Abstract
l-Arabinose (l-Ara) is a plant-specific sugar accounting for 5–10 % of cell wall saccharides in Arabidopsis (Arabidopsis thaliana) and rice (Oryza sativa). l-Ara occurs in pectic arabinan, rhamnogalacturonan II, arabinoxylan, arabinogalactan-protein (AGP), and extensin in the cell walls, as well as in glycosylated signaling peptides like CLAVATA3 and small glycoconjugates such as quercetin 3-O-arabinoside. This review focuses on recent advances towards understanding the generation of l-Ara and the metabolism of l-Ara-containing molecules in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
l-Arabinose (l-Ara) is a plant saccharide that is not found in animals. Like xylose (Xyl—we omit the D-prefix of sugars belonging to the d-series), l-Ara is a pentose comprising five carbons, not six carbon containing hexose like glucose (Glc) and galactose (Gal) (Fig. 1). Although its content in the cell walls varies depending on the plant species, l-Ara can be considered a major sugar. It accounts for 5–10 % of cell wall sugar, for instance, in Arabidopsis (Arabidopsis thaliana) and rice (Oryza sativa) (Konishi et al. 2011; Zablackis et al. 1995). The fact that l-Ara is widely distributed not only in land plants including liverworts and mosses but also found in several Chlorophycean and Charophycean green algae suggests that the metabolic pathway for the synthesis of l-Ara was acquired early by primitive plants (Domozych et al. 2009, 2012; Konno et al. 2010; Lee et al. 2005; Popper and Fry 2003; Roberts et al. 2012; Thomas 1977). The broad range of l-Ara-containing molecules seen in land plants today is likely due to subsequent diversification of the use of l-Ara during plant evolution.
Structure of β-l-Araf and β-l-Arap. Sugars are drawn in the Haworth projection. The furanose β-l-Araf has the shape of a pentagon, whereas the pyranose β-l-Arap forms a hexagon. The structure of β-l-Arap is similar to that of α-Gal and the C-4 epimer of α-Xyl. l-Ara and Xyl are pentoses, whereas Gal and Glc are hexoses
l-Ara may be useful as a natural pharmaceutical. Monomeric l-Ara inhibits intestinal maltase and sucrase (α-glucosidase hydrolyzing sucrose) activities in vitro (Seri et al. 1996). In rats, dietary sucrose increases the insulin level in blood and triacylglycerol levels in blood plasma and the liver, but feeding l-Ara together with sucrose can significantly reduce the increase in these levels (Osaki et al. 2001; Seri et al. 1996). Recently, the effect of l-Ara on controlling insulin and blood-Glc levels was also observed in humans (Kaats et al. 2011). While its effect in humans is still controversial (Halschou-Jensen et al. 2015), the use of l-Ara for these purposes is receiving attention and becoming more wide-spread.
Several excellent reviews have surveyed nucleotide sugar synthesis and sugar metabolism in land plants (Bar-Peled and O’Neill 2011; Bar-Peled et al. 2012; Lagaert et al. 2014; Reiter 2008; Reiter and Vanzin 2001; Seifert 2004). Here we concentrate on recent progress in our understanding of the generation of l-Ara and the synthesis and degradation of l-Ara-containing molecules in land plants.
l-Ara-containing molecules in plants
l-Ara has two ring forms, called l-arabinopyranose (l-Arap, sugars other than l-Ara are in pyranose form unless stated otherwise) and l-arabinofuranose (l-Araf), respectively (Fig. 1). Free l-Ara exists as l-Arap in solution because the pyranose form is more stable than the furanose, but among cell wall polysaccharides and glycoproteins/proteoglycans, l-Araf residues outnumber the l-Arap residues. Representative l-Ara-containing molecules in plants are listed in Table 1.
Pectin is a complex molecule with many different domains, including homogalacturonan, rhamnogalacturonan I (RG-I), and rhamnogalacturonan II (RG-II) (Mohnen 2008; Tan et al. 2013; Willats et al. 2001). Pectic RG-I is a domain to which α-1,3:1,5-arabinan (pectic arabinan) and type I arabinogalactan (AG) are attached (Table 1, Fig. 2a) (Mohnen 2008). The pectic arabinan consists of α-l-Araf residues and is a major l-Ara-containing molecule in the cell walls in many plants (Levigne et al. 2004). ARABINAN DEFICIENT 1 (ARAD1) and ARAD2 are glycosyltransferases associated with the synthesis of pectic arabinan (Harholt et al. 2012). Based on amino acid sequence and structural similarity, ARADs are categorized into glycosyltransferase family (GTF) 47 (Campbell et al. 1997; Coutinho et al. 2003) (Table 2). The importance of pectic arabinan in the regulation of stomata opening was demonstrated in a study using endo-α-1,5-arabinanase specifically degrading α-1,5-arabinan main chains (Jones et al. 2003). α-l-Araf residues also exist in type I AG, another type of RG-I side chain, where they appear as non-reducing terminal residues (Nakamura et al. 2001) (Table 1). RG-II is the most complicated domain comprising more than ten types of sugars and includes β-l-Araf and α-l-Arap residues (Bar-Peled et al. 2012; O’Neill et al. 2001) (Table 1).
Arabinogalactan-proteins (AGPs) constitute a family of plant extracellular proteoglycans with a large carbohydrate moiety rich in l-Ara and Gal. In order to distinguish it from the type I AG of pectin, the glycan of AGP is called type II AG. The basic structure of type I AG is β-1,4-galactan, whereas that of type II AG is β-1,3:1,6-galactan (main chain, β-1,3-galactan; side chain, β-1,6-galactan) (Shimoda et al. 2014; Tan et al. 2004, 2010; Tsumuraya et al. 1988). In AGP, α-l-Araf residues exist as non-reducing terminal residues of type II AG. AGP sometimes has continuous α-l-Araf residues linked through α-1,5-linkages, thus resembling pectic arabinan (Tan et al. 2004; Tryfona et al. 2012). However, it is still unknown whether ARADs also participate in the synthesis of this structure in AGP. In wheat AGP, the α-l-Araf residues are further substituted with β-l-Arap residues (Tryfona et al. 2010) (Table 1; Fig. 2b). The activity of glycosyltransferase (ArapT) catalyzing the transfer of β-l-Arap from UDP-l-Arap onto α-l-Araf residues was detected in the microsomal fraction of mung bean seedlings (Ishii et al. 2005), but the gene encoding this glycosyltransferase has not been identified (Table 2).
In the vegetative tissues of grasses, including rice (Oryza sativa) and wheat (Triticum aestivum), instead of pectic arabinan, arabinoxylan is a major l-Ara-containing molecule in the cell walls (Table 1). The α-l-Araf residues are attached to the β-1,4-xylan main chain through α-1,2- and/or α-1,3-linkages. The α-1,3-l-Araf residues of arabinoxylan are formed by xylan arabinofuranosyltransferase (XAT), which belongs to GTF 61 (Anders et al. 2012). Interestingly, the α-l-Araf residues can be further substituted with ferulic acid, which forms cross-links between arabinoxylans (Grabber et al. 1995; Saulnier et al. 1999). This cross-link formation is physiologically important, as it is regulated by environmental signals including light and osmotic stress and affects cell wall extensibility, thereby controlling growth and development (Parvez et al. 1997; Tan et al. 1992; Wakabayashi et al. 1997, 2015).
Xyloglucan is a major hemicellulosic polysaccharide in many dicotyledonous plants. This polysaccharide usually consists of β-Glc, α-Xyl, β-Gal, and α-l-fucose (α-l-Fuc), but in several plants such as potato and olive, the β-Gal residues are replaced by α-l-Araf residues (Table 1) (Jia et al. 2003; Vierhuis et al. 2001; Vincken et al. 1996; York et al. 1996). The glycosyltransferases catalyzing the transfer of α-l-Araf residues onto the xylosyl (Xyl) residues, xyloglucan S-side chain transferases (XSTs), have been identified. XSTs are members of GTF 47, which also includes Xyloglucan l-side chain galactosylTransferase 2 (XLT2) and MURUS3 catalyzing the transfer of β-Gal residues onto the Xyl residues (Schultink et al. 2013).
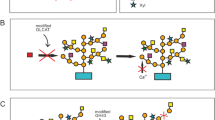
Extensins form a class of cell wall glycoproteins with Hyp-rich core-protein and contain arabino-oligosaccharides consisting of α-l-Araf and β-l-Araf residues (Kieliszewski et al. 1995; Lamport et al. 1973; McNeill et al. 1984) (Fig. 2c; Table 1). Surprisingly, a glycoprotein from Chlamydomonas reinhardtii appears to have similar arabinan chains, that is, the proximal two residues linked to Hyp, β-l-Araf1→2β-l-Araf1→Hyp, are identical to those of extensin (Bollig et al. 2007). This fact suggests that some of Chlorophycean green algae and land plants share the basic mechanism for the synthesis of this arabino-oligosaccharides and points to the possibility that l-arabinofuranosyltransferases and the metabolic pathway for UDP-l-Araf may be conserved.
Extensin-type arabino-oligosaccharides are also attached to glycosylated signaling peptides, the CLAVATA3 (CLV3)/Endosperm surrounding region-related (CLE) peptides (Ohyama et al. 2009; Okamoto et al. 2013; Xu et al. 2015). Using synthetic peptides with or without β-l-Araf1→2β-l-Araf1→2β-l-Araf chain, it has been demonstrated that the arabino-oligosaccharide is necessary for the proper function of CLV3 as a signaling molecule (Ohyama et al. 2009). The transfer of the first β-l-Araf residue onto Hyp is catalyzed by Hyp O-arabinosyltransferases (HPAT) classified into GTF 95 (Table 2). Indeed, loss of function mutations of HPAT genes causes pleiotropic phenotypes in Arabidopsis (Ogawa-Ohnishi et al. 2013). The importance of this arabino-oligosaccharide is further demonstrated by the existence of a tomato inflorescence branching mutant with extra flower and fruit organs, which has defects in a gene encoding GTF77 β-l-arabinofuranosyltransferase that synthesizes the β-l-Araf residues, and can be rescued by treatment with l-arabinofuranosylated CLV3 peptide (Xu et al. 2015).
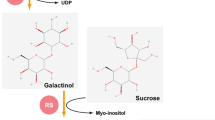
Most l-Ara-containing polysaccharides, proteoglycans, glycoproteins, and secreted peptides are synthesized in the Golgi apparatus, but small l-Ara-containing glycoconjugates are synthesized in the cytosol. Flavonoids are good examples of small glycoconjugates with l-Araf and l-Arap residues (Table 1). In Arabidopsis, a number of l-arabinopyranosylated flavonols have been found (Tueber and Herrmann 1978; Yonekura-Sakakibara et al. 2008). By transcriptome co-expression network analysis using public databases, an l-arabinosyltransferase, UGT78D3, was identified. This enzyme participates in the synthesis of quercetin 3-O-l-arabinoside in Arabidopsis (Yonekura-Sakakibara et al. 2008) (Table 2). Other l-arabinopyranosylated small glycoconjugates were also found. Floratheasaponin in the tea (Camellia sinensis) plant has an α-l-Arap residue in its carbohydrate moiety (Yoshikawa et al. 2005).
Generation of l -Ara
l-Ara is synthesized as a form of UDP-l-Arap from UDP-Xyl by UDP-Xyl 4-epimerases (UXEs) through C-4 epimerization of UDP-Xyl (Fig. 3). This is the only reaction route known to generate l-Ara in plants so far. It is also possible to synthesize UDP-l-Arap from UDP-galacturonic acid (UDP-GalA) through C-6 decarboxylation. However, no UDP-GalA decarboxylase forming UDP-l-Arap from UDP-GalA has been found in plants so far, although UDP-glucuronic acid (UDP-GlcA) decarboxylase (other name, UDP-Xyl synthase, UXS) forming UDP-Xyl from UDP-GlcA exists in many eukaryotes, including plants (Bar-Peled et al. 2001; Harper and Bar-Peled 2002).
Synthesis and degradation of l-Ara-containing molecules. UDP-l-Arap is synthesized in the de novo pathway in the Golgi apparatus and cytosol, which is shown in the left side. UDP-l-Arap is further converted to UDP-l-Araf in the cytosol. UDP-l-Araf and UDP-l-Arap serve as donor substrates in the synthesis of l-Ara-containing molecules. l-Ara-containing molecules undergo degradation by various glycoside hydrolases in the cell walls. The released l-Ara is recycled to UDP-l-Arap in the salvage pathway shown in the right side. The figure combines several reactions routes found in various plant species and is not to be understood to depict pathways found in any one particular plant. The enzymes in the de novo and salvage pathway are listed in Table 3
The reaction synthesizing UDP-l-Arap is part of the de novo pathway for UDP-sugars (Fig. 3). The enzymes constituting the de novo pathway are listed in Table 3. In the de novo pathway, UDP-Glc, the starting substrate of this pathway, is first synthesized from sucrose and UDP by sucrose synthase (SUS) (Baud et al. 2004; Cardini et al. 1955) or from Glc 1-phosphate (Glc 1-P) and UTP by UDP-Glc pyrophosphorylase (UGP) (Meng et al. 2009; Park et al. 2010) or UDP-sugar pyrophosphorylase (USP) (Kotake et al. 2004, 2007; Litterer et al. 2006). UDP-Glc undergoes C-6 oxidation, which turns it into UDP-GlcA by UDP-Glc dehydrogenase (UGD) (Reboul et al. 2011), and then undergoes C-6 decarboxylation to form UDP-Xyl by UXS. Plants have Golgi-localized and cytosolic UXSs, implying that the pathway is dual. In Arabidopsis, UXS1, 2, and 4 with a transmembrane domain at the N-terminus produce UDP-Xyl as Golgi-localized enzymes, whereas UXS3, 5, and 6 catalyze the same reaction in the cytosol. Other enzymes, UDP-apiose/UDP-Xyl synthases (AXSs) also participate in this reaction in the cytosol (Mølhøj et al. 2003). Following l-Ara forming reaction, C-4 epimerization of UDP-Xyl to form UDP-l-Arap, also occurs both in the Golgi apparatus and cytosol. In the Golgi apparatus, the formation of UDP-l-Arap is catalyzed by a Golgi-localized UXE, MURUS4 (MUR4) (Burget and Reiter 1999; Burget et al. 2003), but in the cytosol, it is catalyzed by bifunctional UGE1 and UGE3 in Arabidopsis (Fig. 3). Among five UGEs, only UGE1 and UGE3 possess UXE activity beside UDP-Glc 4-epimerase activity in Arabidopsis (Kotake et al. 2009). An Arabidopsis mur4 mutant shows a 50 % reduction in cell wall l-Ara, but a uge1 uge3 double mutant has normal cell walls (Burget and Reiter 1999; Rösti et al. 2007). These observations suggest that, at least in Arabidopsis, the main reaction to generate UDP-l-Arap occurs in the Golgi apparatus. The role of the cytosolic pathway may be clarified in the future via studies on mur4 uge1 uge3 triple mutants.
Conversion from UDP-l -Arap to UDP-l -Araf
The metabolism of UDP-l-Arap turned out to be more complicated than expected, when the subsequent conversion to UDP-l-Araf was investigated. The interconversion between UDP-l-Arap and UDP-l-Araf is catalyzed by a cytosolic enzyme, UDP-l-Arap mutase/reversibly glycosylated protein (UAM/RGP, Drakakaki et al. 2006; Konishi et al. 2007; Konishi et al. 2011). No Golgi-localized enzyme catalyzing this reaction has been found so far. It thus looks very much as if the main reaction to convert UDP-Xyl to UDP-l-Arap occurs in the Golgi apparatus, but the following conversion of UDP-l-Arap to UDP-l-Araf takes place in the cytosol. For this to work, two specific nucleotide sugar transporters (NSTs) would seem to be necessary: one transporter exporting UDP-l-Arap from the Golgi apparatus and one importing UDP-l-Araf into the Golgi apparatus. To efficiently incorporate synthesized UDP-l-Araf back into the Golgi apparatus, the mutase reaction may occur around the Golgi apparatus in the cytosol. Supporting this view, a recent proteomics analysis has revealed that one of the UAM/RGPs, RGP4, is associated with the Golgi apparatus in Arabidopsis (Nikolovski et al. 2012). NSTs are a family of proteins including 40 members in Arabidopsis, which are categorized into six subgroups (Rautengarten et al. 2014). To date, plant-specific NSTs for GDP-sugars, UDP-GalA/UDP-l-rhamnose, and UDP-Xyl are known (Baldwin et al. 2001; Ebert et al. 2015; Rautengarten et al. 2014), but those for UDP-l-Arap and UDP-l-Araf remain to be identified. While l-Ara mainly exists as a form of l-Araf residues in the cell walls, the level of UDP-l-Araf is lower than that of UDP-l-Arap in plant tissues (Ito et al. 2014; Pabst et al. 2010). It may be derived from the instability of UDP-l-Araf under the experimental condition. It is also conceivable that UDP-l-Araf imported by the transporter into the Golgi apparatus is immediately consumed by l-arabinofuranosyltransferases.
Origin of UDP-Xyl 4-epimerase
As described above, l-Ara is synthesized as a form of UDP-l-Arap by Golgi-localized MUR4 and cytosolic UGEs in land plants. These enzymes have similar amino acid sequences and are all categorized into the Rossmann fold superfamily (Rao and Rossman 1973), which also includes other UDP-sugar metabolizing enzymes: AXS, UXS, UDP-GlcA 4-epimerase, and UDP-l-rhamnose synthase (Diet et al. 2006; Gu and Bar-Peled 2004; Harper and Bar-Peled 2002; Mølhøj et al. 2003). In Arabidopsis, both of MUR4 and the bifunctional UGEs possess UXE activity, but the origin of these proteins seems different. It has thus been suggested that the UXE MUR4 is likely older than the bifunctional UGE.
UGE is one of the most highly conserved enzymes and found not only in eukaryotes but also in prokaryotes. In mammals, it is called UDP-Gal 4-epimerase (GalE), as the enzyme catalyzes interconversion between UDP-Glc and UDP-Gal, which constitutes the Leloir pathway for the detoxification of Gal. Phylogenetic relationships suggest that angiosperm UGEs can be grouped into UGE I and II families (Fig. 4). In Arabidopsis, two out of five UGEs, UGE1 and UGE3 belong to the UGE I family and the other three UGEs, UGE2, UGE4, and UGE5 s to the UGE II family. Biochemical characterization using recombinant UGEs expressed in Escherichia coli showed that Arabidopsis UGE1 and UGE3 have UXE activity beside UDP-Glc 4-epimerase activity but UGE2, UGE4, and UGE5 s have none or only weak UXE activity (Kotake et al. 2009). Interestingly, the Norway spruce (Picea sitchensis, gymnosperm) genome includes a gene encoding UGE corresponding to Arabidopsis UGE1 and UGE3, whereas no UGE from Physcomitrella patens (moss) and Selaginella moellendorffii (spikemoss) was grouped into the UGE I family (Fig. 4). These facts lead to the hypothesis that the UGE I family with UXE activity recently evolved from UGE without UXE activity. To support this conjecture, it would be necessary to characterize other UGEs, particularly UGEs in gymnosperm and moss. In fact, only two MUR4-related proteins, Arabidopsis MUR4 and barley (Hordeum vulgare) UXE1 (Zhang et al. 2010), and three bifunctional UGEs, Arabidopsis UGE1 and UGE3 and pea (Pisam sativum) UGE1, have been shown to possess UXE activity so far (Fig. 4).
Relationships of MUR4 homologues and UGEs. The phylogenetic relationships of MUR4 homologues and UGEs were analyzed using MEGA software (version 6.0, Tamura et al. 2013). The bar indicates substitutions per site. Red circles indicate enzymes that have been shown to possess UXE activity. MUR4-related proteins were taken without their transmembrane domain, which was removed according to the prediction obtained from the TMHMM program (Krogh et al. 2001). Accession numbers for the sequences are listed in Supplemental Table 1
In sharp contrast with the plant UGE I family, close homologues to Arabidopsis MUR4 can be found not only in gymnosperms, ferns, and mosses, but also in green algae. Although the biochemical properties of MUR4 homologues in green algae have not been determined, the existence of l-Ara-containing glycoprotein and UAM catalyzing subsequent conversion of UDP-l-Arap to UDP-l-Araf strongly suggests their role as UXE (Bollig et al. 2007; Kotani et al. 2013). Together with algal homologues, MUR4 and MUR4 homologues in land plants form a subclade apart from plant the UGE I and II families, suggesting that MUR4 is a highly conserved old enzyme in plants.
Degradation of l-Ara containing molecules
l-Ara-containing molecules undergo hydrolysis by glycoside hydrolases (GHs). The α-l-Araf residues of pectic arabinan, arabinoxylan, and AGP are hydrolyzed by α-l-arabinofuranosidases belonging to the GH family (GHF) 3 and GHF 51 in the cell walls in land plants (Fig. 3). Many plant GHF 3 and GHF 51 α-l-arabinofuranosidases are bifunctional enzymes with β-xylosidase activity (Arsovski et al. 2009; Kotake et al. 2006; Lee et al. 2003; Minic et al. 2004; Tateishi et al. 2005). A native GHF 3 α-l-arabinofuranosidase/β-xylosidase purified from radish, RsAraf1, hydrolyzes pectic arabinan, type I AG, AGP (type II AG), and arabinoxylan showing broad substrate specificity toward α-l-Araf residues (Hata et al. 1992). An Arabidopsis mutant with a defect in a GHF 51 α-l-arabinofuranosidase/β-xylosidase gene, araf1, exhibits accumulation of pectic arabinan in vascular tissues (Chávez Montes et al. 2008).
The enzyme hydrolyzing β-l-Arap residues of AGP has not been identified, but candidate genes exist in land plants. Microbial GHF 27 β-l-arabinopyranosidases acting on β-l-Arap residue of larch (Larix laricina) type II AG have been reported (Ichinose et al. 2009; Salama et al. 2012). Based on the similarity of amino acid sequences, four genes in the genome of Arabidopsis are presumed to encode GHF 27 β-l-arabinopyranosidase or α-galactosidase. As the structure of β-l-Arap resembles that of α-Gal (Fig. 1), it is not surprising that β-l-arabinopyranosidase and α-galactosidase exhibit quite similar three dimensional structures (Ichinose et al. 2009).
The hydrolysis of β-l-Araf residues of arabino-oligosaccharides of extensin and CLE peptides has so far remained elusive. A bacterial β-l-arabinofuranosidase including a domain of unknown function (DUF) 1680 has been identified in Bifidobacterium longum (Fujita et al. 2014). Several proteins of Arabidopsis have this domain, but the similarity of amino acid sequences is low (identity at amino acid level, <15 %). It is necessary to examine whether these proteins act on the β-l-Araf residues. No α-l-arabinopyranosidase acting on α-l-Arap residues of RG-II is known at all. It is conceivable that to some extent, α-l-Arap residues are hydrolyzed by GHF 35 β-galactosidase that widely exists in land plants, because α-l-Arap and β-Gal are structurally similar.
Recycling of free l -Ara released in the degradation
Free l-Ara released during the degradation and metabolism of l-Ara-containing molecules is recycled in the salvage pathway for the generation of nucleotide sugars. The enzymes constituting the salvage pathway are listed in Table 3. l-Arap is first phosphorylated by l-arabinokinase1 (ARA1) and turned into l-Arap 1-P (Fig. 3) (Dolezal and Cobbet 1991; Gy et al. 1998; Sherson et al. 1999). l-Arap 1-P is then converted to UDP-l-Arap by USP in the cytosol (Kotake et al. 2004, 2007; Litterer et al. 2006). This metabolic pathway probably functions as a third pathway for the generation of UDP-l-Arap parallel to the dual de novo pathways occurring in the Golgi apparatus and cytosol (Fig. 3). It is interesting that two plant aldopentoses, l-Ara and Xyl, undergo different metabolism although they are C-4 epimer sugars of each other (Fig. 1). Free Xyl differs from l-Ara in that it is predicted to be converted to xylulose by Xyl isomerase (Maehara et al. 2013) and metabolized in the pentose phosphate pathway. The salvage pathway for free l-Ara is implicated in the detoxification of l-Ara: an Arabidopsis ara1 mutant shows a severe growth defect in the presence of a high concentration of l-Ara (Dolezal and Cobbett 1991). Although no homozygous usp mutant has been analyzed—because USP is necessary for pollen development in Arabidopsis—, Geserick and Tenhaken (2013) have demonstrated the physiological importance of USP in vegetative tissue using USP-knock down (kd-usp) Arabidopsis. The kd-usp plant exhibited dwarf phenotype accumulating much free l-Ara and Xyl.
Physiological importance of the l-Ara salvage pathway in pollen development
Observing the remarkable reduction of cell wall l-Ara in Arabidopsis mur4 mutant (Burget and Reiter 1999; Burget et al. 2003), one is tempted to conclude that UDP-l-Arap is mainly synthesized in the de novo pathway. However, several lines of evidence indicate the physiological importance of the salvage pathway for UDP-l-Arap in developing pollens (Table 4). First, the rice l-arabinokinase named Collapsed Abnormal Pollen 1 (CAP1) has been shown to be necessary for normal development of pollens (Ueda et al. 2013), unfortunately, the effect of Arabidopsis ara1 mutation on pollen development has not been studied. Second, the pollen development also appears to be influenced by lack of USP that catalyzes the subsequent conversion of l-Ara 1-P to UDP-l-Arap as an Arabidopsis heterozygous usp mutant did not give any homozygous usp mutant (Kotake et al. 2007). In addition, collapsed pollens were observed in the anthers of heterozygous usp mutants and kd-usp plants (Geserick and Tenhaken 2013; Schnurr et al. 2006). As is the case in the vegetative tissues, UDP-l-Arap generated in the salvage pathway is probably converted to UDP-l-Araf by the action of UAM/RGPs in developing pollens. UAM/RGP was first identified as a factor necessary for the development of pollen in Arabidopsis (Drakakaki et al. 2006). Consistent with the phenotype of the rice cap1 mutant and the Arabidopsis usp mutant, collapsed pollen grains were also observed in the anthers of homo/hetero rgp1 rgp2 double mutants (rgp1/rgp1 RGP2/rgp2 mutant) (Drakakaki et al. 2006). Furthermore, RNA-interference of the rice UAM3 gene results in the formation of abnormal pollens lacking starch inside (Sumiyoshi et al. 2015). The importance of pectic arabinan for pollen development has been shown in the potato (Cankar et al. 2012), therefore it is highly probable that for pollen development the synthesis of a physiologically important portion of pectic arabinan depends on the salvage pathway.
Conclusion and future prospects
The plant-specific sugar l-Ara is generated as a form of UDP-l-Arap through C-4 epimerization of UDP-Xyl in the de novo pathway. This reaction is catalyzed by MUR4 in the Golgi apparatus and by bifunctional UGE in the cytosol. However, the exact extent of the cytosolic contribution to the synthesis of l-Ara-containing molecules and the physiological role of this alternate pathway are not quite clear. l-Ara appears as α-l-Araf, β-l-Araf, α-l-Arap, and β-l-Arap residues in plants. Various GTs and GHs are involved in the synthesis and degradation of these residues, but many pieces of the puzzle, in particular enzymes dealing with β-l-Araf and α-l-Arap residues, remain to be identified. Reasons for slow progress on this question may include the difficulty in the preparation of l-Ara-containing molecules as substrates. Given the physiological importance of l-Araf, it is not surprising that it is involved in various metabolic pathways. However, it is still unclear what significance in the evolution of plants the diversification of l-Ara use and the later emergence of a new pathway in the cytosol had. It would be of great interest to determine the relationship between other evolutionary events and the diversification of the l-Ara metabolism.
Change history
21 February 2018
The article “Metabolism of l-arabinose in plants”, written by “Toshihisa Kotake, Yukiko Yamanashi, Chiemi Imaizumi, Yoichi Tsumuraya”, was originally published Online First without open access. After publication in volume129, issue 5, page 781–792 the Botanical Society of Japan decided to opt for Open Choice and to make the article an open access publication.
References
Akiyama Y, Mori M, Kato K (1980) 13C-NMR analysis of hydroxyl-proline arabinosides from Nicotiana tabacum. Agric Biol Chem 44:2487–2489
Anders N, Wilkinson MD, Lovegrove A, Freeman J, Tryfona T, Pellny TK, Weimar T, Mortimer JC, Stott K, Baker JM, Defoin-Platel M, Shewry PR, Dupree P, Mitchell RAC (2012) Glycosyl transferases in family 61 mediate arabinofuranosyl transfer onto xylan in grasses. Proc Natl Acad Sci 109:989–993
Arsovski AA, Popma TM, Haughn GW, Carpita NC, McCann MC, Western TL (2009) AtBXL1 encodes a bifunctional β-d-xylosidase/α-l-arabinofuranosidase required for pectic arabinan modification in Arabidopsis mucilage secretory cells. Plant Physiol 150:1219–1234
Baldwin TC, Handford MG, Yuseff MI, Orellana A, Dupree P (2001) Identification and characterization of GONST1, a golgi-localized GDP-mannose transporter in Arabidopsis. Plant Cell 13:2283–2295
Bar-Peled M, O’Neill MA (2011) Plant nucleotide sugar formation, interconversion, and salvage by sugar recycling. Annu Rev Plant Biol 62:127–155
Bar-Peled M, Griffith CL, Doering TL (2001) Functional cloning and characterization of a UDP-glucuronic acid decarboxylase: the pathogenic fungus Cryptococcus neoformans elucidates UDP-xylose synthesis. Proc Natl Acad Sci 98:12003–12008
Bar-Peled M, Urbanowicz BR, O’Neill MA (2012) The synthesis and origin of the pectic polysaccharide rhamnogalacturonan II—insights from nucleotide sugar formation and diversity. Front Plant Sci 3:92
Baud S, Vaultier MN, Rochat C (2004) Structure and expression profile of the sucrose synthase multigene family in Arabidopsis. J Exp Bot 55:397–409
Bollig K, Lamshöft M, Schweimer K, Marner FJ, Budzikiewicz H, Waffenschmidt S (2007) Structural analysis of linear hydroxyproline-bound O-glycans of Chlamydomonas reinhardtii-conservation of the inner core in Chlamydomonas and land plants. Carbohydr Res 342:2557–2566
Burget EG, Reiter W-D (1999) The mur4 mutant of Arabidopsis is partially defective in the de novo synthesis of uridine diphospho l-arabinose. Plant Physiol 121:383–389
Burget EG, Verma R, Mølhøj M, Reiter W-D (2003) The biosynthesis of l-arabinose in plants: molecular cloning and characterization of a Golgi-localized UDP-d-xylose 4-epimerase encoded by the MUR4 gene of Arabidopsis. Plant Cell 15:523–531
Campbell JA, Davies GJ, Bulone V, Henrissat B (1997) A classification of nucleotide-diphospho-sugar glycosyltransferases based on amino acid sequence similarities. Biochem J 326:929–939
Cankar K, Kortstee A, Toonen MAJ, Wolters-Arts M, Houbein R, Mariani C, Ulvskov P, Jorgensen B, Schols HA, Visser RG, Trindade LM (2012) Pectic arabinan side chains are essential for pollen cell wall integrity during pollen development. Plant Biotechnol J 12:492–502
Cardini CE, Leloir LF, Chiriboga J (1955) The biosynthesis of sucrose. J Biol Chem 214:149–155
Chávez Montes RA, Ranocha P, Martinez Y, Minic Z, Jouanin L, Marquis M, Saulnier L, Fulton LM, Cobbett CS, Bitton F, Renou JP, Jauneau A, Goffner D (2008) Cell wall modifications in Arabidopsis plants with altered α-l-arabinofuranosidase activity. Plant Physiol 147:63–77
Coutinho PM, Deleury E, Davies GJ, Henrissat B (2003) An evolving hierarchical family classification for glycosyltransferases. J Mol Biol 328:307–317
Diet A, Link B, Seifert GJ, Schellenberg B, Wagner U, Pauly M, Reiter W-D, Ringli C (2006) The Arabidopsis root hair cell wall formation mutant lrx1 is suppressed by mutations in the RHM1 gene encoding a UDP-l-rhamnose synthase. Plant Cell 18:1630–1641
Dolezal O, Cobbett CS (1991) Arabinose kinase-deficient mutant of Arabidopsis thaliana. Plant Physiol 96:1255–1260
Domozych DS, Sørensen I, Willats WG (2009) The distribution of cell wall polymers during antheridium development and spermatogenesis in the Charophycean green alga, Chara corallina. Ann Bot 104:1045–1056
Domozych DS, Ciancia M, Fangel JU, Mikkelsen MD, Ulvskov P, Willats WG (2012) The cell walls of green algae: a journey through evolution and diversity. Front Plant Sci 3:82
Drakakaki G, Zabotina O, Delgado I, Robert S, Keegstra K, Raikhel N (2006) Arabidopsis reversibly glycosylated polypeptides 1 and 2 are essential for pollen development. Plant Physiol 142:1480–1492
Ebert B, Rautengarten C, Guo X, Xiong G, Stonebloom S, Smith-Moritz AM, Herter T, Chan LJG, Adams PD, Petzold CJ, Pauly M, Willats WGT, Heazlewood JL, Scheller HV (2015) Identification and characterization of a Golgi-localized UDP-xylose transporter family from Arabidopsis. Plant Cell 27:1218–1227
Fujita K, Takashi Y, Obuchi E, Kitahara K, Suganuma T (2014) Characterization of a novel β-l-arabinofuranosidase in Bifidobacterium longum: functional elucidation of a DUF1680 protein family member. J Biol Chem 289:5240–5249
Geserick C, Tenhaken R (2013) UDP-sugar pyrophosphorylase is essential for arabinose and xylose recycling, and is required during vegetative and reproductive growth in Arabidopsis. Plant J 74:239–247
Grabber JH, Hatfield RD, Ralph J, Zon J, Amrhein N (1995) Ferulate cross-linking in cell walls isolated from maize cell suspensions. Phytochemistry 40:1077–1082
Gu X, Bar-Peled M (2004) The biosynthesis of UDP-galacturonic acid in plants. Functional cloning and characterization of Arabidopsis UDP-d-glucuronic acid 4-epimerase. Plant Physiol 136:4256–4264
Gy I, Aubourg S, Sherson S, Cobbett CS, Cheron A, Kreis M, Lecharny A (1998) Analysis of a 14-kb fragment containing a putative cell wall gene and a candidate for the ARA1, arabinose kinase, gene from chromosome IV of Arabidopsis thaliana. Gene 209:201–210
Halabalaki M, Urbain A, Paschali A, Mitakou S, Tillequin F, Skaltsounis AL (2011) Quercetin and kaempferol 3-O-[α-l-rhamnopyranosyl-(1→2)-α-l-arabinopyranoside]-7-O-α-l-rhamnopyranosides from Anthyllis hermanniae: structure determination and conformational studies. J Nat Prod 74:1939–1945
Halschou-Jensen K, Knudsen KE, Nielsen S, Bukhave K, Andersen JR (2015) A mixed diet supplemented with l-arabinose does not alter glycaemic or insulinaemic responses in healthy human subjects. Br J Nutr 113:82–88
Harholt J, Jensen JK, Sørensen SO, Orfila C, Pauly M, Scheller HV (2006) ARABINAN DEFICIENT 1 is a putative arabinosyltransferase involved in biosynthesis of pectic arabinan in Arabidopsis. Plant Physiol 140:49–58
Harholt J, Jensen JK, Verhertbruggen Y, Søgaard C, Bernard S, Nafisi M, Poulsen CP, Geshi N, Sakuragi Y, Driouich A, Knox JP, Scheller HV (2012) ARAD proteins associated with pectic Arabinan biosynthesis form complexes when transiently overexpressed in planta. Planta 236:115–128
Harper AD, Bar-Peled M (2002) Biosynthesis of UDP-xylose. Cloning and characterization of a novel Arabidopsis gene family, UXS, encoding soluble and putative membrane-bound UDP-glucuronic acid decarboxylase isoforms. Plant Physiol 130:2188–2198
Hata K, Tanaka M, Tsumuraya Y, Hashimoto Y (1992) α-l-Arabinofuranosidase from radish (Raphanus sativus L.) seeds. Plant Physiol 100:388–396
Ichinose H, Fujimoto Z, Honda M, Harazono K, Nishimoto Y, Uzura A, Kaneko S (2009) A β-l-arabinopyranosidase from Streptomyces avermiilis is a novel member of glycoside hydrolase family 27. J Biol Chem 284:25097–25106
Ishii T, Konishi T, Ito Y, Ono H, Ohnishi-Kameyama M, Maeda I (2005) A β-(1→3)-arabinopyranosyltransferase that transfers a single arabinopyranose onto arabino-oligosaccharides in mung bean (Vigna radiate) hypocotyls. Phytochemistry 66:2418–2425
Ito J, Herter T, Baidoo EE, Lao J, Vega-Sánchez ME, Michelle Smith-Moritz A, Adams PD, Keasling JD, Usadel B, Petzold CJ, Heazlewood JL (2014) Analysis of plant nucleotide sugars by hydrophilic interaction liquid chromatography and tandem mass spectrometry. Anal Biochem 448:14–22
Jia Z, Qin Q, Darvill AG, York WS (2003) Structure of the xyloglucan produced by suspension-cultured tomato cells. Carbohydr Res 338:1197–1208
Jones L, Milne JL, Ashford D, McQueen-Mason SJ (2003) Cell wall arabinan is essential for guard cell function. Proc Natl Acad Sci 100:11783–11788
Kaats GR, Keith SC, Keith PL, Leckie RB, Perricone NV, Preuss HG (2011) A combination of l-arabinose and chromium lowers circulating glucose and insulin levels after an acute oral sucrose challenge. Nutr J 10:42
Kieliszewski MJ, O’Neill M, Leykam J, Orlando R (1995) Tandem mass spectrometry and structural elucidation of glycopeptides from a hydroxyproline-rich plant cell wall glycoprotein indicate that contiguous hydroxyproline residues are the major sites of hydroxyproline O-arabinosylation. J Biol Chem 270:2541–2549
Konishi T, Takeda T, Miyazaki Y, Ohnishi-Kameyama M, Hayashi T, O’Neill MA, Ishii T (2007) A plant mutase that interconverts UDP-arabinofuranose and UDP-arabinopyranose. Glycobiology 17:345–354
Konishi T, Aohara T, Igasaki T, Hayashi N, Miyazaki Y, Takahashi A, Hirochika H, Iwai H, Satoh S, Ishii T (2011) Down-regulation of UDP-arabinopyranose mutase reduces the proportion of arabinofuranose present in rice cell walls. Phytochemistry 72:1962–1968
Konno H, Nakashima S, Katoh K (2010) Metal-tolerant moss Scopelophila cataractae accumulates copper in the cell wall pectin of the protonema. J Plant Physiol 167:358–364
Kotake T, Yamaguchi D, Ohzono H, Hojo S, Kaneko S, Ishida HK, Tsumuraya Y (2004) UDP-sugar pyrophosphorylase with broad substrate specificity toward various monosaccharide 1-phosphates from pea sprouts. J Biol Chem 279:45728–45736
Kotake T, Tsuchiya K, Aohara T, Konishi T, Kaneko S, Igarashi K, Samejima M, Tsumuraya Y (2006) An α-l-arabinofuranosidase/β-d-xylosidase from immature seeds of radish (Raphanus sativus L.). J Exp Bot 57:2353–2362
Kotake T, Hojo S, Yamaguchi D, Aohara T, Konishi T, Tsumuraya Y (2007) Properties and physiological functions of UDP-sugar pyrophosphorylase in Arabidopsis. Biosci Biotechnol Biochem 71:761–771
Kotake T, Takata R, Verma R, Takaba M, Yamaguchi D, Orita T, Kaneko S, Matsuoka K, Koyama T, Reiter W-D, Tsumuraya Y (2009) Bifunctional cytosolic UDP-glucose 4-epimerases catalyse the interconversion between UDP-d-xylose and UDP-l-arabinose in plants. Biochem J 424:169–177
Kotani A, Tsuji M, Azama Y, Ishii T, Takeda T, Yamashita T, Shimojima M, Konishi T (2013) Purification and characterization of UDP-arabinopyranose mutase from Chlamydomonas reinhardtii. Biosci Biotechnol Biochem 77:1874–1878
Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Lagaert S, Pollet A, Courtin CM, Volckaert G (2014) β-Xylosidases and α-l-arabinofuranosidases: accessory enzymes for arabinoxylan degradation. Biotechnol Adv 32:316–332
Lamport DTA, Katona L, Roerig S (1973) Galactosylserine in extensin. Biochem J 133:125–132
Lee RC, Hrmova M, Burton RA, Lahnstein J, Fincher GB (2003) Bifunctional family 3 glycoside hydrolases from barley with α-l-arabinofuranosidase and β-d-xylosidase activity. Characterization, primary structures, and COOH-terminal processing. J Biol Chem 278:5377–5387
Lee KJ, Sakata Y, Mau SL, Pettolino F, Bacic A, Quatrano RS, Knight CD, Knox JP (2005) Arabinogalactan proteins are required for apical cell extension in the moss Physcomitrella patens. Plant Cell 17:3051–3065
Levigne SV, Ralet MCJ, Quéméner BC, Pollet BNL, Lapierre C, Thibault JFJ (2004) Isolation from sugar beet cell walls of arabinan oligosaccharides esterified by two ferulic acid monomers. Plant Physiol 134:1173–1180
Litterer LA, Schnurr JA, Plaisance KL, Storey KK, Gronwald JW, Somers DA (2006) Characterization and expression of Arabidopsis UDP-sugar pyrophosphorylase. Plant Physiol Biochem 44:171–180
Maehara T, Takabatake K, Kaneko S (2013) Expression of Arabidopsis thaliana xylose isomerase gene and its effect on ethanol production in Flammulina velutipes. Fungal Biol 117:776–782
McNeil M, Darvill AG, Fry SC, Albersheim P (1984) Structure and function of the primary cell walls of plants. Annu Rev Biochem 53:625–663
Meng M, Geisler M, Johansson H, Harholt J, Scheller HV, Mellerowicz EJ, Kleczkowski LA (2009) UDP-glucose pyrophosphorylase is not rate limiting, but is essential in Arabidopsis. Plant Cell Physiol 50:998–1011
Minic Z, Rihouey C, Do CT, Lerouge P, Jouanin L (2004) Purification and characterization of enzymes exhibiting β-d-xylosidase activities in stem tissues of Arabidopsis. Plant Physiol 135:867–878
Mohnen D (2008) Pectin structure and biosynthesis. Curr Opin Plant Biol 11:266–277
Mølhøj M, Verma R, Reiter WD (2003) The biosynthesis of the branched-chain sugar d-apiose in plants: functional cloning and characterization of a UDP-d-apiose/UDP-d-xylose synthase from Arabidopsis. Plant J 35:693–703
Nakamura A, Furuta H, Maeda H, Nagamatsu Y, Yoshimoto A (2001) Analysis of structural components and molecular construction of soybean soluble polysaccharides by stepwise enzymatic degradation. Biosci Biotechnol Biochem 65:2249–2258
Nikolovski N, Rubtsov D, Segura MP, Miles GP, Stevens TJ, Dunkley TPJ, Munro S, Lilley KS, Dupree P (2012) Putative glycosyltransferases and other plant Golgi apparatus proteins are revealed by LOPIT proteomics. Plant Physiol 160:1037–1051
Ogawa-Ohnishi M, Matsushita W, Matsubayashi Y (2013) Identification of three hydroxyproline O-arabinosyltransferases in Arabidopsis thaliana. Nat Chem Biol 9:726–730
Ohyama K, Shinohara H, Ogawa-Ohnishi M, Matsubayashi Y (2009) A glycopeptide regulating stem cell fate in Arabidopsis thaliana. Nat Chem Biol 5:578–580
Okamoto S, Shinohara H, Mori T, Matsubayashi Y, Kawaguchi M (2013) Root-derived CLE glycopeptides control nodulation by direct binding to HAR1 receptor kinase. Nat Commun 4:2191
O’Neill MA, Eberhard S, Albersheim P, Darvill AG (2001) Requirement of borate cross-linking of cell wall rhamnogalacturonan II for Arabidopsis growth. Science 294:846–849
Osaki S, Kimura T, Sugimoto T, Hizukuri S, Iritani N (2001) l-Arabinose feeding prevents increases due to dietary sucrose in lipogenic enzymes and triacylglycerol levels in rats. J Nutr 131:796–799
Pabst M, Grass J, Fischl R, Léonard R, Jin C, Hinterkörner G, Borth N, Altmann F (2010) Nucleotide and nucleotide sugar analysis by liquid chromatography-electrospray ionization-mass spectrometry on surface-conditioned porous graphitic carbon. Anal Chem 82:9782–9788
Park JI, Ishimizu T, Suwabe K, Sudo K, Masuko H, Hakozaki H, Nou IS, Suzuki G, Watanabe M (2010) UDP-glucose pyrophosphorylase is rate limiting in vegetative and reproductive phases in Arabidopsis thaliana. Plant Cell Physiol 51:981–996
Parvez MM, Wakabayashi K, Hoson T, Kamisaka S (1997) White light promotes the formation of diferulic acid in maize coleoptile cell walls by enhancing PAL activity. Physiol Plant 99:39–48
Popper ZA, Fry SC (2003) Primary cell wall composition of bryophytes and charophytes. Ann Bot 91:1–12
Rao ST, Rossmann MG (1973) Comparison of super-secondary structures in proteins. J Mol Biol 76:241–256
Rautengarten C, Ebert B, Moreno I, Temple H, Herter T, Link B, Doñas-Cofré D, Moreno A, Saéz-Aguayo S, Blanco F, Mortimer JC, Schultink A, Reiter W-D, Dupree P, Pauly M, Heazlewood JL, Scheller HV, Orellana A (2014) The Golgi localized bifunctional UDP-rhamnose/UDP-galactose transporter family of Arabidopsis. Proc Natl Acad Sci 111:11563–11568
Reboul R, Geserick C, Pabst M, Frey B, Wittmann D, Lütz-Meindl U, Léonard R, Tenhaken R (2011) Down-regulation of UDP-glucuronic acid biosynthesis leads to swollen plant cell walls and severe developmental defects associated with changes in pectic polysaccharides. J Biol Chem 286:39982–39992
Reiter W-D (2008) Biochemical genetics of nucleotide sugar interconversion reactions. Curr Opin Plant Biol 11:236–243
Reiter W-D, Vanzin GF (2001) Molecular genetics of nucleotide sugar interconversion pathways in plants. Plant Mol Biol 47:95–113
Roberts AW, Roberts EM, Haigler CH (2012) Moss cell walls: structure and biosynthesis. Front Plant Sci 3:166
Rösti J, Barton CJ, Albrecht S, Dupree P, Pauly M, Findlay K, Roberts K, Seifert GJ (2007) UDP-glucose 4-epimerase isoforms UGE2 and UGE4 cooperate in providing UDP-galactose for cell wall biosynthesis and growth of Arabidopsis thaliana. Plant Cell 19:1565–1579
Salama R, Alalouf O, Tabachnikov O, Zolotnitsky G, Shoham G, Shoham Y (2012) The abp gene in Geobacillus stearothermophilus T-6 encodes a GH27 β-l-arabinopyranosidase. FEBS Lett 586:2436–2442
Saulnier L, Crepeau M-J, Lahaye M, Thibault J-F, Garcia-Conesa MT, Kroon PA, Williamson G (1999) Isolation and structural determination of two 5,5′-diferuloyl oligosaccharides indicate that maize heteroxylans are covalently cross-linked by oxidatively coupled ferulates. Carbohydr Res 320:82–92
Schnurr JA, Storey KK, Jung HJG, Somers DA, Gronwald JW (2006) UDP-sugar pyrophosphorylase is essential for pollen development in Arabidopsis. Planta 224:520–532
Schultink A, Cheng K, Park YB, Cosgrove DJ, Pauly M (2013) The identification of two arabinosyltransferases from tomato reveals functional equivalency of xyloglucan side chain substituents. Plant Physiol 163:86–94
Seifert GJ (2004) Nucleotide sugar interconversions and cell wall biosynthesis: how to bring the inside to the outside. Curr Opin Plant Biol 7:277–284
Seri K, Sanai K, Matsuo N, Kawakubo K, Xue C, Inoue S (1996) l-Arabinose selectively inhibits intestinal sucrase in an uncompetitive manner and suppresses glycemic response after sucrose ingestion in animals. Metabolism 45:1368–1374
Sherson S, Gy I, Medd J, Schmidt R, Dean C, Kreis M, Lecharny A, Cobbett C (1999) The arabinose kinase, ARA1, gene of Arabidopsis is a novel member of the galactose kinase gene family. Plant Mol Biol 39:1003–1012
Shimoda R, Okabe K, Kotake T, Matsuoka K, Koyama T, Tryfona T, Liang HC, Dupree P, Tsumuraya Y (2014) Enzymatic fragmentation of carbohydrate moieties of radish arabinogalactan-protein and elucidation of the structures. Biosci Biotechnol Biochem 78:818–831
Sumiyoshi M, Inamura T, Nakamura A, Aohara T, Ishii T, Satoh S, Iwai H (2015) UDP-arabinopyranose mutase 3 is required for pollen wall morphogenesis in rice (Oryza sativa). Plant Cell Physiol 56:232–241
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tan KS, Hoson T, Masuda Y, Kamisaka S (1992) Involvement of cell wall-bound diferulic acid in lightinduced decrease in growth rate and cell wall extensibility of Oryza coleoptiles. Plant Cell Physiol 33:103–108
Tan L, Qiu F, Lamport DTA, Kieliszewski MJ (2004) Structure of a hydroxyproline (Hyp)-arabinogalactan polysaccharide from repetitive Ala-Hyp expressed in transgenic Nicotiana tabacum. J Biol Chem 279:13156–13165
Tan L, Varnai P, Lamport DTA, Yuan C, Xu J, Qiu F, Kieliszewski MJ (2010) Plant O-hydroxyproline arabinogalactans are composed of repeating trigalactosyl subunits with short bifurcated side chains. J Biol Chem 285:24575–24583
Tan L, Eberhard S, Pattathil S, Warder C, Glushka J, Yuan C, Hao Z, Zhu X, Avci U, Miller JS, Baldwin D, Pham C, Orlando R, Darvill A, Hahn MG, Kieliszewski MJ, Mohnen D (2013) An Arabidopsis cell wall proteoglycan consists of pectin and arabinoxylan covalently linked to an arabinogalactan protein. Plant Cell 25:270–287
Tateishi A, Mori H, Watari J, Nagashima K, Yamaki S, Inoue H (2005) Isolation, characterization, and cloning of α-l-arabinofuranosidase expressed during fruit ripening of Japanese pear. Plant Physiol 138:1653–1664
Teuber H, Herrmann K (1978) Flavonol glycosides of leaves and fruits of dill (Anethum graveolens L.). II. Phenolics of spices. Z Lebensm Unters Forsch 167:101–104
Thomas RJ (1977) Wall analyses of Lophocolea seta cells (bryophyta) before and after elongation. Plant Physiol 59:337–340
Torres-Mendoza D, González J, Ortega-Barría E, Heller MV, Capson TL, McPhail K, Gerwick WH, Cubilla-Rios L (2006) Weakly antimalarial flavonol arabinofuranosides from Calycolpus warszewiczianus. J Nat Prod 69:826–828
Tryfona T, Liang H-C, Kotake T, Kaneko S, Marsh J, Ichinose H, Lovegrove A, Tsumuraya Y, Shewry PR, Stephens E, Dupree P (2010) Carbohydrate structural analysis of wheat flour arabinogalactan protein. Carbohydr Res 345:2648–2656
Tryfona T, Liang H-C, Kotake T, Tsumuraya Y, Stephens E, Dupree P (2012) Structural characterisation of Arabidopsis leaf arabinogalactan polysaccharides. Plant Physiol 160:653–666
Tsumuraya Y, Ogura K, Hashimoto Y, Mukoyama H, Yamamoto S (1988) Arabinogalactan-proteins from primary and mature roots of radish (Raphanus sativus L.). Plant Physiol 86:155–160
Ueda K, Yoshimura F, Miyao A, Hirochika H, Nonomura K, Wabiko H (2013) COLLAPSED ABNORMAL POLLEN1 gene encoding the arabinokinase-like protein is involved in pollen development in rice. Plant Physiol 162:858–871
Vierhuis E, York WS, Kolli VSK, Vincken JP, Schols HA, Van Alebeek GWM, Voragen AGJ (2001) Structural analyses of two arabinose containing oligosaccharides derived from olive fruit xyloglucan: XXSG and XLSG. Carbohydr Res 332:285–297
Vincken JP, Wijsman AJM, Beldman G, Niessen WMA, Voragen AGJ (1996) Potato xyloglucan is built from XXGG-type subunits. Carbohydr Res 288:219–232
Virkki L, Maina HN, Johansson L, Tenkanen M (2008) New enzyme-based method for analysis of water-soluble wheat arabinoxylans. Carbohydr Res 343:521–529
Wakabayashi K, Hoson T, Kamisaka S (1997) Osmotic stress suppresses cell wall stiffening and the increase in cell wall-bound ferulic and diferulic acids in wheat coleoptiles. Plant Physiol 113:967–973
Wakabayashi K, Soga K, Hoson T, Kotake T, Yamazaki T, Higashibata A, Ishioka N, Shimazu T, Fukui K, Osada I, Kasahara H, Kamada M (2015) Suppression of hydroxycinnamate network formation in cell walls of rice shoots grown under microgravity conditions in space. PLoS One 10:e0137992
Westphal Y, Kühnel S, de Waard P, Hinz SWA, Schols HA, Voragen AGJ, Gruppen H (2010) Branched arabino-oligosaccharides isolated from sugar beet arabinan. Carbohydr Res 345:1180–1189
Willats WG, McCartney L, Mackie W, Knox JP (2001) Pectin: cell biology and prospects for functional analysis. Plant Mol Biol 47:9–27
Xu C, Liberatore KL, MacAlister CA, Huang Z, Chu YH, Jiang K, Brooks C, Ogawa-Ohnishi M, Xiong G, Pauly M, Van Eck J, Matsubayashi Y, van der Knaap E, Lippman ZB (2015) A cascade of arabinosyltransferases controls shoot meristem size in tomato. Nat Genet 47:784–792
Yonekura-Sakakibara K, Tohge T, Matsuda F, Nakabayashi R, Takayama H, Niida R, Watanabe-Takahashi A, Inoue E, Saito K (2008) Comprehensive flavonol profiling and transcriptome coexpression analysis leading to decoding gene-metabolite correlations in Arabidopsis. Plant Cell 20:2160–2176
York WS, Kumar Kolli VS, Orlando R, Albersheim P, Darvill AG (1996) The structures of arabinoxyloglucans produced by solanaceous plants. Carbohydr Res 285:99–128
Yoshikawa M, Morikawa T, Yamamoto K, Kato Y, Nagatomo A, Matsuda H (2005) Floratheasaponins A-C, acylated oleanane-type triterpene oligoglycosides with anti-hyperlipidemic activities from flowers of the tea plant (Camellia sinensis). J Nat Prod 68:1360–1365
Zablackis E, Huang J, Müller B, Darvill AG, Albersheim P (1995) Characterization of the cell-wall polysaccharides of Arabidopsis thaliana leaves. Plant Physiol 107:1129–1138
Zhang Q, Shirley NJ, Burton RA, Lahnstein J, Hrmova M, Fincher GB (2010) The genetics, transcriptional profiles, and catalytic properties of UDP-α-d-xylose 4-epimerases from barley. Plant Physiol 153:555–568
Author information
Authors and Affiliations
Corresponding author
Additional information
The original version of this article was revised due to a retrospective open access order.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kotake, T., Yamanashi, Y., Imaizumi, C. et al. Metabolism of l-arabinose in plants. J Plant Res 129, 781–792 (2016). https://doi.org/10.1007/s10265-016-0834-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-016-0834-z