Abstract
Generative adversarial networks (GAN) in medicine are valuable techniques for augmenting unbalanced rare data, anomaly detection, and avoiding patient privacy issues. However, there were limits to generating high-quality endoscopic images with various characteristics, such as peristalsis, viewpoints, light sources, and mucous patterns. This study used the progressive growing of GAN (PGGAN) within the normal distribution dataset to confirm the ability to generate high-quality gastrointestinal images and investigated what barriers PGGAN has to generate endoscopic images. We trained the PGGAN with 107,060 gastroscopy images from 4165 normal patients to generate highly realistic 5122 pixel-sized images. For the evaluation, visual Turing tests were conducted on 100 real and 100 synthetic images to distinguish the authenticity of images by 19 endoscopists. The endoscopists were divided into three groups based on their years of clinical experience for subgroup analysis. The overall accuracy, sensitivity, and specificity of the 19 endoscopist groups were 61.3%, 70.3%, and 52.4%, respectively. The mean accuracy of the three endoscopist groups was 62.4 [Group I], 59.8 [Group II], and 59.1% [Group III], which was not considered a significant difference. There were no statistically significant differences in the location of the stomach. However, the real images with the anatomical landmark pylorus had higher detection sensitivity. The images generated by PGGAN showed highly realistic depictions that were difficult to distinguish, regardless of their expertise as endoscopists. However, it was necessary to establish GANs that could better represent the rugal folds and mucous membrane texture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastrointestinal (GI) endoscopy is an essential tool for diagnosing and treating various GI diseases, including cancer which remains a global health burden and one of the important causes of cancer-related deaths worldwide [1, 2]. However, given the essential nature of this tool, it is still challenging to diagnose GI neoplasms by endoscopy [3, 4] because the detection and diagnosis depend on the endoscopist’s experience [5].
Several studies have recently reported the performance of deep learning-based artificial intelligence (AI) systems using endoscopic images to compensate for endoscopist experience. These reports have shown favorable performance for detecting GI neoplasms, predicting invasion depth, or classifying neoplasms [6,7,8,9,10,11,12]. The computer-aided detection and diagnosis (CAD) system for colonic neoplasms has well-established and proven clinical efficacy [13,14,15]. However, the CAD system for gastric neoplasms has demonstrated relatively poor performance compared to colonic neoplasms [16]. The reason is that the background mucosa of gastric neoplasms accompanies chronic inflammation, such as in atrophic gastritis and intestinal metaplasia [11, 16]. Given the sensitivity of deep learning to background noise, these studies have significant limitations. They needed a large dataset of real-world and high-quality images. Such images have been associated with high costs, increased time, and ethical issues such as privacy concerns [17].
In research to overcome the vulnerability of deep learning to background noise or adversarial samples, Goodfellow et al. [18] introduced generative adversarial networks (GANs), a neural network to generate the realistic distribution of the training dataset. GAN is a model suitable for medical imaging research that can collect a large number of normal data through health checkups. In particular, the possibility of detecting abnormalities using GAN has been dramatically expanded, and many studies on abnormality detection using GAN have been reported in the medical field [19,20,21,22,23,24]. However, high-resolution and highly represented endoscopic images must be generated to obtain high-sensitivity screening performance. Many previous studies employing GANs in endoscopic images have focused on their application to low-quality images due to the limited performance of the image-generating models. However, we have focused our research on the high image-generating ability of the endoscope to build a more sensitive screening of CAD from a disease detection perspective.
Progressive growing of the GAN (PGGAN) [25] is one of the GAN that can create high-definition and high-quality images. This network has been applied in image processing and the medical field rather than matching endoscopic camera resolution. Park et al. reported that PGGAN could synthesize high-resolution body CT images, proven using the visual Turing test [26]. The result of the Turing test found a lack of accuracy in the thoracic abdominal junction and anatomical details. We speculated that PGGAN would have problems generating specific structures in the same way on endoscopic imaging.
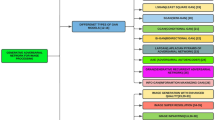
As described above, in the field of endoscopy, various studies using GAN, such as lesion synthesis and abnormality detection techniques, have been attempted. However, most of the research has been done with low-quality images. A GAN that produces low-quality images does not very well reflect small polyps or mucosal patterns. For this reason, we tried to analyze why it is challenging to generate an endoscopic image of high quality. Therefore, we employed a PGGAN to generate highly realistic-looking gastroscopy images and performed a visual Turing test as the first step towards a GANs-based AI system endoscopy. The overall process of our visual Turing test of the gastroscopic images was summarized in Fig. 1.
Process of gastric endoscopy image Turing test, A randomly generated 100 gastroscopy images using Progressive growing of GAN (PGGAN), B randomly extracted 100 images from health checkup data, C Turing test on 19 endoscopists to classify 100 normal images and 100 synthetic images, D Analyzing the diagnostic results of the Turing test
Methods
Data Acquisition
Electronic medical records were retrospectively reviewed to find patients who underwent gastroscopy in the Health Screening and Promotion Center of Asan Medical Center. A total of 166,997 images from 4,165 normal patients were obtained. However, various artifacts were existed in endoscopic imaging, such as motion-blur and light reflections. Also, both the endoscopic images of the gastric cardia and the endoscope itself in the retroflexion view were excluded, as the endoscope's distortion in the generated image is frequently identified as an abnormal structure. Since such artifacts may interfere with the learning representation of gastric endoscopy, 1000 images containing artifacts were selected manually and trained using supervised learning techniques to differentiate them. Data cleansing will be described in more detail in the next Data cleansing session. In this manner, 59,937 images were cleansed as there were motion-blurred or color-blurred images in the captured video. Finally, 107,060 images were used for training. Figure S1 demonstrated a schematic diagram of the process of collecting the training dataset by extracting an endoscopic image; the images were upsampled from 3502 pixels to 5122 pixel-sized images using OpenCV’s resize transform with third-order spline interpolation. This is since a \({2}^{n}\) sized image is required to train the PGGAN generator.
PGGAN was used to generate high-resolution gastroscopic images up to 5122 pixels, generated by applying the progressive growth technique to the generator. We used an official implementation code of PGGAN in this study. Since the training of PGGAN proceeds as the size of the input image of n power 2 gradually increases, the training image size gradually increased from 42 to 5122 pixels, and the batch size decreased from 512 to 16, respectively, as the computational load increased with increasing resolution.
The learning rate of PGGAN was set to 1e-3 while training. Figure 2 shows the process of PGGAN’s step-by-step generation of 100 images for the Turing test. The PGGAN training assessment was based on Fréchet inception distance(FID) [27] scores and expert visual inspection in every 4000 images on the PGGAN. During training PGGAN, the FID score was converged at an average of 6.0 after 6 million images were shown. Finally, the training weight was selected according to the visualized verification result of one expert who was excluded from the Turing test. The training took 5 days with two NVIDIA Titan RTX GPUs.
Data Cleansing
We discovered that the acquired normal endoscopic images contained various artifacts. Assuming that low-quality original images would result in low-quality generated images, we aimed to exclude artifact-contained images. Since it would require a significant effort to examine the quality of a total of 166,997 images, we planned to construct a small image quality dataset to perform data cleansing. We annotated 1000 high-quality and low-quality endoscopic images each from our real dataset. These images were randomly split into training, validation, and testing sets in a 6:2:2 ratio. We used a ResNet18 [28] model pre-trained on ImageNet to train a binary classification model, which achieved an accuracy of 95% in the testing dataset. We applied this model to the remaining 164,997 images to cleanse the dataset, a total of 59,937 images were cleansed using ResNet18.
Image Turing Test on the Realism of Synthetic Endoscopy Images
The validation set consisted of 200 gastroscopy images (100 synthetic images and 100 real images). All synthetic images in the validation set were automatically generated by our modified PGGAN model and were not individually selected by the researchers to avoid selection bias. The real images were taken at the Asan Medical Center, which did not otherwise participate in the realism assessment study. A website was created to upload the validation set, with the posting and displaying of the 200 images being performed randomly as Figure S2. Nineteen endoscopists (twelve endoscopists with < 5 years of experience [Group I]; three endoscopists with 5–10 years of experience [Group II]; and four endoscopists with > 10 years of experience [Group III]) independently evaluated each of the 200 images, and visually decide on whether each image was real or fake with no time limit and no prior information on the number of actual or fake images. To investigate the features of obviously artificial images, we defined artificial images as those synthetic images that the majority of readers (more than ten readers) decided were fake.
Statistical Analysis
The mean accuracy, sensitivity, and specificity of the 19 readers were calculated. Sensitivity is the probability of detecting real images among the real images, and specificity is the proportion of identification of generated images. General estimation equations were used to compare the accuracy, sensitivity, and specificity across the reader groups with different experience levels (Group I, Group II, and Group III), and across the anatomical subgroups. Since there were only 19 participants in our Turing test, a nonparametric Kruskal–Wallis test [29] was performed to analyze whether the difference between the three groups of the independent variable level was significant. The significance level was corrected for multiple comparisons using the Bonferroni-holm correction [30], and inter-reader agreement was evaluated using the intraclass correlation coefficient [31]. The R version 3.6.3 was used for statistical analysis, with a significance level of P < .05.
Results
Overall Assessment of the Visual Turing Test
The mean accuracy, sensitivity, and specificity of the 19 endoscopists were 61.3%, 70.3%, and 52.4%, respectively. Table 1 and Fig. 3 summarize the results of the realistic assessment of all images by 19 endoscopists with different experience levels (Group I: 62.5, Group II: 59.8, and Group III: 59.1; P > .05). There was no correlation between endoscopist groups since the standard deviation of mean sensitivity and mean specificity was tremendous. The average sensitivity tended to be about 17.9% higher than the average specificity.
Subgroup Analysis According to the Location
To analyze whether PGGAN generates images for each region of the stomach and duodenum, subgroup analysis by location was conducted. Figure 4 shows sample endoscopic images generated by PGGAN for each part of the stomach and duodenum. The number of images for each part of the 100 randomly selected real images and generated images were shown in Fig. 5. Although the distribution of the generated image is not precisely the same, it was confident that the generated image with a trend similar to the actual distribution. Table 2 shows the analysis results for each anatomical location. As there was no difference in distinguishing the actual gastroscopy image and the fake image according to experience, we analyzed only the differences for each region. There was no significant difference in diagnostic performance according to the anatomical location and endoscopic view. However, there was a statistically significant difference in sensitivity according to anatomical landmarks; the sensitivity to the pylorus was higher (P = .002).
Sample images of synthesized high-resolution gastroscopy. A Cardia and fundus in retroflexion view. B Lesser curvature and anterior wall of a high body in retroflexion view. C High body and mid body in retroflexion view. D Mid body and low body in forward view. E Angle and antrum in retroflexion view. F Antrum and pylorus in forward view. G Pylorus in forward view. H 2nd portion of duodenum in forward view)
Inter-Observer Agreement in the Synthetic and Real Images
Intraclass correlation coefficients were used to analyze the agreement between the three groups to identify generated images. The interobserver agreement for all endoscopists was observed to be 0.74 for the entire image set, as shown in Table 3. However, when subgroup analysis was performed according to clinical endoscopic experience, Group II and Group III showed poor average Intraclass correlation coefficients below 0.3.
Analysis of the Features of Obviously Artificial Images
One hundred of the randomly chosen synthetic images were included in the image Turing test. The mean number of endoscopists who correctly answered fake for each synthetic image was 9.95 ± 3.08 (CI 95%, 9.34–10.56). Ten of the synthetic images were detected as fake by more than 10 out of 19 endoscopists. One experienced endoscopist reported the key features used to detect the images as fake for these synthetic images. The key features of 10 synthetic images were described in Fig. 6; overall, they showed uneven structure and an absence of typical structure.
Synthetic images that the majority of readers decided were fake. A Irregular transverse folds in the lesser curvature. B Absence of normal gastric folds on the greater curvature of the lower body and scattered hyperemic mucosa and asymmetry between anterior wall and posterior wall of the angle. C Abrupt discontinuation and fusion of gastric folds in the greater curvature. D Absence of normal gastric folds on the greater curvature of the low body and asymmetry between anterior wall and posterior wall of the angle. E Abrupt discontinuation of gastric folds in the greater curvature. F Absence of an endoscope in the esophagogastric junction. G Discontinuation of circular folds on the duodenum. H Abrupt discontinuation and fusion of gastric folds in the greater curvature. I Absence of gastric folds and fundus. J Absence of gastric folds and fundus)
Discussion
In this study, we used the PGGAN model to generate realistic gastroscopy images and evaluated the realism of the images generated by 19 endoscopy physicians with the image Turing test. In the Turing test results, 19 endoscopists generally identified fake images with sensitivity and less specificity. Accuracy does not make a meaningful distinction between real and fake images. Also, in a subgroup analysis of endoscopic experience, there was no significant difference in the discrimination between real and synthetic endoscopic images.
Recently, many health experts are conducting regular checkups, and developments in endoscopic data storage have made it easy to access common imaging datasets to many researchers and clinicians. Normal and abnormal endoscopic images can, therefore, be taken frequently; however, normal images are more effective and useful for collecting huge datasets than abnormal images since they are easier to curate and provide more data. To date, supervised deep learning-based models have been successfully applied in the detection and classification of specific diseases in various medical fields [32]. However, deep learning techniques may be challenged in endoscopy due to the large range of normal variations and non-neoplastic conditions such as inflammation. Neoplastic lesions can be disregarded or unnoticed during the screening endoscopy, leading to a missed diagnosis of both the index lesion and any simultaneous lesions [3, 4, 33]. Therefore, meticulous inspection through the deep learning-based AI system for gastroscopy may improve diagnostic performance.
We believe that creating these highly realistic images may be the first step in applying GAN to the development of models applicable in the medical fields described above. This is the first study that systematically evaluated the realism of GAN-generated synthetic gastroscopy images through the intuition of numerous endoscopists. Therefore, the realistic nature of the synthetic images used in the previous studies was not confirmed. In addition, most previous studies focused on improving deep learning-based task performance resulting from the generation of images with diseases, using synthetic abnormal images to adjust for data imbalances; they did not consider the generation of normal images.
Recently, several studies have shown the favorable performance of deep learning-based AI systems in endoscopy and there were promising results in colonoscopy. However, AI systems for the detection of gastric neoplasms have exhibited poor performance compared to that of colonic neoplasms. In a previous study, a real-time AI system using white light endoscopic images showed high sensitivity (94.2%) similar to expert endoscopists in detecting upper GI cancer; however, the positive predictive value of the AI system (81.4%) was relatively lower than that of experts (93.2%) [8]. Hirasawa et al. also reported a low positive predictive value (30.6%) in deep learning-based AI systems for gastric cancer [16]. The causes of false positives by AI systems in the study included benign lesions such as gastritis, atrophy, or intestinal metaplasia and normal structures such as the gastric angle, pylorus, amounts of mucus, and elevation of gastric wall during peristalsis. Our PGGAN could create realistic gastric mucosa and a specific typical structure, including the gastric angle, pylorus, and the endoscope itself. The detection of gastric neoplasms using GANs will help to improve performance by reducing these errors. A previous deep learning based-AI study attempted a binary classification of gastric ulcers and gastric cancer but showed lower accuracy (77.1%) [34]. This performance is possible because they used a relatively smaller number of cases (220 benign ulcer images and 367 cancer images) than other deep learning-based studies. The need for a large dataset of real-world and high-quality images is one of the main limitations of deep learning. The multi-institutional data sharing and research agreements through efforts to address ethical issues in medical AI research are important for obtaining a large dataset of real-world and high-quality images. Also, the development of GAN can increase data quantity and quality with relatively few ethical issues and low costs. Our pre-trained GAN for healthy gastroscopy images will be helpful to investigate related research.
Our study has some limitations. First, the visual Turing test was conducted using 5122 resolution images. Recently, since clinical endoscopists deal with full high-definition endoscopic images, it is necessary to evaluate them in higher-quality images. The future work will increase GAN output resolution further to match technical advances. Secondly, the sensitivity according to anatomical landmarks was statistically significant in the subgroup analysis, specifically for the pylorus. We did not give endoscopists any prior information on the number of actual or fake images in the image Turing test for a precise validation test. Nineteen endoscopists tended to indicate the test image as real, as it was difficult to distinguish real or fake images. As a result, thirteen out of nineteen endoscopists had higher sensitivity than specificity. The specificity according to anatomical landmarks showed a low tendency (45.9–55.1%). Third, our evaluation was not performed with various modern generative networks other than PGGAN. We chose PGGAN as it was the most recent state-of-the-art at the time of our Turing Test analysis. We should determine if we can overcome the lack of generative performance by employing cutting-edge networks that are superior to PGGAN at generating high-quality images. Finally, to test PGGAN's capacity to generate realistic images, real endoscopic images with color jittering and motion blur were removed. Since many low-quality images were included in the actual endoscopic image, data bias may have been introduced while removing them.
In conclusion, our PGGAN model generated highly realistic endoscopic images of the stomach and duodenum, difficult to distinguish regardless of endoscopist experience. Since this PGGAN model can produce high-quality endoscopic images, it may help mitigate the limitations of deep learning, including class imbalances, challenges of data-sharing, and privacy concerns. In addition, if the expression of folds and texture, which the PGGAN lacks, is improved, it may be sufficient to detect abnormal lesions of the upper GI.
Data Availability
The datasets are not publicly available due to restrictions in the data-sharing agreements with the data sources. Ethics approval for using these de-identified slides in this study was granted by the Institutional Review Board of Asan Medical Center.
References
DeWitt J, Van Dam J: Development of Endoscopy- Gastroenterology Diamond Jubilee Review. Gastroenterology 155:237-240, 2018
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A: Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries 68:394–424, 2018
Lee HL, et al.: When do we miss synchronous gastric neoplasms with endoscopy? Gastrointestinal endoscopy 71:1159-1165, 2010
Ahn SB, Han DS, Bae JH, Byun TJ, Kim JP, Eun CS: The Miss Rate for Colorectal Adenoma Determined by Quality-Adjusted, Back-to-Back Colonoscopies. Gut and liver 6:64-70, 2012
Yamazato T, et al.: Two years' intensive training in endoscopic diagnosis facilitates detection of early gastric cancer. Internal medicine (Tokyo, Japan) 51:1461-1465, 2012
Cho BJ, et al.: Automated classification of gastric neoplasms in endoscopic images using a convolutional neural network. Endoscopy 51:1121-1129, 2019
Li L, et al.: Convolutional neural network for the diagnosis of early gastric cancer based on magnifying narrow band imaging. Gastric cancer : official journal of the International Gastric Cancer Association and the Japanese Gastric Cancer Association 23:126-132, 2020
Luo H, et al.: Real-time artificial intelligence for detection of upper gastrointestinal cancer by endoscopy: a multicentre, case-control, diagnostic study. The Lancet Oncology 20:1645-1654, 2019
Nakagawa K, et al.: Classification for invasion depth of esophageal squamous cell carcinoma using a deep neural network compared with experienced endoscopists. Gastrointestinal endoscopy 90:407-414, 2019
Ohmori M, et al.: Endoscopic detection and differentiation of esophageal lesions using a deep neural network. Gastrointestinal endoscopy 91:301-309, 2019
Yoon HJ, et al.: A Lesion-Based Convolutional Neural Network Improves Endoscopic Detection and Depth Prediction of Early Gastric Cancer. Journal of clinical medicine 8:1310, 2019
Zhu Y, et al.: Application of convolutional neural network in the diagnosis of the invasion depth of gastric cancer based on conventional endoscopy. Gastrointestinal endoscopy 89:806-815.e801, 2019
Wang P, et al.: Real-time automatic detection system increases colonoscopic polyp and adenoma detection rates: a prospective randomised controlled study. Gut 68:1813-1819, 2019
Misawa M, et al.: Characterization of Colorectal Lesions Using a Computer-Aided Diagnostic System for Narrow-Band Imaging Endocytoscopy. Gastroenterology 150:1531-1532.e1533, 2016
Song EM, et al.: Endoscopic diagnosis and treatment planning for colorectal polyps using a deep-learning model. Scientific reports 10:30, 2020
Hirasawa T, et al.: Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastric cancer : official journal of the International Gastric Cancer Association and the Japanese Gastric Cancer Association 21:653-660, 2018
Chartrand G, et al.: Deep Learning: A Primer for Radiologists. Radiographics : a review publication of the Radiological Society of North America, Inc 37:2113–2131, 2017
Goodfellow I, et al.: Generative Adversarial Nets. Adv Neural Inf Process Syst 2672–2680, 2014
Schlegl T, Seeböck P, Waldstein SM, Langs G, Schmidt-Erfurth U: f-AnoGAN: Fast unsupervised anomaly detection with generative adversarial networks. Medical image analysis 54:30-44, 2019
Zhang J, et al.: Viral Pneumonia Screening on Chest X-rays Using Confidence-Aware Anomaly Detection. IEEE transactions on medical imaging, 2020
Kazeminia S, et al.: GANs for medical image analysis. Artificial Intelligence in Medicine 101938, 2020
Kim M, et al.: Deep learning in medical imaging. Neurospine 16:657, 2019
Han C, et al.: MADGAN: unsupervised medical anomaly detection GAN using multiple adjacent brain MRI slice reconstruction. BMC bioinformatics 22:1-20, 2021
Zhou K, et al.: Sparse-gan: Sparsity-constrained generative adversarial network for anomaly detection in retinal oct image. Proc. 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI): City
Karras T, Aila T, Laine S, Lehtinen J: Progressive growing of gans for improved quality, stability, and variation. arXiv preprint arXiv 171010196, 2017
Park HY, et al.: Realistic High-Resolution Body Computed Tomography Image Synthesis by Using Progressive Growing Generative Adversarial Network: Visual Turing Test. JMIR Medical Informatics 9:e23328, 2021
Heusel M, Ramsauer H, Unterthiner T, Nessler B, Klambauer G, Hochreiter S: GANs Trained by a Two Time-Scale Update Rule Converge to a Nash Equilibrium, 2017
He K, Zhang X, Ren S, Sun J: Deep residual learning for image recognition. Proc. Proceedings of the IEEE conference on computer vision and pattern recognition: City
McKight PE, Najab J: Kruskal‐wallis test. The corsini encyclopedia of psychology 1–1, 2010
Lesack K, Naugler C: An open-source software program for performing Bonferroni and related corrections for multiple comparisons. Journal of pathology informatics 2, 2011
Shrout PE, Fleiss JL: Intraclass correlations: uses in assessing rater reliability. Psychological bulletin 86:420, 1979
Lee JG, et al.: Deep Learning in Medical Imaging: General Overview. Korean J Radiol 18:570-584, 2017
Yalamarthi S, Witherspoon P, McCole D, Auld CD: Missed diagnoses in patients with upper gastrointestinal cancers. Endoscopy 36:874-879, 2004
Lee JH, et al.: Spotting malignancies from gastric endoscopic images using deep learning. Surgical Endoscopy 33:3790-3797, 2019
Funding
This study was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (HI18C0022, and HI18C2383).
Author information
Authors and Affiliations
Contributions
K.W.S.: methodology, software, formal analysis, investigation, data curation, writing—original draft, and visualization. J.S.L.: methodology, investigation, writing—original draft, and validation. J.Y.L.: writing—review and editing, and validation. H.S.L.: writing—review and editing. J.S.K.: writing—review and editing. J.S.B.: writing—review and editing. H.Y.J.: writing—review and editing. D.H.K.: conceptualization, resources, writing—review and editing. N.K.K.: conceptualization, writing—review and editing, supervision, project administration, and funding acquisition.
Corresponding authors
Ethics declarations
Ethics Approval
This retrospective study was conducted according to the principles of the Declaration of Helsinki and was performed in accordance with current scientific guidelines. The study protocol was approved by the Institutional Review Board Committee of Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea (IRB No. 2020–0568). The requirement for informed patient consent was waived by the Institutional Review Board Committee of Asan Medical Center.
Conflict of Interest
The authors except Namkug Kim have no conflicts of interest to disclose. Namkug Kim is a stakeholder of Promedius Inc.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shin, K., Lee, J.S., Lee, J.Y. et al. An Image Turing Test on Realistic Gastroscopy Images Generated by Using the Progressive Growing of Generative Adversarial Networks. J Digit Imaging 36, 1760–1769 (2023). https://doi.org/10.1007/s10278-023-00803-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-023-00803-2