Abstract
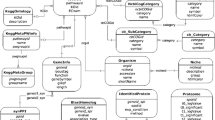
Under stressful conditions, the non-model marine microalga Tetraselmis subcordiformis can accumulate a substantial amount of starch, making it a potential feedstock for the production of fuel ethanol. Investigating the interactions of the enzymes and the regulatory factors involved in starch metabolism will provide potential genetic manipulation targets for optimising the starch productivity of T. subcordiformis. For this reason, the proteome of T. subcordiformis was utilised to predict the first protein–protein interaction (PPI) network for this marine alga based on orthologous interactions, mainly from the general PPI repositories. Different methods were introduced to evaluate the credibility of the predicted interactome, including the confidence value of each PPI pair and Pfam-based and subcellular location-based enrichment analysis. Functional subnetworks analysis suggested that the two enzymes involved in starch metabolism, starch phosphorylase and trehalose-phosphate synthase may be the potential ideal genetic engineering targets.




Similar content being viewed by others
References
Barabasi AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113. doi:10.1038/nrg1272
Beer LL, Boyd ES, Peters JW, Posewitz MC (2009) Engineering algae for biohydrogen and biofuel production. Curr Opin Biotech 20:264–271. doi:10.1016/j.copbio.2009.06.002
Boersema PJ, Raijmakers R, Lemeer S, Mohammed S, Heck AJR (2009) Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat Protocols 4:484–494. doi:10.1038/nprot.2009.21
Boxem M, Maliga Z, Klitgord N, Li N, Lemmens I, Mana M, de Lichtervelde L, Mul JD, van de Peut D, Devos M, Simonis N, Yildirim MA, Cokol M, Kao H-L, de Smet A-S, Wang H, Schlaitz A-L, Hao T, Milstein S, Fan C, Tipsword M, Drew K, Galli M, Rhrissorrakrai K, Drechsel D, Koller D, Roth FP, Iakoucheva LM, Dunker AK, Bonneau R, Gunsalus KC, Hill DE, Piano F, Tavernier J, van den Heuvel S, Hyman AA, Vidal M (2008) A protein domain-based interactome network for C. elegans early embryogenesis. Cell 134:534–545. doi:10.1016/j.cell.2008.07.009
Chatr-aryamontri A, Breitkreutz B-J, Heinicke S, Boucher L, Winter A, Stark C, Nixon J, Ramage L, Kolas N, O’Donnell L, Reguly T, Breitkreutz A, Sellam A, Chen D, Chang C, Rust J, Livstone M, Oughtred R, Dolinski K, Tyers M (2013) The BioGRID interaction database: 2013 update. Nucleic Acids Res 41:D816–D823. doi:10.1093/nar/gks1158
Chatr-aryamontri A, Ceol A, Palazzi LM, Nardelli G, Schneider MV, Castagnoli L, Cesareni G (2007) MINT: the molecular interaction database. Nucleic Acids Res 35:D572–D574. doi:10.1093/nar/gkl950
Chou K-C, Shen H-B (2010) Plant-mPLoc: a top-down strategy to augment the power for predicting plant protein subcellular localization. PLoS One 5:e11335. doi:10.1371/journal.pone.0011335
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. doi:10.1093/bioinformatics/bti610
Florez A, Park D, Bhak J, Kim B-C, Kuchinsky A, Morris J, Espinosa J, Muskus C (2010) Protein network prediction and topological analysis in Leishmania major as a tool for drug target selection. BMC Bioinform 11:484. doi:10.1186/1471-2105-11-484
Gandhi TKB, Zhong J, Mathivanan S, Karthick L, Chandrika KN, Mohan SS, Sharma S, Pinkert S, Nagaraju S, Periaswamy B, Mishra G, Nandakumar K, Shen B, Deshpande N, Nayak R, Sarker M, Boeke JD, Parmigiani G, Schultz J, Bader JS, Pandey A (2006) Analysis of the human protein interactome and comparison with yeast, worm and fly interaction datasets. Nat Genet 38:285–293. doi:10.1038/ng1747
Geisler-Lee J, O’Toole N, Ammar R, Provart NJ, Millar AH, Geisler M (2007) A predicted interactome for Arabidopsis. Plant Physiol 145:317–329. doi:10.1104/pp.107.103465
Guo J, Li H, Chang J-W, Lei Y, Li S, Chen L–L (2013) Prediction and characterization of protein–protein interaction network in Xanthomonas oryzae pv. oryzae PXO99A. Res Microbiol 164:1035–1044. doi:10.1016/j.resmic.2013.09.001
He F, Zhang Y, Chen H, Zhang Z, Peng Y-L (2008) The prediction of protein–protein interaction networks in rice blast fungus. BMC Genom 9:519. doi:10.1186/1471-2164-9-519
Hennen-Bierwagen TA, Lin Q, Grimaud F, Planchot V, Keeling PL, James MG, Myers AM (2009) Proteins from multiple metabolic pathways associate with starch biosynthetic enzymes in high molecular weight complexes: a model for regulation of carbon allocation in maize amyloplasts. Plant Physiol 149:1541–1559. doi:10.1104/pp.109.135293
Hennen-Bierwagen TA, Liu F, Marsh RS, Kim S, Gan Q, Tetlow IJ, Emes MJ, James MG, Myers AM (2008) Starch biosynthetic enzymes from developing maize endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908. doi:10.1104/pp.108.116285
Ho C-L, Wu Y, Shen H-b, Provart N, Geisler M (2012) A predicted protein interactome for rice. Rice 5:1–14. doi:10.1186/1939-8433-5-15
Jeong H, Mason SP, Barabasi AL, Oltvai ZN (2001) Lethality and centrality in protein networks. Nature 411:41–42. doi:10.1038/35075138
Kötting O, Kossmann J, Zeeman SC, Lloyd JR (2010) Regulation of starch metabolism: the age of enlightenment? Curr Opin Plant Biol 13:320–328. doi:10.1016/j.pbi.2010.01.003
Lee DY, Park J–J, Barupal DK, Fiehn O (2012) System response of metabolic networks in Chlamydomonas reinhardtii to total available ammonium. Mol Cell Proteomics 11:973–988. doi:10.1074/mcp.M111.016733
Lemeer S, Jopling C, Gouw J, Mohammed S, Heck AJR, Slijper M, den Hertog J (2008) Comparative phosphoproteomics of zebrafish Fyn/Yes morpholino knockdown embryos. Mol Cell Proteomics 7:2176–2187. doi:10.1074/mcp.M800081-MCP200
Liu Z-P, Chen L (2012) Proteome-wide prediction of protein–protein interactions from high-throughput data. Protein Cell 3:508–520. doi:10.1007/s13238-012-2945-1
Östlund G, Schmitt T, Forslund K, Köstler T, Messina DN, Roopra S, Frings O, Sonnhammer ELL (2010) InParanoid 7: new algorithms and tools for eukaryotic orthology analysis. Nucleic Acids Res 38:D196–D203. doi:10.1093/nar/gkp931
Pignolet O, Jubeau S, Vaca-Garcia C, Michaud P (2013) Highly valuable microalgae: biochemical and topological aspects. J Ind Microbiol Biotechnol 40:781–796. doi:10.1007/s10295-013-1281-7
Procházková G, Brányiková I, Zachleder V, Brányik T (2013) Effect of nutrient supply status on biomass composition of eukaryotic green microalgae. J Appl Phycol pp 1–19. doi:10.1007/s10811-013-0154-922
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J, Heger A, Holm L, Sonnhammer ELL, Eddy SR, Bateman A, Finn RD (2012) The Pfam protein families database. Nucleic Acids Res 40:D290–D301. doi:10.1093/nar/gkr1065
Saito R, Smoot ME, Ono K, Ruscheinski J, Wang P, Lotia S, Pico AR, Bader GD, Ideker T (2012) A travel guide to cytoscape plugins. Nat Meth 9:1069–1076. doi:10.1038/nmeth.2212
Smoot ME, Ono K, Ruscheinski J, Wang P, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27:431–432. doi:10.1093/bioinformatics/btq675
Tetlow IJ, Beisel KG, Cameron S, Makhmoudova A, Liu F, Bresolin NS, Wait R, Morell MK, Emes MJ (2008) Analysis of protein complexes in wheat amyloplasts reveals functional interactions among starch biosynthetic enzymes. Plant Physiol 146:1878–1891. doi:10.1104/pp.108.116244
Tetlow IJ, Wait R, Lu Z, Akkasaeng R, Bowsher CG, Esposito S, Kosar-Hashemi B, Morell MK, Emes MJ (2004) Protein phosphorylation in amyloplasts regulates starch branching enzyme activity and protein–protein interactions. Plant Cell 16:694–708. doi:10.1105/tpc.017400
van Breukelen B, van den Toorn HWP, Drugan MM, Heck AJR (2009) StatQuant: a post-quantification analysis toolbox for improving quantitative mass spectrometry. Bioinformatics 25:1472–1473. doi:10.1093/bioinformatics/btp181
Walhout AJM, Sordella R, Lu X, Hartley JL, Temple GF, Brasch MA, Thierry-Mieg N, Vidal M (2000) Protein interaction mapping in C. elegans using proteins involved in vulval development. Science 287:116–122. doi:10.1126/science.287.5450.116
Xenarios I, Salwínski L, Duan XJ, Higney P, Kim S-M, Eisenberg D (2002) DIP, the database of interacting proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res 30:303–305. doi:10.1093/nar/30.1.303
Y ao C, Ai J, Cao X, Xue S (2013) Characterization of cell growth and starch production in the marine green microalga Tetraselmis subcordiformis under extracellular phosphorus-deprived and sequentially phosphorus-replete conditions. Appl Microbiol Biot 97:6099–6110. doi:10.1007/s00253-013-4983-x
Yao C, Ai J, Cao X, Xue S (2013) Salinity manipulation as an effective method for enhanced starch production in the marine microalga Tetraselmis subcordiformis. Biores Technol 146:663–671. doi:10.1016/j.biortech.2013.07.134
Yao C, Ai J, Cao X, Xue S, Zhang W (2012) Enhancing starch production of a marine green microalga Tetraselmis subcordiformis through nutrient limitation. Bioresource Technol 118:438–444. doi:10.1016/j.biortech.2012.05.030
Acknowledgments
The authors gratefully acknowledge the support of the Hundred Talent Program of the Chinese Academy of Sciences (No. A1097) and thank Dr. Hongwei Liu and Mr. Keyue Wang for the proteomic data collection.
Author information
Authors and Affiliations
Corresponding authors
Additional information
C. Ji and X. Cao have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ji, C., Cao, X., Yao, C. et al. Protein–protein interaction network of the marine microalga Tetraselmis subcordiformis: prediction and application for starch metabolism analysis. J Ind Microbiol Biotechnol 41, 1287–1296 (2014). https://doi.org/10.1007/s10295-014-1462-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-014-1462-z