Abstract
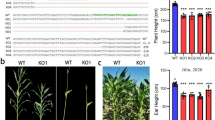
Medicinal Salvia miltiorrhiza contains two main groups of active pharmaceutical ingredients: lipid-soluble tanshinones and water-soluble phenolic acids, including rosmarinic acid and salvianolic acid B. Phenylalanine ammonia-lyase (PAL) catalyzes the first step in the phenylpropanoid pathway and is assumed to be closely related to the accumulation of rosmarinic acid and its derivatives. We selected a 217-bp fragment, located at the 3′ end of the coding region of PAL1, to establish an RNA interference construct that was introduced into S. miltiorrhiza via Agrobacterium tumefaciens-mediated transformation. PAL-suppressed plants exhibited several unusual phenotypes such as stunted growth, delayed root formation, altered leaves, and reduced lignin deposition. The total phenolic content was decreased by 20–70% in PAL-suppressed lines, and was accompanied by lower PAL activity. Down-regulation of PAL also affected the expression of C4H, 4CL2, and TAT, which are related genes in the rosmarinic acid pathway. Moreover, rosmarinic acid and salvianolic acid B were markedly reduced in PAL-suppressed lines, as detected by HPLC analysis. Our results indicate that PAL is very important for the synthesis of major water-soluble pharmaceutical ingredients within S. miltiorrhiza.






Similar content being viewed by others
References
Achnine L, Blancaflor EB, Rasmussen S, Dixon RA (2004) Colocalization of l-phenylalanine ammonia-lyase and cinnamate 4-hydroxylase for metabolic channeling in phenylpropanoid biosynthesis. Plant Cell 16:3098–3109
Bate NJ, Orr J, Ni W, Meromi A, Nadler-Hassar T, Doerner PW, Dixon RA, Lamb CJ, Elkind Y (1994) Quantitative relationship between phenylalanine ammonia-lyase levels and phenylpropanoid accumulation in transgenic tobacco identifies a rate-determining step in natural product synthesis. Proc Natl Acad Sci USA 91:7608–7612
Blount JW, Korth KL, Masoud SA, Rasmussen S, Lamb C, Dixon RA (2000) Altering expression of cinnamic acid 4-hydroxylase in transgenic plants provides evidence for a feedback loop at the entry point into the phenylpropanoid pathway. Plant Physiol 122:107–116
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Campbell MM, Sederoff RR (1996) Variation in lignin content and composition, mechanisms of control and implications for the genetic improvement of plants. Plant Physiol 110:3–13
Chen H, Chen F (2000) Effect of yeast elicitor on the secondary metabolism of Ti-transformed Salvia miltiorrhiza cell suspension cultures. Plant Cell Rep 19:710–717
Chen F, Reddy MS, Temple S, Jackson L, Shadle G, Dixon RA (2006) Multi-site genetic modulation of monolignol biosynthesis suggests new routes for formation of syringyl lignin and wall-bound ferulic acid in alfalfa (Medicago sativa L.). Plant J 48:113–124
Cochrane FC, Davin LB, Lewis NG (2004) The Arabidopsis phenylalanine ammonia lyase gene family: kinetic characterization of the four PAL isoforms. Phytochemistry 65:1557–1564
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085–1097
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Elkind Y, Edwards R, Mavandad M, Hedrick SA, Ribak O, Dixon RA, Lamb CJ (1990) Abnormal plant development and down-regulation of phenylpropanoid biosynthesis in transgenic tobacco containing a heterologous phenylalanine ammonia-lyase gene. Proc Natl Acad Sci USA 81:9057–9061
Gleave AP (1992) A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Goujon T, Sibout R, Eudes A, Mackay J, Jouanin L (2003) Genes involved in the biosynthesis of lignin precursors in Arabidopsis thaliana. Plant Physiol Biochem 41:677–687
Hahlbrock K, Scheel D (1989) Physiology and molecular biology of phenylpropanoid metabolism. Annu Rev Plant Physiol Plant Mol Biol 40:347–369
Higuchi M, Yoshizumi T, Kuriyama T, Hara H, Akagi C, Shimada H, Matsui M (2009) Simple construction of plant RNAi vectors using long oligonucleotides. J Plant Res 122:477–482
Hu P, Luo G, Zhao Z, Jiang Z (2005) Quality assessment of Radix Salviae Miltiorrhizae. Chem Pharm Bull (Tokyo) 53:481–486
Hu YS, Zhang L, Di P, Chen WS (2009) Cloning and induction of phenylalanine ammonia-lyase gene from Salvia miltiorrhiza and its effect on hydrophilic phenolic acids levels. Chin J Nat Med 7:449–457
Liu AH, Li L, Xu M, Lin YH, Guo HZ, Guo DA (2006a) Simultaneous quantification of six major phenolic acids in the roots of Salvia miltiorrhiza and four related traditional Chinese medicinal preparations by HPLC-DAD method. J Pharm Biomed Anal 41:48–56
Liu R, Xu S, Li J, Hu Y, Lin Z (2006b) Expression profile of a PAL gene from Astragalus membranaceus var mongholicus and its crucial role in flux into flavonoid biosynthesis. Plant Cell Rep 25:705–710
Lois R, Dietrich A, Hahlbrock K, Schulz W (1989) A phenylalanine ammonia-lyase gene from parsley: structure, regulation and identification of elicitor and light responsive cis-acting elements. EMBO J 8:1641–1648
Lu Y, Foo LY (2002) Polyphenolics of Salvia—a review. Photochemistry 59:117–140
Ma L, Zhang X, Guo H, Gan Y (2006) Determination of four water-soluble compounds in Salvia miltiorrhiza Bunge by high-performance liquid chromatography with a coulometric electrode array system. J Chromatogr B Analyt Technol Biomed Life Sci 833:260–263
MacDonald MJ, D’Cunha GB (2007) A modern view of phenylalanine ammonia lyase. Biochem Cell Biol 85:273–282
Mahesh V, Rakotomalala JJ, Le Gal L, Vigne H, de Kochko A, Hamon S, Noirot M, Campa C (2006) Isolation and genetic mapping of a Coffea canephora phenylalanine ammonia-lyase gene (CcPAL1) and its involvement in the accumulation of caffeoyl quinic acids. Plant Cell Rep 25:986–992
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue culture. Physiol Plant 15:473–497
Olsen KM, Lea US, Slimestad R, Verheul M, Lillo C (2008) Differential expression of four Arabidopsis PAL genes; PAL1 and PAL2 have functional specialization in abiotic environmental-triggered flavonoid synthesis. J Plant Physiol 165:1491–1499
Petersen M, Simmonds MS (2003) Rosmarinic acid. Phytochemistry 62:121–125
Petersen M, Haüsler E, Karwatzki B, Meinhard J (1993) Proposed biosynthetic pathway for rosmarinic acid in cell cultures of Coleus blumei Benth. Planta 189:10–14
Razzaque A, Ellis BE (1977) Rosmarinic acid production in Coleus cell cultures. Planta 137:287–291
Rohde A, Morreel K, Ralph J, Goeminne G, Hostyn V, De Rycke R, Kushnir S, Van Doorsselaere J, Joseleau JP, Vuylsteke M, Van Driessche G, Van Beeumen J, Messens E, Boerjan W (2004) Molecular phenotyping of the pal1 and pal2 mutants of A. thaliana reveals far-reaching consequences on phenylpropanoid, amino acid, and carbohydrate metabolism. Plant Cell 16:2749–2771
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Sewalt V, Ni W, Blount JW, Jung HG, Masoud SA, Howles PA, Lamb C, Dixon RA (1997) Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of l-phenylalanine ammonia-lyase or cinnamate 4-hydroxylase. Plant Physiol 115:41–50
Shadle GL, Wesley SV, Korth KL, Chen F, Lamb C, Dixon RA (2003) Phenylpropanoid compounds and disease resistance in transgenic tobacco with altered expression of L-phenylalanine ammonia-lyase. Phytochemistry 64:153–161
Song J, Wang Z (2009) Molecular cloning, expression and characterization of a phenylalanine ammonia-lyase gene (SmPAL1) from Salvia miltiorrhiza. Mol Biol Rep 36:939–952
Wanner LA, Li G, Ware D, Somssich IE, Davis KR (1995) The phenylalanine ammonialyase gene family in Arabidopsis thaliana. Plant Mol Biol 27:327–338
Wesley SV, Helliwell CA, Smith NA, Wang MB, Rouse DT, Liu Q, Gooding PS, Singh SP, Abbott D, Stoutjesdijk PA, Robinson SP, Gleave AP, Green AG, Waterhouse PM (2001) Construct design for efficient, effective and high throughput gene silencing in plants. Plant J 27:581–590
Yan Y, Wang Z (2007) Genetic transformation of the medicinal plant Salvia miltiorrhiza by Agrobacterium tumefaciens-mediated method. Plant Cell Tissue Organ Cult 88:175–184
Yan Q, Shi M, Ng J, Wu JY (2006) Elicitor-induced rosmarinic acid accumulation and secondary metabolism enzyme activities in Salvia miltiorrhiza hairy roots. Plant Sci 170:853–858
Zhang H, Yu C, Jia JY, Leung SW, Siow YL, Man RY, Zhu DY (2002) Contents of four active components in different commercial crude drugs and preparations of Danshen (Salvia miltiorrhiza). Acta Pharmacol Sin 23:1163–1168
Zhao SJ, Hu ZB, Liu D, Leung FC (2006) Two divergent members of 4-coumarate:coenzyme A ligase from Salvia miltiorrhiza Bunge: cDNA cloning and functional study. J Integr Plant Biol 48:1355–1364
Zheng W, Wang SY (2001) Antioxidant activity and phenolic compounds in selected herbs. J Agric Food Chem 49:5165–5170
Zhou L, Zuo Z, Chow MS (2005) Danshen: an overview of its chemistry, pharmacology, pharmacokinetics, and clinical use. J Clin Pharmacol 45:1345–1359
Acknowledgments
This work was supported by the 10th–11th “five-year technique project” of the Ministry of Science and Technology of the People’s Republic of China (2004BA701A35, 2006BAI06A12-04).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Following files are unfortunately not in the Publisher's archive anymore:
-
Supplementary material 1 (TIFF 104 kb)
-
Supplementary material 2 (TIFF 430 kb)
-
Supplementary material 3 (TIFF 284 kb)
Rights and permissions
About this article
Cite this article
Song, J., Wang, Z. RNAi-mediated suppression of the phenylalanine ammonia-lyase gene in Salvia miltiorrhiza causes abnormal phenotypes and a reduction in rosmarinic acid biosynthesis. J Plant Res 124, 183–192 (2011). https://doi.org/10.1007/s10265-010-0350-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-010-0350-5