Abstract
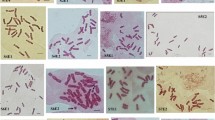
Fusarium oxysporum is an ascomycete fungus including plant pathogenic and nonpathogenic strains. Genome analyses have indicated that the karyotype of F. oxysporum is diverse among isolates. Here we used the germ tube burst method (GTBM), a more reliable method than conventional cytology or pulsed field gel electrophoretis, to karyotype isolates of F. oxysporum ff. spp. lycopersici and conglutinans and nonpathogenic F. oxysporum. In this first application of GTBM for F. oxysporum, pathogenic isolates were found to have more chromosomes than in nonpathogenic isolates. We also used a ribosomal DNA probe and fluorescence in situ hybridization (FISH) to analyze chromosome structure.




Similar content being viewed by others

References
Alves-Santos FM, Benito EP, Eslava AP, Díaz-Mínguez JM (1999) Genetic diversity of Fusarium oxysporum strains from common bean fields in Spain. Appl Environ Microbiol 65:3335–3340
Boehm EWA, Ploetz RC, Kistler HC (1994) Statistical analysis of electrophoretic karyotype variation among vegetative compatibility groups of Fusarium oxysporum f. sp. cubense. Mol Plant Microbe Interact 7:196–207
Garmaroodi HS, Taga M (2015) Meiotic inheritance of a fungal supernumerary chromosome and its effect on sexual fertility in Nectria haematococca. Fungal Biol 119:929–939
Imai S, Teraoka T, Arie T (2008) Generation of mating type locus-replaced transformant in Fusarium oxysporum (abstract in Japanese). Jpn J Phytopathol 74:39
Inami K, Yoshioka-Akiyama C, Morita Y, Yamasaki M, Teraoka T, Arie T (2012) A genetic mechanism for emergence of races in Fusarium oxysporum f. sp. lycopersici: inactivation of avirulence gene AVR1 by transposon insertion. PLoS One 7:e44101
Inami K, Kashiwa T, Kawabe M, Onokubo-Okabe A, Ishikawa N, Pérez ER, Hozumi T, Caballero LA, De Baldarrago FC, Roco MJ, Madadi KA, Peever TL, Teraoka T, Kodama M, Arie T (2014) The tomato wilt fungus Fusarium oxysporum f. sp. lycopersici shares common ancestors with nonpathogenic F. oxysporum isolated from wild tomatoes in the Peruvian Andes. Microbe Environ 29:200–210
Kashiwa T, Inami K, Fujinaga M, Ogiso H, Yoshida T, Teraoka T, Arie T (2013) An avirulence gene homologue in the tomato wilt fungus Fusarium oxysporum f. sp. lycopersici race 1 functions as a virulence gene in the cabbage yellows fungus F. oxysporum f. sp. conglutinans. J Gen Plant Pathol 79:412–421
Kashiwa T, Kozaki T, Ishii K, Turgeon BG, Teraoka T, Komatsu K, Arie T (2017) Sequencing of individual chromosomes of plant pathogenic Fusarium oxysporum. Fungal Genet Biol 98:46–51
Kawabe M, Kobayashi Y, Okada G, Yamaguchi I, Teraoka T, Arie T (2005) Three evolutionary lineages of tomato wilt pathogen, Fusarium oxysporum f. sp. lycopersici, based on sequences of IGS, MAT1, and pg1, are each composed of isolates of a single mating type and a single or closely related vegetative compatibility group. J Gen Plant Pathol 71:263–272
Kim DH, Martyn RD, Magill CW (1993) Chromosome polymorphism in Fusarium oxysporum f. sp. niveum. Phytopathology 83:1209–1216
Kuroiwa T, Kojima H, Miyakawa I, Sando N (1984) Meiotic karyotype of the yeast Saccharomyces cerevisiae. Exp Cell Res 153:259–265
Lu BCK (1996) Chromosomes, mitosis, and meiosis. In: Bos CJ (ed) Fungal genetics: principles and practice. Marcel Dekker, New York, NY, USA, pp 119–176
Ma LJ, van der Does HC, Borkovich KA, Coleman JJ, Daboussi MJ, Di Pietro A, Dufresne M, Freitag M, Grabherr M, Henrissat B, Houterman PM, Kang S, Shim WB, Woloshuk C, Xie X, Xu JR, Antoniw J, Baker SE, Bluhm BH, Breakspear A, Brown DW, Butchko RAE, Chapman S, Coulson R, Coutinho PM, Danchin EGJ, Diener A, Gale LR, Gardiner DM, Goff S, Hammond-Kosack KE, Hilburn K, Hua-Van A, Jonkers W, Kazan K, Kodira CD, Koehrsen M, Kumar L, Lee YH, Li L, Manners JM, Miranda-Saavedra D, Mukherjee M, Park G, Park J, Park SY, Proctor RH, Regev A, Ruiz-Roldan MC, Sain D, Sakthikumar S, Sykes S, Schwartz DC, Turgeon BG, Wapinski I, Yoder O, Young S, Zeng Q, Zhou S, Galagan J, Cuomo CA, Kistler HC, Rep M (2010) Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464:367–373
Mahmoud AM. Taga M (2012) Cytological karyotyping and characterization of a 410 kb minichromosome in Nectria haematococca MPI. Mycologia 104:845–856
Mehrabi R, Taga M, Aghaee M, de Wit PJGM, Kema GHJ (2012) Karyotyping methods for fungi. In: Bolton M, Thomma B (eds) Plant fungal pathogens. Methods in molecular biology (methods and protocols), vol 835. Humana Press, New York, NY, USA, pp 591–602
Migheli Q, Berio T, Gullino ML (1993) Electrophoretic karyotypes of Fusarium spp. Exp Mycol 17:329–337
Migheli Q, Berio T, Gullino ML, Garibaldi A (1995) Electrophoretic karyotype variation among pathotypes of Fusarium oxysporum f. sp. dianthi. Plant Pathol 44:308–315
Min BR (1995) Comparison of electrophoretic karyotypes in Fusarium. J Microbiol 33:334–338
Min BR, Kim KA, Choi YK (1998) Electrophoretic karyotypes of Fusarium oxysporum formae speciales. J Microbiol 36:14–19
Momol EA, Kistler HC (1992) Mitochondrial plasmids do not determine host range in crucifer-infecting strains of Fusarium oxysporum. Plant Pathol 41:103–112
Ogawa K, Komada H (1984) Biological control of Fusarium wilt of sweet potato by non-pathogenic Fusarium oxysporum. Ann Phytopathol Soc Jpn 50:1–9
Punithalingam E (1975) Cytology of some Fusarium species. Nova Hedwigia 26:275–303
Rosewich UL, Pettway RE, Katan T, Kistler HC (1999) Population genetic analysis corroborates dispersal of Fusarium oxysporum f. sp. radicis-lycopersici from Florida to Europe. Phytopathology 89:623–630
Shahi S, Beerens B, Bosch M, Linmans J, Rep M (2016) Nuclear dynamics and genetic rearrangement in heterokaryotic colonies of Fusarium oxysporum. Fungal Genet Biol 91:20–31
Shirane N, Masuko M, Hayashi Y (1988) Nuclear behavior and division in germinating conidia of Botrytis cinerea. Phytopathology 78:1627–1630
Shirane N, Masuko M, Hayashi Y (1989) Light microscopic observation of nuclei and mitotic chromosomes of Botrytis species. Phytopathology 79:728–730
Shishido M, Miwa C, Usami T, Amemiya Y, Johnson KB (2005) Biological control efficiency of Fusarium wilt of tomato by nonpathogenic Fusarium oxysporum Fo-B2 in different environments. Phytopathology 95:1072–1080
Taga M, Murata M (1994) Visualisation of mitotic chromosomes in filamentous fungi by fluorescence staining and fluorescence in situ hybridization. Chromosoma 103:408–413
Taga M, Murata M, Saito H (1998) Comparison of different karyotyping methods in filamentous ascomycetes—a case study of Nectria haematococca. Mycol Res 102:1355–1364
Taga M, Murata M, VanEtten HD (1999) Visualization of a conditionally dispensable chromosome in the filamentous ascomycete Nectria haematococca by fluorescence in situ hybridization. Fungal Genet Biol 26:169–177
Taga M, Tsuchiya D, Murata M (2003) Dynamic changes of rDNA condensation state during mitosis in filamentous fungi revealed by fluorescence in situ hybridisation. Mycol Res 107:1012–1020
Taga M, Tanaka K, Kato S, Kubo Y (2015) Cytological analyses of the karyotypes and chromosomes of three Colletotrichum species, C. orbiculare, C. graminicola and C. higginsianum. Fungal Genet Biol 82:238–250
To-Anun C, Nelson H, Ouchi S (1995) Electrophoretic karyotyping of Fusarium oxysporum. Ann Phytopathol Soc Jpn 61:350–356
Tsuchiya D, Taga M (2001) Cytological karyotyping of three Cochliobolus spp. by the germ tube burst method. Phytopathology 91:354–360
Tsuchiya D, Taga M (2010) Fluorescence in situ hybridization for molecular cytogenetic analysis in filamentous fungi. In: Sharon A (ed) Molecular and cell biology methods for fungi. Methods in molecular biology (methods and protocols), vol 638. Humana Press, New York, NY, USA, pp 235–257
Tsuge T, Kobayashi H, Nishimura S (1989) Organization of ribosomal RNA genes in Alternaria alternata Japanese pear pathotype, a host-selective AK-toxin-producing fungus. Curr Genet 16:267–272
van Dam P, Fokkens L, Ayukawa Y, van der Gragt M, ter Horst A, Brankovics B, Houterman PM, Arie T, Rep M (2017) A mobile pathogenicity chromosome in Fusarium oxysporum for infection of multiple cucurbit species. Sci Rep 7:9042
Vlaardingerbroek I, Beerens B, Rose L, Fokkens L, Cornelissen BJC, Rep M (2016a) Exchange of core chromosomes and horizontal transfer of lineage-specific chromosomes in Fusarium oxysporum. Environ Microbiol 18:3702–3713
Vlaardingerbroek I, Beerens B, Schmidt SM, Cornelissen BJC, Rep M (2016b) Dispensable chromosomes in Fusarium oxysporum f. sp. lycopersici. Mol Plant Pathol 17:1455–1466
Acknowledgements
This study was partly supported by Grants-in-Aid from The Japan Society for the Promotion of Sciences (JSPS) to TA (16H02536 and 18380030) and Building of Consortia for the Development of Human Resources in Science and Technology (Innovation Advancement Organization, TUAT) from Ministry of Education, Culture, Sports, Science and Technology of Japan. We thank ASKA Pharmaceutical (Tokyo, Japan) for providing Driselase. We are grateful to Dr. Yoshimiki Amemiya (Chiba University, Matsudo, Japan) for providing a fungal isolate and Dr. Takashi Tsuge (Nagoya University, Nagoya, Japan) for providing plasmid DNA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ayukawa, Y., Komatsu, K., Taga, M. et al. Cytological karyotyping of Fusarium oxysporum by the germ tube burst method (GTBM). J Gen Plant Pathol 84, 254–261 (2018). https://doi.org/10.1007/s10327-018-0784-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10327-018-0784-5