Abstract
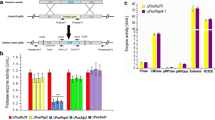
Gluconobacter oxydans is capable of rapidly incomplete oxidation of many sugars and alcohols, which means the strain has great potential for industrial purposes. Strong promoters are one of the essential factors that can improve strain performance by overexpression of specific genes. In this study, a pipeline for screening strong promoters by proteomics analysis was established. Based on the procedure, a new strong promoter designated as P B932_2000 was identified in G. oxydans WSH-003. The promoter region was characterized based on known genome sequence information using BPROM. The strength of P B932_2000 was further assessed by analysis of enhanced green fluorescent protein (egfp) expression and comparison with egfp expression by two commonly used strong promoters, P E. coli_tufB and P G. oxydans_tufB . Both quantitative real-time PCR and fluorescence intensities for egfp gene expression showed that P B932_2000 promoter is stronger than the other two. Overexpression of d-sorbitol dehydrogenase (sldh) by P B932_2000 in G. oxydans WSH-003 enhanced the titer and productivity of l-sorbose synthesis from d-sorbitol by 12.0 % and 33.3 %, respectively. These results showed that proteomics analysis is an efficient way to identify strong promoters. The isolated promoter P B932_2000 could further facilitate the metabolic engineering of G. oxydans.




Similar content being viewed by others
References
Anderson NL, Anderson NG (1998) Proteome and proteomics: new technologies, new concepts, and new words. Electrophoresis 19:1853–1861
Battey AS, Schaffner DW (2001) Modelling bacterial spoilage in cold-filled ready to drink beverages by Acinetobacter calcoaceticus and Gluconobacter oxydans. J Appl Microbiol 91:237–247
Blackstock WP, Weir MP (1999) Proteomics: quantitative and physical mapping of cellular proteins. Trends Biotechnol 17:121–127
Bryksin AV, Matsumura I (2010) Overlap extension PCR cloning: a simple and reliable way to create recombinant plasmids. Biotechniques 48:463–465
Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC (1994) Green fluorescent protein as a marker for gene-expression. Science 263:802–805
Cui Z, Zhang X, Zhang Z, Li S (2004) Construction and application of a promoter-trapping vector with methyl parathion hydrolase gene mpd as the reporter. Biotechnol Lett 26(14):1115–1118
De Muynck C, Pereira C, Naessens M, Parmentier S, Soetaert W, Vandamme E (2007) The genus Gluconobacter oxydans: comprehensive overview of biochemistry and biotechnological applications. Crit Rev Biotechnol 27:147–171
Fukui K, Koseki C, Yamamoto Y, Nakamura J, Sasahara A, Yuji R, Hashiguchi K, Usuda Y, Matsui K, Kojima H, Abe K (2011) Identification of succinate exporter in Corynebacterium glutamicum and its physiological roles under anaerobic conditions. J Biotechnol 154:25–34
Gao L, Hu Y, Liu J, Du G, Zhou J, Chen J (2014) Stepwise metabolic engineering of Gluconobacter oxydans WSH-003 for the direct production of 2-keto-l-gulonic acid from d-sorbitol. Metab Eng 24:30–37
Gao L, Zhou J, Liu J, Du G, Chen J (2012) Draft genome sequence of Gluconobacter oxydans WSH-003, a strain that is extremely tolerant of saccharides and alditols. J Bacteriol 194(16):4455–4456
Gatgens C, Degner U, Bringer-Meyer S, Herrmann U (2007) Biotransformation of glycerol to dihydroxyacetone by recombinant Gluconobacter oxydans DSM 2343. Appl Microbiol Biotechnol 76:553–559
Gupta A, Singh VK, Qazi GN, Kumar A (2001) Gluconobacter oxydans: its biotechnological application. J Mol Microbiol Biotechnol 3:445–456
He J, Thomas K, Leif JJ (2012) Comparative proteome analysis of Saccharomyces cerevisiae: a global overview of in vivo targets of the yeast activator protein 1. BMC Genom 13:230
Ikeda M, Mizuno Y, Awane S, Hayashi M, Mitsuhashi S, Takeno S (2011) Identification and application of a different glucose uptake system that functions as an alternative to the phosphotransferase system in Corynebacterium glutamicum. Appl Microbiol Biotechnol 90:1443–1451
Ji AG, Gao PJ (2001) Substrate selectivity of Gluconobacter oxydans for production of 2,5-diketo-d-gluconic acid and synthesis of 2-keto-l-gulonic acid in a multienzyme system. Appl Biochem Biotechnol 94:213–223
Jopcik M, Bauer M, Moravcikova J, Boszoradova E, Matusikova I, Libantova J (2013) Plant tissue-specific promoters can drive gene expression in Escherichia coli. Plant Cell Tissue Organ Cult 113:387–396
Kovach ME, Elzer PH, Hill DS, Robertson GT, Farris MA, Roop RM, Peterson KM (1995) 4 new derivatives of the broad-host-range cloning vector PBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166:175–176
Lee JS, An G, Friesen JD, Fill NP (1981) Location of the tufB promoter of E. coli: cotranscription of tufB with four transfer RNA genes. Cell 25:251–258
Lu L, Wei L, Zhu K, Wei D, Hua Q (2012) Combining metabolic engineering and adaptive evolution to enhance the production of dihydroxyacetone from glycerol by Gluconobacter oxydans in a low-cost way. Bioresour Technol 117:317–324
Matsushita K, Toyama H, Adachi O (1994) Respiratory chains and bioenergetics of acetic acid bacteria. Adv Microb Physiol 36:247–301
Merfort M, Herrmann U, Bringer-Meyer S, Sahm H (2006) High-yield 5-keto-d-gluconic acid formation is mediated by soluble and membrane-bound gluconate-5-dehydrogenases of Gluconobacter oxydans. Appl Microbiol Biotechnol 73:443–451
Neuhoff V, Arold N, Taube D, Ehrhardt W (1988) Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis 9:255–262
Pátek M, Nešvera J (2012) Promoters and plasmid vectors of Corynebacterium glutamicum. In: Yukawa H, Inui M (eds) Biology and biotechnology of Corynebacterium glutamicum. Springer, Berlin, pp 51–88
Patek M, Holatko J, Busche T, Kalinowski J, Nesvera J (2013) Corynebacterium glutamicum promoters: a practical approach. Microb Biotechnol 6:103–117
Patterson GH, Knobel SM, Sharif WD, Kain SR, Piston DW (1997) Use of the green fluorescent protein and its mutants in quantitative fluorescence microscopy. Biophys J 73:2782–2790
Peters B, Junker A, Brauer K, Muhlthaler B, Kostner D, Mientus M, Liebl W, Ehrenreich A (2013) Deletion of pyruvate decarboxylase by a new method for efficient markerless gene deletions in Gluconobacter oxydans. Appl Microbiol Biotechnol 97:2521–2530
Prust C, Hoffmeister M, Liesegang H, Wiezer A, Fricke WF, Ehrenreich A, Gottschalk G, Deppenmeier U (2005) Complete genome sequence of the acetic acid bacterium Gluconobacter oxydans. Nat Biotechnol 23:195–200
Rhodius VA, Mutalik VK (2010) Predicting strength and function for promoters of the Escherichia coli alternative sigma factor, σE. Proc Natl Acad Sci USA 107(7):2854–2859
Richard G (2010) Prokaryotic transcription. In: Kebs Jocelyn E, Kilpatrick Stephen T, Goldstein Elliott S (eds) Lewein’s gene X. Jones and Bartlett Publishers, Massachusetts, pp 514–515
Saito Y, Ishii Y, Hayashi H, Imao Y, Akashi T (1997) Cloning genes coding for l-sorbose and l-sorbosone dehydrogenase from Gluconobacter oxydans and microbial production of 2-keto-l-gulonate, a precursor of l-ascorbic acid, in a recombinant G. oxydans strain. Appl Environ Microbiol 63:454–460
Schleyer U, Bringer-Meyer S, Sahm H (2008) An easy cloning and expression vector system for Gluconobacter oxydans. Int J Food Microbiol 125:91–95
Schmid R, Gerloff DL (2004) Functional properties of the alternative NADH:ubiquinone oxidoreductase from E. coli through comparative 3-D modelling. FEBS Lett 578:163–168
Sharma CM, Hoffmann S, Darfeuille F, Reignier J, Findeiss S, Sittka A, Chabas S, Reiche K, Hackermuller J, Reinhardt R, Stadler PF, Vogel J (2010) The primary transcriptome of the major human pathogen Helicobacter pylori. Nature 464(7286):250–255
Shi L, Li K, Zhang H, Liu X, Lin J, Wei D (2014) Identification of a novel promoter gHp0169 for gene expression in Gluconobacter oxydans. J Biotechnol 175:69–74
Siegl T, Tokovenko B, Myronovskyi M, Luzhetskyy A (2013) Design, construction and characterisation of a synthetic promoter library for fine-tuned gene expression in actinomycetes. Metab Eng 19:98–106
Siekevitz M, Feinberg MB, Holbrook N, Wong-Staal F, Greene WC (1987) Activation of interleukin 2 and interleukin 2 receptor (Tac) promoter expression by the trans-activator (tat) gene product of human T-cell leukemia virus, type I. Proc Natl Acad Sci USA 84:5389–5393
Singh BR, Al-Khedhairy AA, Alarifi SA, Musarrat J (2009) Regulatory elements in the 5′region of 16SrRNA gene of Bacillus sp. strain SJ-101. Bioinformation 3:375–380
Soboleski MR, Oaks J, Halford WP (2005) Green fluorescent protein is a quantitative reporter of gene expression in individual eukaryotic cells. FASEB J 19:440–442
Solovyev V, Salamov A (2011) Automatic annotation of microbial genomes and metagenomic sequences. In: Li RW (ed) Metagenomics and its applications in agriculture, biomedicine and environmental studies. Nova Science Publishers, New York, pp 61–78
Solovyev VV, Shahmuradov IA, Salamov AA (2010) Identification of promoter regions and regulatory sites. Methods Mol Biol 674:57–83
Srivastava S, Singh V, Kumar V, Verma PC, Srivastava R, Basu V, Gupta V, Rawat AK (2008) Identification of regulatory elements in 16S rRNA gene of Acinetobacter species isolated from water sample. Bioinformation 3:173–176
Wang J, Ai X, Mei H, Fu Y, Chen B, Yu Z, He J (2013) High-throughput identification of promoters and screening of highly active promoter-5′-UTR DNA region with different characteristics from Bacillus thuringiensis. PLoS ONE 8(5):e62960
Xu Y, Liu Q, Zhou L, Yang Z, Zhang Y (2008) Surface display of GFP by Pseudomonas syringae truncated ice nucleation protein in attenuated Vibrio anguillarum strain. Mar Biotechnol 10:701–708
Yang M (2013) Generation of an artificial double promoter for protein expression in Bacillus subtilis through a promoter trap system. PLoS ONE 8(2):e56321
Yang X, Wei L, Lin J, Yin B, Wei D (2008) Membrane-bound pyrroloquinoline quinone-dependent dehydrogenase in Gluconobacter oxydans M5, responsible for production of 6-(2-hydroxyethyl) amino-6-deoxy-l-sorbose. Appl Environ Microbiol 74:5250–5253
Zhou J, Du G, Chen J (2012) Metabolic engineering of microorganisms for vitamin C production. Subcell Biochem 64:241–259
Zhou J, Wang K, Xu S, Wu J, Liu P, Du G, Li J, Chen J (2015) Identification of membrane proteins associated with phenylpropanoid tolerance and transport in Escherichia coli BL21. J Proteomics 113:15–28
Zhu Y, Liu J, Liu J, Du G, Zhou J, Chen J (2012) A high throughput method to screen companion bacterium for 2-keto-l-gulonic acid biosynthesis by co-culturing Ketogulonicigenium vulgare. Process Biochem 47:1428–1432
Acknowledgments
This work was supported by grants from the National High Technology Research and Development Program of China (863 Program, 2012AA022103), the Major State Basic Research Development Program of China (973 Program, 2014CB745103), the Major Program of National Natural Science Foundation of China (21390204), the Program for New Century Excellent Talents in University (NCET-12-0876), the Foundation for the Author of National Excellent Doctoral Dissertation of PR China (FANEDD, 201256), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and the 111 Project (111-2-06).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hu, Y., Wan, H., Li, J. et al. Enhanced production of l-sorbose in an industrial Gluconobacter oxydans strain by identification of a strong promoter based on proteomics analysis. J Ind Microbiol Biotechnol 42, 1039–1047 (2015). https://doi.org/10.1007/s10295-015-1624-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-015-1624-7