Abstract
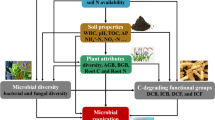
Effects of applying composted aquatic plants on soil chemistry, soil microbes (fungi and bacteria), and the growth of cultivated plant were demonstrated. To identify drivers promoting cultivated plant growth on soil amended with composted aquatic plant, empirical data of pot experiments were incorporated into structural equation models by hypothesizing causal relationships between the application of composted aquatic plants, soil chemistry, soil microbes, and cultivated plant growth. Cultivated plant growth, total carbon content, and bacterial and fungal richness in soil increased on soil applied with composted aquatic plants, and the composition of bacterial and fungal assemblages in soil were significantly different among the application treatments. Structural equation models explicitly demonstrated the relative importance of bacterial assemblages compared to soil chemistry as a promoter of cultivated plant growth in response to the application of composted aquatic plants. The present study is the first to demonstrate that the positive effects of composted aquatic plants on terrestrial plant growth are mediated by soil microbial processes. Our results could provide basic insights into the transfer and cycling of nutrients from aquatic to terrestrial systems through human activities.


Similar content being viewed by others
References
Antunes LP, Martins LF, Pereira RV, Thomas AM, Barbosa D, Lemos LN, Silva GMM, Moura LMS, Epamino GWC, Digiampietri LA, Lombardi KC, Ramos PL, Quaggio RB, de Oliveira JCF, Pascon RC, da Cruz JB, da Silva AM, Setubal JC (2016) Microbial community structure and dynamics in thermophilic composting viewed through metagenomics and metatranscriptomics. Sci Rep 6:38915
Ban S, TodaT KM, Ishikawa K, Kohzu A, Imai A (2019) Modern lake ecosystem management by sustainable harvesting and effective utilization of aquatic macrophytes. Limnology 20:93–100
Browne MW, Cudeck R (1993) Alternative ways of assessing model fit. In: Bollen KA, Long JS (eds) Testing structural equation models. Sage Publications, Newbury Park, pp 136–162
Chambers PA, Lacoul P, Murphy KJ, Thomaz SM (2008) Global diversity of aquatic macrophytes in freshwater. Hydrobiologia 595:9–26
Chao A, Jost L (2012) Coverage-based rarefaction and extrapolation: standardizing samples by completeness rather than size. Ecology 93:2533–2547
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Fujii S, Mori AS, Koide D, Makoto K, Matsuoka S, Osono T, Isbell F (2017) Disentangling relationships between plant diversity and decomposition processes under forest restoration. J Appl Ecol 54:80–90
Gardes M, Bruns TD (1993) ITS primer with enhanced specificity for basidiomycetes: application to the identification of mycorrhizae and rust. Mol Ecol 2:113–118
Gratton C, Zanden MJV (2009) Productivity to land: comparison of lentic and lotic ecosystems. Ecology 90:2689–2699
Gunnarsson CC, Petersen CM (2007) Water hyacinths as a resource in agriculture and energy production: a literature review. Waste Manag 27:117–129
Hamady M, Walker JJ, Harris JK, Gold NJ, Knight R (2008) Error-correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat Methods 5:235–237
Hasegawa S (1939) Utilization and effect of aquatic plants at lakesides of Lake Biwa (in Japanese). Shiga Prefecture Agricultural Experiment Station, Shiga
Hooper D, Coughlan J, Mullen M (2008) Structural equation modelling: guidelines for determining model fit. Electron J Bus Res Methods 6:53–60
Li W, Fu L, Niu B, Wu S, Wooley J (2012) Ultrafast clustering algorithms for metagenomic sequence analysis. Brief Bioinform 13:656–668
Lin Y, Ye G, Kuzyakov Y, Liu D, Fan J, Ding W (2019) Long-term manure application increases soil organic matter and aggregation, and alters microbial community structure and keystone taxa. Soil Biol Biochem 134:187–196
Liu Z, Lozupone C, Hamady M, Bushman FD, Knight R (2007) Short pyrosequencing reads suffice for accurate microbial community analysis. Nucleic Acids Res 35:e120
Lori M, Symnaczik S, Mӓder P, De Deyn G, Gattinger A (2017) Organic farming enhance soil microbial abundance and activity—a meta-analysis and meta-regression. PLoS ONE 12:e0180442
Martínez-Viveros O, Jorquera MA, Crowley DE, Gajardo G, Mora ML (2010) Mechanisms and practical considerations involved in plant growth promotion by rhizobacteria. J Soil Sci Plant Nutr 10:293–319
Matsuoka S, Mori AS, Kawaguchi E, Hobara S, Osono T (2016a) Disentangling the relative importance of host tree community, abiotic environment, and spatial factors on ectomycorrhizal fungal assemblages along an elevation gradient. FEMS Microb Ecol 92:fiw044
Matsuoka S, Kawaguchi E, Osono T (2016b) Temporal distance decay of similarity of ectomycorrhizal fungal community composition in a subtropical evergreen forest in Japan. FEMS Microb Ecol 92:fiw061
Ohtsuka T, Kuwabara Y, Haga H (2004) Estimated distribution and biomass of a submerged macrophyte community in Lake Biwa’s South Basin using an echo-sounder (in Japanese with English abstract). Jpn J Limnol 65:13–20
Osono T (2014) Metagenomic approach growths insights into fungal diversity and functioning. In: Sota T, Kagata H, Ando Y, Utsumi S, Osono T (eds) Species diversity and community structure. Springer, Berlin, pp 1–23
Osono T, Takeda H (2004) Potassium, calcium, and magnesium dynamics during litter decomposition in a cool temperate forest. J For Res 9:23–31
Schindler DE, Smits AP (2017) Subsidies of aquatic resources in terrestrial ecosystems. Ecosystems 20:78–93
Tanabe AS (2016) Claident pipeline v0.2.2016.07.05, software distributed by the author at https://www.fifthdimension.jp/. Accessed 21 June 2019
Tanabe AS, Toju H (2013) Two new computational methods for universal DNA barcoding: a benchmark using barcode sequences of bacteria, archaea, animals, fungi, and land plants. PLoS ONE 8:e76910
Truog E (1930) The determination of the readily available phosphorus of soils. J Am Soc Agro 22:874–882
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Yabuzaki I, Kaneko M, Osono T, Hobara S (2014) Effects of composting years on chemical properties of waterweed compost and growth of Japanese mustard spinach (in Japanese with Engilsh abstract). J Rakuno Gakuen Univ 39:87–92
Zhang H, Sekiguchi Y, Hanada S, Hugenholtz P, Kim H, Kamagata Y, Nakamura K (2003) Gemmatimonas aurantiaca gen. nov., sp. nov., a gram-negative, aerobic, polyphosphate-accumulating micro-organism, the first cultured representative of the new bacterial phylum Gemmatimonadetes phyl. nov. Int J Syst Evol Microbiol 53:1155–1163
Acknowledgements
We thank Dr. N. Okuda, Dr. Y. Sakai, Dr. S. Asano, Dr. I. Tayasu, Dr. S. Yachi, Dr. K. Wakita, Dr. T. Iwata, and Dr. S. Ban for useful comments and suggestions on experimental design, data analyses, and interpretations; Dr. S. Fujinaga and members of Center for Ecological Research, Kyoto University for assistance in laboratory works and pot experiments; staffs of Lake Biwa Policy Division of Shiga Prefecture and Ohmi Environment Conservation Foundation for providing composted aquatic plants and soil used in the pot experiments; Dr. H. Doi for critical comments on the manuscript; and Dr. E. Nakajima for critical reading of the manuscript. This study received partial financial support from Research Project (D06-14200119) of the Research Institute for Humanity and Nature (RIHN), Ohmi Environment Conservation Foundation, and from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (MEXT) (No. 17K15199 and 18K05731).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Handling Editor: Ichiro Tayasu.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Matsuoka, S., Kobayashi, Y., Hobara, S. et al. Identifying microbial drivers promoting plant growth on soil amended with composted aquatic plant: insight into nutrient transfer from aquatic to terrestrial systems. Limnology 21, 443–452 (2020). https://doi.org/10.1007/s10201-020-00613-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10201-020-00613-3