Abstract
This work aimed to investigate whether global radiomic features (GRFs) from mammograms can predict difficult-to-interpret normal cases (NCs). Assessments from 537 readers interpreting 239 normal mammograms were used to categorise cases as 120 difficult-to-interpret and 119 easy-to-interpret based on cases having the highest and lowest difficulty scores, respectively. Using lattice- and squared-based approaches, 34 handcrafted GRFs per image were extracted and normalised. Three classifiers were constructed: (i) CC and (ii) MLO using the GRFs from corresponding craniocaudal and mediolateral oblique images only, based on the random forest technique for distinguishing difficult- from easy-to-interpret NCs, and (iii) CC + MLO using the median predictive scores from both CC and MLO models. Useful GRFs for the CC and MLO models were recognised using a scree test. The CC and MLO models were trained and validated using the leave-one-out-cross-validation. The models’ performances were assessed by the AUC and compared using the DeLong test. A Kruskal–Wallis test was used to examine if the 34 GRFs differed between difficult- and easy-to-interpret NCs and if difficulty level based on the traditional breast density (BD) categories differed among 115 low-BD and 124 high-BD NCs. The CC + MLO model achieved higher performance (0.71 AUC) than the individual CC and MLO model alone (0.66 each), but statistically non-significant difference was found (all p > 0.05). Six GRFs were identified to be valuable in describing difficult-to-interpret NCs. Twenty features, when compared between difficult- and easy-to-interpret NCs, differed significantly (p < 0.05). No statistically significant difference was observed in difficulty between low- and high-BD NCs (p = 0.709). GRF mammographic analysis can predict difficult-to-interpret NCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among women globally, breast cancer is the most prevalent cancer type and leading cause of cancer-related death [1]. While mammography screening has played a major role in the early detection of breast cancer for decades [2,3,4], mammographic image interpretation is an extremely complex process with reported false-positive (FP) error rates of up to 16% and estimated cumulative risk of FP results of up to 63% in women 50–69 years of age after ten biennial mammographic screening rounds [5, 6]. A study reported that 80% of women having abnormal mammographic findings at screening were recalled for further diagnostic procedures that eventually resolved with normal outcomes and 40% of those went through invasive biopsy which subsequently followed in benign findings [7]. These unnecessary recalls (i.e., FP findings) have placed huge burdens on patients and healthcare systems, causing considerable psychological distress and anxiety as well as unwarranted medical costs (e.g., USD$2.8 billion each year in the USA) [8,9,10,11]. Considering that approximately 99.5% of cases in screening programs are normal, the ability of radiologists to interpret and detect normal cases (NCs) correctly is extremely important [12, 13].
Educational test sets are powerful web-based platforms shown to improve radiologists’ performance in interpreting mammograms [14,15,16,17,18,19,20,21,22]. Since NCs often constitute almost 70% of the cases in these educational test sets, the capability to identify normal mammographic features can be well assessed [19, 20, 23]. However, a consistent difficulty for normal mammographic cases across different test sets is important in order to facilitate a reliable and efficient objective measurement of FP errors and the performance of individual reader, since test set based education is widely used worldwide as a means of monitoring clinical performance [16,17,18,19,20, 22,23,24,25]. This implies that if a radiologist’s performance changes over time, it should be a function of the radiologist, not the test set conditions. Considering readers’ performance in educational test sets can largely signify their clinical performance; measured by clinical audits, a consistent difficulty of NCs in test set based education is also vital for clinical screening programs. It can not only enable uniform empirical measurement of the screening program’s recall rate performance over time, but also facilitate the establishment of national standards of recall rate that could be used to compare with the recall rate performance of individual breast reader as part of a quality improvement [15, 20, 22,23,24,25].
To date, there appears to be no reliable method of standardising test set difficulty for NCs since test set generation currently relies on experts’ assessment of case difficulty, where suitable NCs are selected manually by expert radiologists, often based on the traditional breast imaging and reporting data system (BI-RADS) breast density (BD) category and inclusion of benign features [18, 26,27,28]. Studies, however, have revealed that the difficulty of NCs subjectively defined by experts or perceived by individuals based on the BD classification rarely aligns to the actual difficulty based on readers’ performance in the test set [28,29,30]. Although a retrospective assessment of normal case difficulty is performed by calculating the numbers of errors made by readers engaging with the test set, this is rare, and indeed when it occurs, it relies on the efficacy of the readers of that test set which is still not a good descriptor of the actual difficulty of NCs [18, 26, 28, 29]. Consequently, there is an urgent need for an alternative standardised method of predicting NCs’ difficulty so that readers’ performance and individual specific FP errors can be better monitored, identified, addressed, and improved.
This study aims to provide a reliable method for automatically describing the difficulty of NCs (i.e., an objective measure of difficulty based on readers’ error rate [31]) using global image features [32] derived from radiomics (extracting and analysing a large number of quantitative features from medical images and converting them into multi-dimensional data) [33,34,35,36,37,38]. Radiomics in mammography, to date, has focused on predicting readers’ error making patterns using local radiomic features of specific tumour/FP areas, achieving up to 0.61 AUC [39, 40]. Previous studies have also shown that radiologists detect the gist of “abnormal” based on overall image appearance immediately after image onset [41,42,43]. Considering the importance of the initial global impression in guiding a visual search and making a diagnostic decision [41,42,43,44,45] as well as the value of radiomics in mammography [38], we hypothesised that global image statistics (i.e., global radiomic features/GRFs) of difficult-to-interpret NCs differed from easy-to-interpret ones. However, little attempt has been made to predict difficult-to-interpret NCs based on GRFs. This study addresses this deficiency. Moreover, since test set curation often depends on the traditional BI-RADS BD classification and higher BD cases have previously been reported to be more difficult to interpret [28, 30, 46], this work also investigates if there is a difference in difficulty level between low-BD (BI-RADS BD category A = fatty and B = fibroglandular) vs high-BD (BI-RADS BD category C = heterogeneous and D = extremely dense) NCs [47].
Materials and Methods
Ethical approval was obtained for this study from the Human Research Ethics Committee of the University of Sydney [2019/013 and 2017/028] with informed consent acquired from each reader. This study only involved retrospective analysis of existing and non-identifiable data collected as part of the BreastScreen Reader Assessment Strategy (BREAST) databank (https://breast-australia.sydney.edu.au/) [48, 49], which is funded by the Commonwealth of Australia to improve observer performance in BreastScreen Australia.
Readers
A total of 537 readers were included in this study, with expert breast Australian and New Zealand radiologists (n = 382), breast physicians (n = 18, medical practitioners specializing in interpreting breast images such as mammograms, and diagnosis and management of benign and malignant breast disease, in partnership with breast surgeons, radiologists, oncologists, pathologists and geneticists), and radiology trainees (n = 137) who engaged with the BREAST program from 04 September 2014 to 17 March 2021. These readers were recruited either through a radiology and/or breast cancer workshop/conference or a direct registration via the online BREAST platform. Additionally, radiologists may be referred to complete the BREAST test set by their lead BreastScreen Australia radiologist. The number of readers who read each test set varied from 33 to 206, in accordance with the time release of the test sets between 2014 to 2021. There was a mean value of 130 readers per test set, and each reader had an average of 10 years’ experience in reading mammograms. Out of 537, 471 readers provided their age details, giving a mean age of 46. While the number of female (44%) and male (41%) readers (15% did not specify their gender) and the number of readers who spent less than 4 h (50%) and 4–30 h per week (48%) reading mammograms were similar, 46% of readers read 20–200 cases of mammograms per week, 26% completed a fellowship which lasted for 3–6 months, and 42% currently read for breast screening program (Table 1).
Normal Cases and Reading Conditions
Given the aim of the study is to predict the difficulty of NCs using GRFs, only NCs were included.NCs came from a screening population; hence, patients were all females, aged between 50 to 75, and generally asymptomatic. Truth for each normal case was validated by the consensus reading of at least two expert radiologists, followed by a subsequent negative screen outcome confirmed by two expert radiologists [14]. A total of 361 normal mammography cases from all the nine existing BREAST test sets were used, with each case including bilateral craniocaudal (CC) and mediolateral oblique (MLO) views—a total of four DICOM images per case. The NCs of each test set were viewed by the 537 readers via the BREAST platform, either at a workshop/conference in a simulated reading room (n = 320), or online at their usual clinical setting (n = 217). Reading conditions in both settings were comparable with DICOM images being displayed on a pair of 5-megapixel medical standard monitors and recommended ambient light levels range of 20–40 lux [17, 19, 48, 50, 51]. The mammogram examinations were performed using mammography provided by Hologic (Hologic, Inc., Marlborough, MA, USA), GE (GE Healthcare, Chicago, IL, USA), Siemens (Munich, Germany), Sectra (Sectra, Linköping, Sweden), Philips (Philips Healthcare, Amsterdam, the Netherlands), and Fujifilm (Fujifilm Corporation, Minato City, Tokyo, Japan).
Difficult Normal Cases and Their Breast Density
To identify the difficulty of NCs, difficulty scores (i.e., percentage of incorrect reports per case defined as an objective measure of difficulty [31]) were firstly calculated for each of the 361 NCs using the Royal Australian and New Zealand College of Radiologists (RANZCR) rating (scale of 2 to 5 while 1 represents a normal case [48]) provided by each of the 537 readers. Similar to the American College of Radiology’s numbered BI-RADS mammographic assessment categories, a RANZCR score of 2 was given when readers thought the case was benign lesion, while a score of 3, 4, or 5 was given when readers believed a malignant lesion was present with greater number indicating a higher confidence of malignancy. For cases with more than one rating from the same reader (this happened when a reader believed there were multiple abnormalities), only the highest rating was used. Difficulty scores were derived by dividing the number of readers who gave an incorrect report (i.e., rated a normal case as cancer with a rating of 3, 4, or 5) by the number of total readers who read the test set. The difficulty scores were then categorised into three classes, giving 120 difficult-to-interpret, 122 intermediate-to-interpret, and 119 easy-to-interpret NCs based on one-third of the total 361 cases having the highest, intermediate, and lowest difficulty scores, respectively. However, only cases (total of 239) in the difficult- (n = 120) and easy-to-interpret (n = 119) categories were included for the analysis. The 122 intermediate NCs were excluded to enable a clear distinction between the highest and lowest difficulty NCs. These two categories also resulted in a total of 115 low-BD (58 difficult- and 57 easy-to-interpret) and 124 high-BD NCs (62 difficult- and 62 easy-to-interpret).
Radiomic Analysis
Extraction of Global Radiomic Features
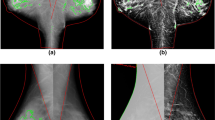
The first step in the radiomics analysis pipeline was to obtain input mammographic images and binary (black and white) masks. In total, 956 mammography DICOM images (4 images per case × 239 NCs: 120 difficult- and 119 easy-to-interpret) were acquired. Using a standard gray level thresholding value of 100 (all pixel intensities above the threshold value were converted to white while the rest were converted to black), binary masks were generated from 956 DICOM images to extract the breast region from the surrounding image background, removing undesirable labels, markers, and other artifacts. Where necessary, a manual adjustment on the thresholding value was applied to ensure the binary masks were created accurately. Next, for computational efficiency purposes, the DICOM images and masks were converted to a TIFF file format, and all the right CC and MLO views of mammograms were flipped to the left so that all mammograms had consistent chest wall on the left side. The TIFF images and binary masks were then cropped based on the maximum size of breast region and used as input images for the radiomic analysis (Fig. 1).
Study workflow. First, images and masks were acquired and used as input images (a). Using the lattice- (ROI) and squared-based (SQ) approaches (b), 34 global radiomic features (GRFs) per image were then extracted from the region of interest (yellow region) and normalised (image gray intensity mean = 0 and standard deviation = 1) (c). Lastly, three machine learning models for predicting difficult-to-interpret mammographic normal cases were built and evaluated (d)
Extraction of GRFs was performed using the handcrafted features approach [38] based on region of interests defined through two methods: (1) multiple regions of interests defined by the lattice-based approach covering the entire breast image (ROI) [52, 53] and (2) largest square rectangular box inscribed within breast (SQ) [54,55,56]. Unlike the previous studies’ techniques [52,53,54,55,56], we included the pectoral muscle for our radiomic analysis as many of our cases had FP annotations near the pectoral muscle area.
A total of 34 GRFs per image (gray level co-occurrence matrix/GLCM-based Haralick texture features (n = 30) [57, 58], first-order statistics/FOS features (n = 2) [33, 58], and neighbourhood gray tone difference matrix/NGTDM texture features (n = 2) [58, 59], Table 2) were extracted using our in-house MATLAB programs. These features were selected as they have been shown to be valuable descriptors of mammographic appearances in measuring the contrast values of spatial inter-relationships between neighbouring pixels of the image (GLCM and NGTDM) and the deviation of single pixel intensity value within the image region of interest (FOS) [33, 57,58,59,60,61,62]. The extracted features were then normalised using a z-score algorithm to calibrate image gray intensity mean to 0 and standard deviation values to 1 [63,64,65]. The ROI features were analysed using MATLAB distinct block processing method with block size 214 × 214 pixels and summarised using the standard deviation approach.
Model Building
Bivariate analysis comparing potential GRFs to the difficult- and easy-to-interpret NCs was conducted.
To predict difficult- from easy-to-interpret NCs, three binary machine learning (ML) classification models were built: (1) CC, (2) MLO, and (3) CC + MLO model. Firstly, in constructing the CC and MLO classification models, the 34 GRFs, obtained from the corresponding right and left CC and MLO view images only, were fed into MATLAB ensemble of 500 decision trees (i.e., random forest) enhanced with LogitBoost ensemble aggregation algorithm (i.e., adaptive logistic regression) [66,67,68]. We selected the random forest technique because a random feature selection is embedded within the model which can reduce the features overfitting problem while producing interpretable models with automatic estimation of feature importance [68]. To recognise useful GRFs for the predictive CC and MLO models, importance scores for each feature were computed through a feature importance analysis using MATLAB’s predictor importance algorithm. Importance scores indicate how useful each feature was in the construction of the decision trees within the model. The higher the value, the more useful the features were in making the prediction for the model. Finally, in building the CC + MLO model, we utilized a late fusion method [69] which consisted of taking the median predictive scores from the constructed CC and MLO models (Fig. 2).
Statistical Analysis and Validation
To assess the performance of the models on our dataset, we needed to measure how well the predictions made by the model matched the observed data. Thus, we trained and validated the CC and MLO models using the leave-one-out-cross-validation approach, a reliable, unbiased, and accurate validation method for evaluating the performance of a machine learning model, assuring no data leakage from the training to the testing phase [62, 70]. This means that each time the model was trained, one case (all images belonging to that case) was left out and used once as a test set to validate the predictive performance of the model, while the remaining cases were used to train the model. This process was repeated for all cases until each case was left out once. The performances of the three models for differentiating difficult- from easy-to-interpret NCs were evaluated by the area under receiver operating characteristic curve (AUC). A DeLong test [71, 72] was used to determine if there was a statistically significant difference in the performance between the three models (CC vs CC + MLO, MLO vs CC + MLO, and CC vs MLO model). A two-sided p value of < 0.05 was used as the criterion of statistically significant difference.
A scree test of exploratory factor analysis [73] was used to determine useful GRFs based on their total importance scores from the CC and MLO models.
Using a Kruskal–Wallis test, we explored if the 34 GRFs differed between difficult- vs easy-to-interpret NCs and if the median difficulty level differed among low- vs high-BD NCs (difficult- and easy-to-interpret). A p value of < 0.05 was considered statistically significant.
All radiomics analysis pipeline and statistical analysis were conducted using MATLAB R2021a (MathWorks, Natick, MA, USA) apart from the DeLong test which was performed using pROC package [74] in RStudio 2021.09.0.351 (RStudio Team, Boston, MA, USA)/R 4.1.2 (R Core Team, Vienna, Austria).
Results
Radiomics Predicting Difficult Normal Cases
Figure 3 demonstrates the comparisons of the overall classification performance. When only GRFs from either CC or MLO views were included, corresponding CC and MLO models achieved similar 0.66 AUCs (95% confidence interval/CI: 0.61–0.71). However, when GRFs from both CC and MLO views were combined using the median predictive scores from both CC and MLO models, the CC + MLO model reached a higher AUC value of 0.71 (95% CI: 0.64–0.78). Nevertheless, the DeLong test shows that there was no statistically significant difference in the performance between the three models (Z = − 1.561, p = 0.119, CC vs CC + MLO; Z = − 1.846, p = 0.065, MLO vs CC + MLO; and Z = 0.198, p = 0.843, CC vs MLO model).
Area under the receiver operating characteristic curve (AUC) for three classifiers. When differentiating difficult- from easy-to-interpret normal cases, using GRFs from CC or MLO view images only, the performance of CC and MLO models achieved similar AUCs of 0.66 (95% CI: 0.61–0.71). However, when fusing GRFs from both CC and MLO images using the median predictive scores from both CC and MLO models, the combined CC + MLO model yielded a higher AUC of 0.71 (95% CI: 0.64–0.78). CC = craniocaudal, CI = confidence interval, MLO = mediolateral oblique
Significant Global Radiomic Features
Among the 34 extracted GRFs, 20 features (Table 2) showed a statistically significant difference between difficult- vs easy-to-interpret NCs (p < 0.05). When the combined 34 features were used for the ML predictive models, six significant features from predictive CC and MLO models demonstrated highest importance scores when compared between the 34 features based on the scree test (Fig. 4). The three most important features from CC model were ROI_Std_Sum_entropy_3, SQ_Difference_variance_9, and SQ_Range_all with mean importance score of 1.25. The three most important features from MLO model were ROI_Std_Correlation_3, ROI_Std_Homogeneity_3, and SQ_Coarseness_9 including 1.39 mean importance score (Table 3).
Significant global radiomic features based on their importance scores derived from the predictive CC and MLO models. a Among the 34 global radiomic features, three features with highest importance scores from CC model were ROI_Std_Sum_entropy_3 (features no. 11) and SQ_Difference_variance_9 (features no. 21) of GLCM, and SQ_Range_all (features no. 33) of FOS, determined based on b the scree plot indicating the steep slope occurred after the third highest features (shown by the red arrow). Similarly, three highest importance score features from MLO model were ROI_Std_Correlation_3 (features no. 1) and ROI_Std_Homogeneity_3 (features no. 7) of GLCM, and SQ_Coarseness_9 (features no. 30) of NGTDM according to c the scree plot with the red arrow showing the steep slope occurred after the third highest features. CC = craniocaudal, FOS = first order statistics, GLCM = gray level co-occurrence matrix, MLO = mediolateral oblique, NGTDM = neighbourhood gray tone difference matrix, ROI = multiple regions of interests defined by the lattice-based approach covering the entire breast image, SQ = largest square rectangular box inscribed within breast, Std = standard deviation
Breast Density of Difficult Normal Cases
Figure 5 illustrates the comparison of the traditional BD-based difficulty level between low- vs high-BD NCs (difficult- and easy-to-interpret). The result from the Kruskal–Wallis test showed no statistically significant difference in the difficulty level between the two groups (H(1) = 0.14, p = 0.709).
Comparison of difficulty level between low-BD (A&B) vs high-BD (C&D) normal cases. No statistically significant difference in the difficulty level was found between the two groups (H(1) = 0.14, p = 0.709). A = Fatty: 0–25% glandular, B = Fibroglandular: 25–50% glandular, BD = breast density, C = Heterogenous: 51–75% glandular, D = Extremely dense: 76–100% glandular
Discussion
This first exploratory study investigated if by employing a radiomic approach to describe the global mammographic features, difficult-to-interpret NCs can be predicted. Our work based on the three ML models (CC, MLO, and CC + MLO) exhibited that GRFs had the ability to accurately differentiate difficult- from easy-to-interpret NCs (AUCs up to 0.71). A few earlier studies indicated the value of local radiomic features from specific tumor/FP regions in predicting radiology trainees’ error making patterns (AUCs up to 0.61) [39, 40]. However, we focused on GRFs’ efficacy in predicting difficult-to-interpret NCs (i.e., an objective measure of difficulty based on readers’ error rate [31]) based on the significance of radiomics in describing mammographic characteristics [38] and global image processing (i.e., global gist/radiological first impression) in diagnostic decision making [41,42,43,44]. Furthermore, our combined CC + MLO model particularly integrating features from both CC and MLO views of mammograms (using the median predictive scores from both CC and MLO models) showed higher performance (0.71 AUC, 0.64–0.78) when compared to CC or MLO model alone (0.66 AUCs, 0.61–0.71) using only features from corresponding CC or MLO images (Fig. 3). This finding suggests that GRFs from both CC and MLO views of mammographic images can provide complementary useful information in predicting difficult-to-interpret NCs. Nonetheless, more and larger future works are required to confirm this finding. While the results of comparing CC to MLO model showed a clearer indication of no statistically significant difference between the model’s performance (Z = 0.198, p = 0.843), the comparisons of CC and MLO to the combined CC + MLO model were indefinite (Z = − 1.561, p = 0.119 and Z = − 1.846, p = 0.065, respectively) which suggests more conclusive answers might need a larger sample.
Previous studies reported that the global gist signal without any localised target information could be used to classify normal from abnormal mammograms [41, 42]. An earlier study [75] highlighted the importance of the global gist signal in guiding a visual search and making a diagnosis decision when interpreting difficult cancer cases (unreported retrospectively identified cancers on mammograms). Thus, we hypothesised that difficult-to-interpret NCs have global image characteristics which may resemble an abnormal appearance and differ from easy-to-interpret images. If a difficult normal image seems to be abnormal at first, this often results in prolonged interpretations, and subsequently, the chance of FP errors may increase since FP errors are often reported at the later stages of prolonged visual searches on normal mammograms [76].
Our work suggests that the GRFs (describing the entire image and largest square rectangular inscribed within breast) contained within mammographic images can help to identify difficult-to-interpret NCs. To date the process of assessing difficulty, NCs tends to depend heavily on a manual and subjective evaluation of radiological experts with high-BD cases often perceived as more difficult [19, 26,27,28, 30]. However, similar to previous studies [28, 29], our finding (Fig. 5) demonstrated no statistically significant difference (H(1) = 0.14, p = 0.709) observed in the difficulty level based on the traditional BD class between low- and high-BD NCs, suggesting that BD classifications may not be suitable for estimating difficult NCs. Conversely, our GRFs can provide important details for recognising mammographic features of difficult NCs that could be used to facilitate an automatic and consistent approach for evaluating difficult NCs, building more predictable test sets and consistent educational materials for radiologists [14,15,16, 21]. It can also facilitate a reliable and effective objective measurement of the recall rate performance of each reader as well as screening program over time given that readers’ performance in test set based education can correspond well with their clinical performance [15, 20, 22,23,24,25].
Additionally, our approach may be used as an augmented clinical artificial intelligence (AI) tool to notify clinical managers about difficult-to-interpret NCs so that more experienced readers can be assigned to the case or an appropriate strategy in pairing readers could be employed to optimise diagnostic accuracy. A single reading strategy, instead of a double reading, can also be exploited for easy-to-interpret NCs, reducing readers’ workload since NCs constitute about 99.5% of cases in screening programs [13, 77,78,79]. Nonetheless, more work is required to further explore and verify such an AI tool’s effectiveness and reliability before incorporating it into a clinical practice.
Our ML approach based on GRFs indicated that the overall mammographic appearance is an important factor in analysing difficult-to-interpret NCs. From the total of 34 features, 20 features showed statistically significant difference between difficult- vs easy-to-interpret NCs (p < 0.05, Table 2). Nevertheless, when the combined 34 features were used for the ML-based predictive models, more redundant features were discovered, resulted in six helpful GRFs in describing difficult-to-interpret NCs based on their importance scores (Fig. 4 and Table 3). The MLO- rather than the CC-derived parameters appeared to be more valuable which may be linked to the MLO-based GRFs involving more breast tissue including the pectoral muscle. This finding, though, needs further investigation by larger future studies.
Some limitations of this study should be acknowledged. The prevalence of NCs in the BREAST test sets used in this study was lower than the real clinical practice. Also, this first proof-of-concept study only examined 34 mammography-based handcrafted GRFs’ ability in predicting difficult-to-interpret NCs based on the average difficulty scores of all readers. Moreover, determining important GRFs based on the scree test [73] (Fig. 4b, c) was less clear cut for the CC model when compared to the MLO. Future works should explore other potential useful CC-based GRFs (e.g., feature no. 30, 14, 16, and 27) and type of radiomic features (e.g., local handcrafted or deep learning features) and using equipment from other imaging vendors (e.g., Canon, Canon Medical Systems Corporation, Otawara, Tochigi, Japan). Finally, separate models for predicting difficult cases of readers at various experience levels should be investigated.
Conclusions
Our findings suggested that quantitative GRFs extracted from mammograms are helpful for accurately distinguishing difficult- from easy-to-interpret NCs, denoting the importance of global gist in making a diagnostic decision. Six important features from the classification models were recognised and highlighted. Our findings could be useful for radiology educations and mammography screening programs in creating more predictable test sets, and reliable and effective objective measurement of the recall rate performance of each reader and screening program over time, as well as optimising single and double reading practice.
Data Availability
The data that support the findings of this study are available from BREAST but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available.Data are however available from the authors upon reasonable request and with permission of BREAST.
Abbreviations
- AI:
-
Artificial intelligence
- AUC:
-
Area under receiver operating characteristic curve
- BD:
-
Breast density
- BI-RADS:
-
Breast Imaging and Reporting Data System
- BREAST:
-
BreastScreen Reader Assessment Strategy
- CC:
-
Craniocaudal
- CI:
-
Confidence interval
- DICOM:
-
Digital Imaging and Communications in Medicine
- FOS:
-
First order statistics
- FP:
-
False positive
- GLCM:
-
Gray level co-occurrence matrix
- GRFs:
-
Global radiomic features
- ML:
-
Machine Learning
- MLO:
-
Mediolateral oblique
- NCs:
-
Normal cases
- NGTDM:
-
Neighbourhood gray tone difference matrix
- RANZCR:
-
Royal Australian and New Zealand College of Radiologists
- ROI:
-
Multiple regions of interests defined by the lattice-based approach covering the entire breast image
- SQ:
-
Largest square rectangular box inscribed within breast
References
Sung H, Ferlay J, Siegel RL et al (2021) Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA: A Cancer Journal for Clinicians 71:209–249
Seely JM, Alhassan T (2018) Screening for breast cancer in 2018-what should we be doing today? Current oncology (Toronto, Ont) 25:S115-S124
Njor S, Nystrom L, Moss S et al (2012) Breast cancer mortality in mammographic screening in Europe: a review of incidence-based mortality studies. 19:33-41
Welfare AIoHa (2020) BreastScreen Australia monitoring report 2020. AIHW, Canberra
Castells X, Torá-Rocamora I, Posso M et al (2016) Risk of Breast Cancer in Women with False-Positive Results according to Mammographic Features. Radiology 280:379-386
Román M, Castells X, Hofvind S, Euler‐Chelpin M (2016) Risk of breast cancer after false‐positive results in mammographic screening. Cancer Medicine 5:1298-1306
Ekpo EU, Alakhras M, Brennan P (2018) Errors in Mammography Cannot be Solved Through Technology Alone. Asian Pacific journal of cancer prevention : APJCP 19:291-301
Skaane P (2019) How Can We Reduce Unnecessary Procedures after Screening Mammography? Radiology 291:318-319
Ong M-S, Mandl KD (2015) National Expenditure For False-Positive Mammograms And Breast Cancer Overdiagnoses Estimated At $4 Billion A Year. Health Affairs 34:576-583
Vlahiotis A, Griffin B, Stavros Md FAT, Margolis J (2018) Analysis of utilization patterns and associated costs of the breast imaging and diagnostic procedures after screening mammography. ClinicoEconomics and Outcomes Research Volume 10:157-167
Alcusky M, Philpotts L, Bonafede M, Clarke J, Skoufalos A (2014) The Patient Burden of Screening Mammography Recall. Journal of Women's Health 23:S-11-S-19
Lee W, Peters G (2013) Mammographic screening for breast cancer: A review. Journal of Medical Radiation Sciences 60:35-39
Peintinger F (2019) National Breast Screening Programs across Europe. Breast Care 14:354-358
Brennan PC, Trieu PD, Tapia K, Ryan J, Mello-Thoms C, Lee W (2014) BREAST: A Novel Strategy to Improve the Detection of Breast Cancer. In: Fujita H, Hara T, Muramatsu C, (eds) International Workshop on Digital Mammography 2014. Springer International Publishing Switzerland, Cham, pp 438-443
Gale A, Chen Y (2020) A review of the PERFORMS scheme in breast screening. The British journal of radiology 93:20190908
Suleiman ME, Rickard M, Brennan PC (2020) Perfecting detection through education. Radiography 26:S49-S53
Trieu PD, Tapia K, Frazer H, Lee W, Brennan P (2019) Improvement of Cancer Detection on Mammograms via BREAST Test Sets. Academic Radiology 26:e341–e347
Trieu PD, Lewis SJ, Li T et al (2021) Improving radiologist's ability in identifying particular abnormal lesions on mammograms through training test set with immediate feedback. Scientific reports 11:1-8
Suleiman WI, Rawashdeh MA, Lewis SJ et al (2016) Impact of Breast Reader Assessment Strategy on mammographic radiologists' test reading performance. Journal of Medical Imaging and Radiation Oncology 60:352–358
Chen Y, James JJ, Cornford EJ, Jenkins J (2020) The Relationship between Mammography Readers’ Real-Life Performance and Performance in a Test Set–based Assessment Scheme in a National Breast Screening Program. Radiology: Imaging Cancer 2:e200016
Chen Y, Gale A (2018) Performance Assessment Using Standardized Data Sets: The PERFORMS Scheme in Breast Screening and Other Domains. In: Samei E, Krupinski EA, (eds) The Handbook of Medical Image Perception and Techniques. Cambridge University Press, Cambridge, pp 328-342
Brancato B, Peruzzi F, Saieva C et al (2022) Mammography self-evaluation online test for screening readers: an Italian Society of Medical Radiology (SIRM) initiative. Eur Radiol 32:1624-1633
Soh BP, Lee W, McEntee MF et al (2013) Screening Mammography: Test Set Data Can Reasonably Describe Actual Clinical Reporting. Radiology 268:46-53
Soh BP, Lee WB, Mello‐Thoms C et al (2015) Certain performance values arising from mammographic test set readings correlate well with clinical audit. Journal of Medical Imaging and Radiation Oncology 59:403-410
Qenam BA, Li T, Brennan PC (2021) Test set participation may impact positively on clinical mammographic performanceSPIE 11599, Medical Imaging 2021: Image Perception, Observer Performance, and Technology Assessment. SPIE, Online, pp 1–8
Dong L, Chen Y, Gale A (2013) Breast screening: understanding case difficulty and the nature of errorsSPIE 8673, Medical Imaging 2013: Image Perception, Observer Performance, and Technology Assessment. SPIE, Lake Buena Vista (Orlando Area), Florida, United States, pp 1-8
Scott HJ, Gale AG (2005) Breast screening technologists: When is a difficult case truly difficult and for whom?SPIE 5749, Medical Imaging 2005: Image Perception, Observer Performance, and Technology Assessment. SPIE, Bellingham, WA, pp 557-565
Wong D, Gandomkar Z, Reed W, Lewis SJ, Brennan P (2022) The effect on reader performance of case image characteristics including expert determined difficulty ratings using a mammography test set. Under review
Grimm LJ, Kuzmiak CM, Ghate SV, Yoon SC, Mazurowski MA (2014) Radiology Resident Mammography Training. Academic Radiology 21:888-892
Ang ZZ, Rawashdeh MA, Heard R, Brennan PC, Lee W, Lewis SJ (2017) Classification of normal screening mammograms is strongly influenced by perceived mammographic breast density. Journal of Medical Imaging and Radiation Oncology 61:461-469
Mazurowski MA (2013) Difficulty of mammographic cases in the context of resident training: preliminary experimental dataSPIE 8673, Medical Imaging 2013: Image Perception, Observer Performance, and Technology Assessment SPIE, Lake Buena Vista (Orlando Area), Florida, United States, pp 1–6
Kabbai L, Abdellaoui M, Douik A (2019) Image classification by combining local and global features. The Visual Computer 35:679-693
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: Images Are More than Pictures, They Are Data. Radiology 278:563-577
Gillies RJ, Schabath MB (2020) Radiomics Improves Cancer Screening and Early Detection. Cancer Epidemiology Biomarkers & Prevention 29:2556-2567
Lee S-H, Park H, Ko ES (2020) Radiomics in Breast Imaging from Techniques to Clinical Applications: A Review. Korean Journal of Radiology 21:779-792
Tagliafico AS, Piana M, Schenone D, Lai R, Massone AM, Houssami N (2020) Overview of radiomics in breast cancer diagnosis and prognostication. The Breast 49:74-80
Conti A, Duggento A, Indovina I, Guerrisi M, Toschi N (2020) Radiomics in breast cancer classification and prediction. Seminars in Cancer Biology 72:238-250
Siviengphanom S, Gandomkar Z, Lewis SJ, Brennan PC (2021) Mammography-based Radiomics in Breast Cancer: A Scoping Review of Current Knowledge and Future Needs. Academic Radiology 29:1228-1247
Zhang J, Lo JY, Kuzmiak CM, Ghate SV, Yoon SC, Mazurowski MA (2014) Using computer-extracted image features for modeling of error-making patterns in detection of mammographic masses among radiology residents. Medical Physics 41:091907
Zhang J, Silber JI, Mazurowski MA (2015) Modeling false positive error making patterns in radiology trainees for improved mammography education. Journal of Biomedical Informatics 54:50-57
Evans KK, Georgian-Smith D, Tambouret R, Birdwell RL, Wolfe JM (2013) The gist of the abnormal: Above-chance medical decision making in the blink of an eye. Psychonomic Bulletin & Review 20:1170-1175
Gandomkar Z, Ekpo E, Lewis S et al (2018) Detection of the abnormal gist in the prior mammograms even with no overt sign of breast cancerSPIE 10718, 14th International Workshop on Breast Imaging (IWBI 2018). SPIE, Atlanta, Georgia, United States, pp 1-6
Brennan PC, Gandomkar Z, Ekpo EU et al (2018) Radiologists can detect the 'gist' of breast cancer before any overt signs of cancer appear. Scientific reports 8:8717-8717
Gandomkar Z, Siviengphanom S, Ekpo EU et al (2021) Global processing provides malignancy evidence complementary to the information captured by humans or machines following detailed mammogram inspection. Scientific reports 11:1-12
Gandomkar Z, Mello-Thoms C (2019) Visual search in breast imaging. The British journal of radiology 92:20190057
Lehman CD, White E, Peacock S, Drucker MJ, Urban N (1999) Effect of age and breast density on screening mammograms with false-positive findings. American Journal of Roentgenology 173:1651-1655
Holland K, Van Gils CH, Mann RM, Karssemeijer N (2017) Quantification of masking risk in screening mammography with volumetric breast density maps. Breast Cancer Research and Treatment 162:541-548
Brennan P, Lee W, Tapia K (2018) Breast Screen Reader Assessment Strategy (BREAST): A Research Infrastructure with a Translational Objective. In: Samei E, Krupinski EA, (eds) The Handbook of Medical Image Perception and Techniques. Cambridge University Press, Cambridge, pp 343–356
Brennan PC, Tapia K, Ryan J, Lee W (2013) BREAST: a novel method to improve the diagnostic efficacy of mammographySPIE 8673, Medical Imaging 2013: Image Perception, Observer Performance, and Technology Assessment. SPIE, Lake Buena Vista (Orlando Area), Florida, United States, pp 1–5
Tapia KA, Rickard MT, McEntee MF, Garvey G, Lydiard L, C Brennan P (2020) Impact of breast density on cancer detection: observations from digital mammography test sets. International Journal of Radiology & Radiation Therapy 7:36–41
McEntee M, Brennan P, Evanoff M, Phillps P, O Connor W, Manning D (2006) Optimum ambient lighting conditions for the viewing of softcopy radiological images. SPIE
Zheng Y, Keller BM, Ray S et al (2015) Parenchymal texture analysis in digital mammography: A fully automated pipeline for breast cancer risk assessment. Medical Physics 42:4149-4160
Zheng Y, Wang Y, Keller B, Conant E, Gee J, Kontos D (2013) A fully-automated software pipeline for integrating breast density and parenchymal texture analysis for digital mammograms: parameter optimization in a case-control breast cancer risk assessment studySPIE 8670, Medical Imaging 2013: Computer-Aided Diagnosis SPIE Lake Buena Vista (Orlando Area), Florida, United States 1 7
Wei J, Chan H-P, Wu Y-T et al (2011) Association of Computerized Mammographic Parenchymal Pattern Measure with Breast Cancer Risk: A Pilot Case-Control Study. Radiology 260:42-49
Wu Y-T, Sahiner B, Chan H-P et al (2008) Comparison of mammographic parenchymal patterns of normalsubjects and breast cancer patientsSPIE 6915, Medical Imaging 2008: Computer-Aided Diagnosis. SPIE, San Diego, California, United States, pp 1-8
Gandomkar Z, Suleiman M, Demchig D, Brennan P, McEntee M (2019) BI-RADS density categorization using deep neural networksSPIE 10952, Medical Imaging 2019: Image Perception, Observer Performance, and Technology Assessment. SPIE, pp 1–7
Haralick RM, Shanmugam K, Dinstein IH (1973) Textural Features for Image Classification. IEEE Transactions on Systems, Man, and Cybernetics SMC-3:610–621
Costaridou L (2005) Medical image analysis methods. Taylor & Francis, Boca Raton
Amadasun M, King R (1989) Textural features corresponding to textural properties. IEEE Transactions on Systems, Man, and Cybernetics 19:1264-1274
Cui WJ, Wang C, Jia L et al (2019) Differentiation Between G1 and G2/G3 Phyllodes Tumors of Breast Using Mammography and Mammographic Texture Analysis. Frontiers in Oncology 9:1-9
Jiang X, Zou X, Sun J, Zheng A, Su C (2020) A Nomogram Based on Radiomics with Mammography Texture Analysis for the Prognostic Prediction in Patients with Triple-Negative Breast Cancer. Contrast Media & Molecular Imaging 2020:1-10
La Forgia D, Fanizzi A, Campobasso F et al (2020) Radiomic Analysis in Contrast-Enhanced Spectral Mammography for Predicting Breast Cancer Histological Outcome. Diagnostics 10:1-11
Castaldo R, Pane K, Nicolai E, Salvatore M, Franzese M (2020) The Impact of Normalization Approaches to Automatically Detect Radiogenomic Phenotypes Characterizing Breast Cancer Receptors Status. Cancers 12:1-25
Kotsiantis SB, Kanellopoulos D, Pintelas PE (2007) Data Preprocessing for Supervised Leaning. International Journal of Computer and Information Engineering 1:4091-4096
Haga A, Takahashi W, Aoki S et al (2019) Standardization of imaging features for radiomics analysis. The Journal of Medical Investigation 66:35-37
Friedman J, Hastie T, Tibshirani R (2000) Additive logistic regression: a statistical view of boosting (With discussion and a rejoinder by the authors). The Annals of Statistics 28:337-407
Breiman L (2001) Random Forests. Machine Learning 45:5-32
Couronné R, Probst P, Boulesteix A-L (2018) Random forest versus logistic regression: a large-scale benchmark experiment. BMC Bioinformatics 19:1-14
Lee SL, Zare MR, Muller H (2019) Late fusion of deep learning and handcrafted visual features for biomedical image modality classification. IET Image Processing 13:382-391
Molinaro AM, Simon R, Pfeiffer RM (2005) Prediction error estimation: a comparison of resampling methods. Bioinformatics 21:3301-3307
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the Areas under Two or More Correlated Receiver Operating Characteristic Curves: A Nonparametric Approach. Biometrics 44:837-845
Sun X, Xu W (2014) Fast Implementation of DeLong’s Algorithm for Comparing the Areas Under Correlated Receiver Operating Characteristic Curves. IEEE Signal Processing Letters 21:1389-1393
Ledesma RD, Valero-Mora P, Macbeth G (2015) The Scree Test and the Number of Factors: a Dynamic Graphics Approach. The Spanish Journal of Psychology 18
Robin X, Turck N, Hainard A et al (2011) pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12:77
Nodine CF, Mello-Thoms C, Weinstein SP et al (2001) Blinded Review of Retrospectively Visible Unreported Breast Cancers: An Eye-Position Analysis. Radiology 221:122-129
Mello-Thoms C (2006) How Does the Perception of a Lesion Influence Visual Search Strategy in Mammogram Reading? Academic Radiology 13:275-288
Taylor-Phillips S, Stinton C (2020) Double reading in breast cancer screening: considerations for policy-making. The British journal of radiology 93:20190610
Brennan PC, Ganesan A, Eckstein MP et al (2019) Benefits of Independent Double Reading in Digital Mammography. Academic Radiology 26:717-723
Gandomkar Z, Tay K, Brennan PC, Kozuch E, Mello-Thoms C (2018) Can eye-tracking metrics be used to better pair radiologists in a mammogram reading task? Medical Physics 45:4844-4856
Acknowledgements
The authors thank the BreastScreen Reader Assessment Strategy (BREAST) program at the University of Sydney for access and use of their data for the study. BREAST is funded/supported by the Commonwealth Government Department of Health, with their images kindly provided by BreastScreen Australia, the National Breast Cancer Foundation, and the Cancer Institute New South Wales. We would like to also acknowledge the funding support for this research project provided by the National Breast Cancer Foundation [grant ID: IIRS-22-087] and Cancer Institute New South Wales [grant ID: 2022/ECF1426]. The authors wish to thank Dr Robert Heard for his kind statistical advice as well as the reviewers for their rigorous evaluation of this manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. This work is supported by the National Breast Cancer Foundation [grant ID: IIRS-22-087] and Cancer Institute New South Wales [grant ID: 2022/ECF1426].
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Siviengphanom, S., Gandomkar, Z., Lewis, S.J. et al. Global Radiomic Features from Mammography for Predicting Difficult-To-Interpret Normal Cases. J Digit Imaging 36, 1541–1552 (2023). https://doi.org/10.1007/s10278-023-00836-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-023-00836-7