gariepinus (Collar et al. 1992; Kirkconnell 2012; Taylor et al. 2020), and habitat change engendered by the spread of the invasive Broad-leaved Paperback (Melaleuca quinquenervia). Nowadays, this rail is known from just six closely spaced localities in the northern Ciénaga de Zapata (Matanzas province) in south mainland Cuba, based on 14 specimens collected between 1927 and 1934 (all but one in four US museums; Table S1), and a relatively small number of sight records since 1979, most recently in November 2014 (Kirkconnell et al. 2020). However, fossil material of Cyanolimnas cerverai dating from the Holocene has been identified from Cueva de Pío Domingo, Sumidero, and Cueva El Abrón, Sierra de La Güira (Pinar del Río province), Cueva de Paredones and Cueva de Sandoval (Artemisa province), Calabazar (La Habana province), Cueva de Insunsa, Cuevas de Las Charcas and Cueva del Indio (all in Mayabeque province), near Jagüey Grande (Matanzas province), Cueva de Humboldt and Cueva del Salón (Sancti Spíritus province), and the Sierra de Caballos in the northern Isle of Pines (Olson 1974; Arredondo 1984; Jiménez and Valdés 1995; Rojas-Consuegra et al. 2012; Jiménez and Orihuela 2021; Suárez 2022), i.e., over a much larger area than the known modern range.
Genus Cyanolimnas Barbour and Peters, 1927, has always been considered monospecific, and was described as “A medium-sized ralline with short rounded wing; very short tail, the barbs of the rectrices very sparse; tarsus stout and short, not exceeding middle toe with claw. Bill moderate, somewhat longer than head, swollen basally. … The combination of short wing and stout tarsus suggests relationships with Nesotrochis Wetmore [a genus comprising three large, flightless, extinct species from the Greater Antilles] … but the latter has a tarsus more than twice as long” (Barbour and Peters 1927). Ridgway and Friedmann (1941) proffered a detailed morphological description of this genus, and thought it was apparently flightless. However, while Cyanolimnas has reduced powers of flight, it is volant (AK pers obs), and in any case, flightlessness is now known to have arisen many times in Rallidae and cannot be used as a predictor of relationships (e.g., Olson 1973; Slikas et al. 2002; Kirchman 2012; Gaspar et al. 2020; Garcia-R and Matzke 2021, although Cyanolimnas was incorrectly treated as flightless in the latter). As noted by Olson (1973) and Steadman et al. (2013), in its robust, deep-based bill Cyanolimnas is similar to the two species frequently assigned to Neocrex Sclater & Salvin, 1868, Colombian Crake N. colombiana and Paint-billed Crake N. erythrops (both of which are now often placed in an expanded Mustelirallus Bonaparte, 1856; e.g., Kirchman et al. 2021). Plumage and osteological characters are similar to either Neocrex, or less so to the latter’s sister taxon Pardirallus Bonaparte, 1856, to which genus Cyanolimnas was considered most closely related in the morphological phylogenies of Livezey (1998) and Garcia-R and Matzke (2021). Currently, all four of the global bird checklists place Cyanolimnas close to both Neocrex and Pardirallus (Dickinson and Remsen 2013; del Hoyo et al 2014; Clements et al. 2021; Gill et al. 2022).
Its ecology and natural history are virtually unknown; for example, a nest with eggs ascribed to Cyanolimnas, found in early September 1982 (Bond 1984), seems unlikely to have been identified correctly (Kirkconnell et al. 2020), and even this rail’s voice is unknown. A published sound recording, originally believed to pertain to Cyanolimnas (Reynard and Garrido 1988; Hardy et al. 1996), is now identified as belonging to Spotted Rail Pardirallus maculatus, which is a rather abundant species in the Ciénaga de Zapata (Kirkconnell et al. 2020). During November–December 1998, a survey using the published sound recording as a tool estimated a population of 70–90 individuals of Cyanolimnas (Kirkconnell et al. 1999). Subsequently, it was realized that the recording actually involved P. maculatus (Kirkconnell et al. 2005) and that the description of Cyanolimnas vocalizations in Kirkconnell et al. (1999) was erroneous. Inferences concerning the relationships of Cyanolimnas based on ecology or vocalizations are consequently impossible.
To date, just one molecular phylogenetic study, using ultra-conserved elements (UCEs), has attempted to ascertain the relationships of Cyanolimnas (using a toe pad from AMNH 300416, a female collected by P. Quintaña in April 1934), but was unsuccessful in yielding any UCEs, and consequently failed to place the species (Kirchman et al. 2021). As a result, Kirchman et al. (2021) proposed to treat Cyanolimnas as genus incertae sedis; the aim of the present work is to address this shortfall in knowledge.
Methods
Museum specimens and sampling
Worldwide, there are 14 Zapata Rail specimens housed in museum collections (Table S1). With written permission from the museum, we obtained a ~ 5 mm3 toepad sample of a Zapata Rail from the Museo de Historia Natural ‘Felipe Poey’, La Habana, Cuba (catalog no. MFP 14.000218; Fig. S1) using a clean scalpel. The specimen in question was a male collected in the Ciénaga de Zapata, Cuba, on 17 June 1927 (Table S1).
Primer design and evaluation
We obtained cytochrome oxidase I (COI) sequences of rallid species NCBI/GenBank (https://www.ncbi.nlm.nih.gov/nucleotide/), aligned these sequences using CLC Sequence viewer v6.0 (QIAGEN; UK) (https://digitalinsights.qiagen.com/products/clc-sequence-viewer-direct-download/) and identified regions of high similarity and roughly 50% GC content. We manually designed 21–23 bp long primers, including some degenerated sites to account for alignment sequence variation among, to produce < 200 bp fragments from the degraded DNA (Table 1; Fig. S2). We tested the primer pairs on Common Moorhen Gallinula chloropus and Allen’s Gallinule Porphyrio alleni, because DNA availability was not a restriction for these two rallid species, in order to assess the likelihood of successful amplification of a limited amount of historic DNA from the Zapata Rail. Additional primers (not listed) were tested but did not yield successful amplification/sequencing.
Sampling and laboratory procedures
We extracted DNA from the toepad sample MFP 14.000218 using QIAGEN QIAamp® DNA microkit, following the manufacturer’s instructions, modified for digestion overnight at 56 °C and repeated vortexing. The DNA was eluted in 80 µl Buffer AE, and quantified to 22 ng/μl on a Nanodrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA).
We set up 50 µl PCR reactions using 2.5 µl template DNA and 47.5 µl PCR mastermix made from 5 µl of 10 × OptiBuffer (Bioline Reagents Ltd, UK), 2.5 µl of each primer at 10 μM, 1 µl 50 mM magnesium chloride, 0.6 µl 10 mM DNT mix (Sigma, Poole, UK), 0.5 µl BIO-X-ACT short DNA polymerase (Bioline Reagents Ltd, London, UK), and 35.4 µl ddH2O. We ran samples and negative controls in PCR programs following Hebert et al. (2004), with a low annealing temperature in first 5 cycles to accommodate potential primer–sample mismatches: denaturing 60 s at 94 °C, annealing 60 s at 45 °C, and extension 50 s at 72 °C. For the following 30 cycles, the annealing temperature was increased to 51 °C. We adopted this PCR strategy for all primer combinations.
Four µl 100-bp DNA ladder (Promega, Southampton, UK) and 40 µl of the PCR products were loaded with 5 µl Promega 6 × DNA loading dye on 1.5% agarose gels (peqGOLD universal agarose; Peqlab, Fareham, UK), stained with 1–2 µl ethidium bromide, in 1 × TAE buffer. We ran the electrophoresis at 86 V, 100 mA for 20–40 min, and inspected the gel under UV light. While PCR reactions were specific, primer dimers did form, and due to the relatively small difference in length between these and the PCR product we chose not to perform differential precipitation to purify the PCR product, Instead, we cut the bands of expected size with a scalpel and isolated the PCR product from the gel fragments with a Gel Extraction Kit (Qiagen, Manchester, UK), following the manufacturer’s instructions, and eluted in 30 µl elution buffer EB. We quantified the purified PCR product with a Nanodrop spectrophotometer and diluted the PCR products to 10 µg/µl, after which they were bidirectionally Sanger sequenced by Source Bioscience (Nottingham, UK).
Phylogenetic analyses
We initially scrutinized the sequences using FinchTV (Geospiza Inc.) and then assembled them to the rallid sequence set used for primer design in CLC Sequence Viewer 8 (https://www.qiagenbioinformatics.com/products/clc-sequence-viewer/). This resulted in a 558-bp consensus sequence, with one internal 11-bp and one 18-bp gap consisting of primer sequence, resulting in 529 informative nucleotides (Fig. S2). We then obtained a larger set of 450 COI sequences of extant taxa within the order Gruiformes from GenBank, and additionally included sequence data from ancient DNA of the extinct Nesotrochis steganinos (Oswald et al. 2021), because of its proposed close relationship to the extant taxa included in the analyses in combination with its recent extinction (Oswald et al. 2021 dated bones to 6,430 ± 30 years before present). We aligned sequences with MAFFT (Katoh and Standley 2013) in Geneious v. 10.2.6 (Kearse et al. 2012) and selected ≤ 3 per taxon based on length and quality (for taxa and accession numbers, see Table S2). We further trimmed this dataset to a 962-bp region that maximized overall sequence coverage and fully overlapped the Zapata Rail COI sequence (see “Data availability”). We employed the greedy algorithm (Lanfear et al. 2012) in PartitionFinder2 (Lanfear et al. 2016) to evaluate the full set of DNA substitution models and explore optimal partitioning with regard to codon position, using PhyML (Guindon et al. 2010). The best partitioning scheme by AICc was identified as individual substitution models per codon position: the general time-reversible (GTR; Tavaré 1986) model with the among-site rate variation following a gamma (Γ) distribution and with a proportion of invariant sites (I) for position 1, the transversion model (TVM) + Γ + I for position 2, and the transition model (TiM) + Γ + I for position 3.
We implemented the partitioning scheme in Beast v. 2.6.6 (Bouckaert et al. 2014) using the plugin SSM v. 1.1.0 and set the proportion of invariant sites to be estimated, and the Γ distribution to be estimated across four rate categories. Employing a birth–death speciation model, we set the prior for death rate to follow an exponential distribution with mean 1. We enforced monophyly of the clades Rallidae, Ralloidea, and Gruoidea, and employed dating calibration priors on Gruiformes and stem Rallidae following Chaves et al. (2020). We then ran two iterations of 10 million generations, sampling every 1000 generations, and optimized tuning parameters and operator weights to produce the final xml specification used for our analyses (see “Data availability”). We ran five replicate analyses at different seeds in Beast, sampling every 1000 generation for 40–145 million generations, until inspection in Tracer v. 1.7.1 (Rambaut et al. 2018) revealed stationarity and sufficient effective samples sizes (all parameters > 200). Some substitution model-related parameters for codon position 1 and to some degree 2 (specific substitution rates, gamma shape, and proportion invariant sites) seemed to have dual optima, with shifting within and between replicates. Thus, replicates converged in either of two groups; however, this had no impact on the topology, support, or dating of the focal clade. We calculated maximum clade credibility trees with mean node heights after discarding 5–50% as burn-in.
In addition to the Bayesian inference (BI) with Beast, we also used IQtree v. 2.1.2 (Lanfear et al. 2020) for phylogenetic analyses based on maximum likelihood (ML). We applied the partitioning and substitution model scheme determined with PartitionFinder2 and ran two separate analyses comprising 1000 ultrafast bootstrap replicates.
Results
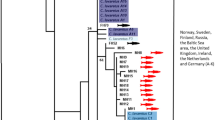
All phylogenetic analyses placed Cyanolimnas as sister to N. erythrops, the pair of which was sister to the Ash-throated Crake Mustelirallus (formerly Porzana) albicollis (Fig. 1). These three taxa formed a sister clade to the genus Pardirallus (Fig. 1). There was near-full support for a Pardirallus–Mustelirallus–Neocrex–Cyanolimnas clade (posterior probability (PP) 1.0 in the BI analyses and 99% ultrafast bootstrap support (UFB) in the ML analyses). The exact placement of Cyanolimnas as sister to Neocrex received full support (UFB 100%) in the ML analyses and PP 0.86 with BI (Fig. 1). There were some differences in inferred topology between BI replicates (see “Data availability”); however, none of these occurred within the focal Pardirallus–Mustelirallus–Neocrex–Cyanolimnas clade.
Phylogenetic tree of Gruiformes, based on mitochondrial cytochrome oxidase I (COI) sequences, analyzed with Bayesian inference (BI) in Beast. Posterior probability is indicated by node color (0.0 = white; 1.0 = black) and specified with labels at nodes for the focal clade in tribe Pardirallini (first number), together with ultrafast bootstrap support values (0–100) from maximum likelihood (ML) analyses with IQtree (second number). Note that many deeper relationships are not very well resolved or well supported by analyses based on a partial COI sequence alone; for example, Gruidae is rendered paraphyletic. The asterisks at the root of Gruiformes and the stem of Rallidae indicate fossil calibration points. Non-focal clades represented by multiple sequences have been collapsed and labeled according to Kirchman et al. (2021) with tribe (for Rallidae) or family (outside Rallidae) followed by species or genus (if multiple species). For non-collapsed replicate BI trees and ML tree, see “Data availability”
The inferred mean age of split between Cyanolimnas and Neocrex was 6.3 MA (95% highest posterior density 2.9–10.0 MA), between those and Mustelirallus 8.8 (4.9–13.1) MA, and between that clade and Pardirallus 11.5 (7.0–16.2) MA (Fig. 1).
In line with Oswald et al. (2021), Nesotrochis was recovered as a sister lineage to Sarothruridae (Fig. 1).
Discussion
We provide the first genetic data for Cyanolimnas, as the species has not been included or successfully sequenced by any previous molecular phylogenetic study of the Rallidae (e.g., Garcia-R et al. 2014; Garcia-R and Matzke 2021; Kirchman et al. 2021), which led Kirchman et al. (2021) to consider its placement ‘incertae sedis’. It should be stressed that the reliability of phylogenies based on single genetic markers is limited, and this is particularly true for mitochondrial markers. The fast-evolving mitochondrion occurs with multiple copies per cell and has historically been widely used for molecular phylogenies, and further remains important for genetic barcoding. However, for several reasons the mitochondrial phylogenetic signal may be discordant from that of the nuclear genome (e.g., through introgression) and reflect an evolutionary trajectory different from ‘true’ speciation events (Toews and Brelsford 2012). In our analyses, for example, the COI tree renders Gruidae paraphyletic with respect to Aramidae (Fig. 1). Nevertheless, mitochondrial data are relevant for a first test of presumed phylogenetic context, and yield useful relative divergence times.
Similar to the whole-mitochondrion analyses by Oswald et al. (2021), our analyses of COI placed Nesotrochis outside Rallidae, separated from Cyanolimnas by some 45 million years (Fig. 1). Our data instead provide strong support for the previous hypotheses rooted in morphological traits that have placed Cyanolimnas in what has been referred to as the ‘Aramides clade’ (Livezey 1998; Garcia-R et al. 2014; Garcia-R and Matzke 2021), better termed tribe Pardirallini sensu Kirchman et al. (2021). Within this grouping, the species appears to be most closely related to N. erythrops and M. albicollis, but less so to the three species of Pardirallus, in partial congruence with prior predictions (Olson 1973; Livezey 1998; Garcia-R & Matzke 2021). Although it is now rather well established that plumage is not necessarily a reliable character for inferring phylogenetic relationships within the Rallidae (e.g., Garcia-R et al. 2014; Stervander et al. 2019; Chaves et al. 2020), in the present case morphology appears to be rather informative.
The key question that arises is whether the separate monospecific genus Cyanolimnas is warranted? The age of the split between Cyanolimnas and Neocrex (6.3 MA) is comparatively young relative to many other Rallidae, a family that contains several old generic clades (e.g., Aramides 7.7 MA, Porzana 10.4 MA, Rallus 11.1 MA, Rallina 12.6 MA, Zapornia 13.8 MA, and Porphyrio 19.0 MA based our COI analyses; Fig. 1). However, some clades contain even younger taxa, for example the Eulabeornis–Cabalus–Gallirallus–Hypotaenidia group of the Rallini (which arose 4.3 MA), all or most of which genera are accepted by some authorities (e.g., Dickinson and Remsen 2013; del Hoyo et al 2014; Kirchman et al. 2021). Nevertheless, in the latter case these four genera are sometimes treated alternatively as a single genus, for which the name Gallirallus Lafresnaye, 1841, has priority (Kirchman 2012).
We consider that the available molecular and morphological evidence in combination strongly supports that C. cerverai be included in either genus Neocrex Sclater & Salvin, 1869, or Mustelirallus, Bonaparte, 1856, given that its most divergent trait from either of these two, (near-)flightlessness, is well accepted to be not taxonomically informative (Olson 1973; Slikas et al. 2002; Kirchman 2012; Gaspar et al. 2020; Garcia-R and Matzke 2021). Although N. colombiana, an exceptionally poorly known rallid of northwestern South America and easternmost Panama, has yet to be sampled genetically, to date, there is no indication that it is not very closely related to N. erythrops (Wetmore 1967; Olson 1973; Taylor 1996). Following the lead of Kirchman et al. (2021), we recommend that, for the present, the most parsimonious approach is to treat all four species—Ash-throated Crake, Paint-billed Crake, Colombian Crake and, now, Zapata Rail—as members of a single genus, for which the name Mustelirallus (masculine) has priority. The new combinations are Mustelirallus albicollis, M. erythrops, M. colombianus, and M. cerverai. Kirchman et al. (2021) incorrectly listed Colombian Crake as M. columbianus, overlooking that Bangs’ (1898) original spelling was Neocrex colombianus as noted by Dickinson and Remsen (2013).
Data availability
The Zapata Rail COI sequence has been deposited at Genbank with accession number OL449772. Supporting material has been deposited at Zenodo and is available at https://doi.org/10.5281/zenodo.6500391. This includes Table S2 (taxon name, classification, and accession numbers for sequences included); sequence alignment nexus file; Beast xml input file; Beast output including maximum clade credibility tree files; log files and raw tree files; and IQtree consensus tree files.
References
Arredondo O (1984) Sinopsis de las aves halladas en depósitos fosilíferos pleisto-holocénicos de Cuba. Rep Invest Inst Zool Acad Cienc Cuba 17:1–35
Bangs O (1898) On some birds from the Sierra Nevada de Santa Marta, Colombia. Proc Biol Soc Wash 12:171–182
Barbour T, Peters JL (1927) Two more remarkable new birds from Cuba. Proc N Engl Zool Club 9:95–97
Bond J (1984) Twenty-fifth supplement to the check-list of birds of the West Indies (1956). Academy of Natural Sciences, Philadelphia
Bouckaert R, Heled J, Kuhnert D, Vaughan T, Wu CH, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. Plos Comput Biol 10:e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Chaves JA, Martinez-Torres PJ, Depino EA, Espinoza-Ulloa S, García-Loor J, Beichman AC, Stervander M (2020) Evolutionary history of the Galápagos rail revealed by ancient mitogenomes and modern samples. Diversity 12:425. https://doi.org/10.3390/d12110425
Clements JF, Schulenberg TS, Iliff MJ, Billerman SM, Fredericks TA, Gerbracht JA, Lepage D, Sullivan BL, Wood CL (2021) The eBird/Clements checklist of birds of the world: v2021. https://www.birds.cornell.edu/clementschecklist/download/
Collar N, Gonzaga L, Krabbe N, Madroño Nieto A, Naranjo L, Parker T III, Wege D (1992) Threatened Birds of the Americas: the ICBP/IUCN Red Data Book. International Council for Bird Preservation, Cambridge
del Hoyo J, Collar NJ, Christie DA, Elliott A, Fishpool LDC (2014) HBW and birdlife international illustrated checklist of the birds of the world, vol 1: non-passerines. Lynx Edicions and BirdLife International, Barcelona
Dickinson E, Remsen Jr JV (eds) (2013) The Howard & Moore Complete Checklist of the Birds of the World, vol 1, 4th edn. Aves Press, Eastbourne
Garcia-R JC, Matzke NJ (2021) Trait-dependent dispersal in rails (Aves: Rallidae): historical biogeography of a cosmopolitan bird clade. Mol Phylogenet Evol 159:107106. https://doi.org/10.1016/j.ympev.2021.107106
Garcia-R JC, Gibb GC, Trewick SA (2014) Deep global evolutionary radiation in birds: diversification and trait evolution in the cosmopolitan bird family Rallidae. Mol Phylogenet Evol 81:96–108. https://doi.org/10.1016/j.ympev.2014.09.008
Gaspar J, Gibb GC, Trewick SA (2020) Convergent morphological responses to loss of flight in rails (Aves: Rallidae). Ecol Evol 10:6186–6207. https://doi.org/10.1002/ece3.6298
Gill F, Donsker D, Rasmussen P (eds) (2022) IOC World Bird List (v12.1). https://doi.org/10.14344/IOC.ML.12.1
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Hardy J, Reynard G, Taylor T (1996) Voices of the New World rails. ARA Records, Gainesville
Hebert PDN, Stoeckle MY, Zemlak TS, Francis CM (2004) Identification of birds through DNA barcodes. PLoS Biol 2:1657–1663
Jiménez O, Orihuela J (2021) Nuevos hallazgos de aves en contextos paleontológicos y arqueológicos de Cuba. Nov Carib 163–176
Jiménez O, Valdés P (1995) Los vertebrados fósiles de la Cueva del Indio, San José de las Lajas, Habana, Cuba. Resúmenes de Congr Intern LV Aniversario de la Soc Espeleológica de Cuba, pp 62–63
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Kirchman JJ (2012) Speciation of flightless rails on islands: a DNA-based phylogeny of the typical rails of the Pacific. Auk 129:56–69. https://doi.org/10.1525/auk.2011.11096
Kirchman JJ, Rotzel McInerney N, Giarla TC, Olson SL, Slikas E, Fleischer RC (2021) Phylogeny based on ultra-conserved elements clarifies the evolution of rails and allies (Ralloidea) and is the basis for a revised classification. Ornithology 138:ukab042. https://doi.org/10.1093/ornithology/ukab042
Kirkconnell A (2012) Cyanolimnas cerverai Barbour y Peters. In: González Alonso H, Rodríguez Schettino L, Rodríguez A, Mancina CA, Ramos García I (eds) Libro Rojo de los Vertebrados de Cuba. Instituto de Ecología y Sistemática, La Habana, Cuba
Kirkconnell A, González O, Alfaro E, Cotayo L (1999) Nuevas localidades para la Gallinuela de Santo Tomás Cyanolimnas cerverai y la Ferminia Ferminia cerverai en la Ciénaga de Zapata. Cuba Cotinga 12:57–60
Kirkconnell A, Stotz DF, Shopland JM (eds) (2005) Cuba: Peninsula de Zapata. Rapid Biological Inventories Report 07. The Field Museum, Chicago
Kirkconnell A, Kirwan G, Garrido O, Mitchell A, Wiley J (2020) The Birds of Cuba, an annotated checklist. British Ornithologists’ Club, Tring
Kirwan GM, Levesque A, Oberle MW, Sharpe CJ (2019) Birds of the West Indies. Lynx Edicions, Barcelona
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2016) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34:772–773. https://doi.org/10.1093/molbev/msw260
Lanfear R, von Haeseler A, Woodhams MD, Schrempf D, Chernomor O, Schmidt HA, Minh BQ, Teeling E (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534. https://doi.org/10.1093/molbev/msaa015
Livezey BC (1998) A phylogenetic analysis of the Gruiformes (Aves) based on morphological characters, with an emphasis on the rails (Rallidae). Philos Trans R Soc Lond B Biol Sci 353:2077–2151. https://doi.org/10.1098/rstb.1998.0353
Olson SL (1973) A classification of the Rallidae. Wilson Bull 85:381–416
Olson SL (1974) A new species of Nesotrochis from Hispaniola, with notes on other fossil rails from the West Indies (Aves: Rallidae). Proc Biol Soc Wash 87:439–450
Oswald JA, Terrill RS, Stucky BJ, LeFebvre MJ, Steadman DW, Guralnick RP, Allen JM (2021) Ancient DNA from the extinct Haitian cave-rail (Nesotrochis steganinos) suggests a biogeographic connection between the Caribbean and Old World. Biol Lett 17:20200760. https://doi.org/10.1098/rsbl.2020.0760
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Reynard G, Garrido O (1988) Bird songs in Cuba. Cornell University Laboratory of Ornithology, Ithaca
Ridgway R, Friedmann H (1941) The birds of North and Middle North America, Part IX. Bull United States Natl Mus. https://doi.org/10.5962/bhl.title.54021
Rojas-Consuegra R, Jiménez O, Condis-Fernández M, Díaz-Franco S (2012) Tafonomía y paleoecología de un yacimiento paleontológico del Cuaternario en la Cueva del Indio, La Habana. Cuba Espelunca Digital 12:1–12
Slikas B, Olson SL, Fleischer RC (2002) Rapid, independent evolution of flightlessness in four species of Pacific Island rails (Rallidae): an analysis based on mitochondrial sequence data. J Avian Biol 33:5–14. https://doi.org/10.1034/j.1600-048X.2002.330103.x
Steadman D, Morris J, Wright N (2013) A new species of Late Pleistocene rail (Aves: Rallidae) from Abaco, the Bahamas. Paleontol J 47:1355–1364
Stervander M, Ryan PG, Melo M, Hansson B (2019) The origin of the world’s smallest flightless bird, the Inaccessible Island Rail Atlantisia rogersi (Aves: Rallidae). Mol Phylogenet Evol 130:92–98. https://doi.org/10.1016/j.ympev.2018.10.007
Suárez W (2022) Catalogue of Cuban fossil and subfossil birds. Bull Br Ornithol Club 142:10–74. https://doi.org/10.25226/bboc.v142i1.2022.a3
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. Lect Math Life Sci 17:57–86
Taylor PB (1996) Family Rallidae (rails, gallinules and coots). In: del Hoyo J, Elliott A, Sargatal J (eds) Handbook of the birds of the world, vol 3. Hoatzin to Auks. Lynx Edicions, Barcelona, pp 108–209
Taylor B, Kirwan GM, Sharpe C (2020) Zapata Rail (Cyanolimnas cerverai), v 1.0. In: del Hoyo J, Elliott A, Sargatal J, Christie D, de Juana E (eds) Birds of the world. Cornell Lab of Ornithology, Ithaca. https://doi.org/10.2173/bow.zaprai1.01
Toews DPL, Brelsford A (2012) The biogeography of mitochondrial and nuclear discordance in animals. Mol Ecol 21:3907–3930. https://doi.org/10.1111/j.1365-294X.2012.05664.x
Wetmore A (1967) Further systematic notes on the avifauna of Panama. Proc Biol Soc Wash 80:229–242
Acknowledgements
GMK and AK are grateful to staff, particularly Ianela García-Lau, Manolo Barro and volunteers at the Museo de Historia Natural ‘Felipe Poey’, La Habana, Cuba, for access to relevant specimens. This study was funded by University of Aberdeen (AB) and The Sound Approach Ph.D. Studentship (TJS).
Author information
Authors and Affiliations
Contributions
Conceived and managed project: GMK, JMC, and MS. Provided biological sample: AK and GMK. Designed primers, performed molecular lab work: YL, AFB, TJS, and JMC. Assembled COI sequence: AFB and TJS. Performed phylogenetic analyses: MS. Wrote the manuscript: GMK and MS.
Corresponding authors
Additional information
Communicated by M. Wink.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Brown, A.F., Lawrie, Y., Shannon, T.J. et al. First genetic data for the critically endangered Cuban endemic Zapata Rail Cyanolimnas cerverai, and the taxonomic implications. J Ornithol 163, 945–952 (2022). https://doi.org/10.1007/s10336-022-02004-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10336-022-02004-0