Abstract
Background
Fabry disease is an X-linked inherited lysosomal storage disorder caused by mutations in the gene encoding α-galactosidase A. Males are usually severely affected, while females have a wide range of disease severity. This variability has been assumed to be derived from organ-dependent skewed X-chromosome inactivation (XCI) patterns in each female patient. Previous studies examined this correlation using the classical methylation-dependent method; however, conflicting results were obtained. This study was established to ascertain the existence of skewed XCI in nine females with heterozygous pathogenic variants in the GLA gene and its relationship to the phenotypes.
Methods
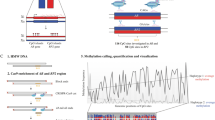
We present five female patients from one family and four individual female patients with Fabry disease. In all cases, heterozygous pathogenic variants in the GLA gene were detected. The X-chromosome inactivation patterns in peripheral blood leukocytes and cells of urine sediment were determined by both classical methylation-dependent HUMARA assay and ultra-deep RNA sequencing. Fabry Stabilization Index was used to determine the clinical severity.
Results
Skewed XCI resulting in predominant inactivation of the normal allele was observed only in one individual case with low ⍺-galactosidase A activity. In the remaining cases, no skewing was observed, even in the case with the highest total severity score (99.2%).
Conclusion
We conclude that skewed XCI could not explain the severity of female Fabry disease and is not the main factor in the onset of various clinical symptoms in females with Fabry disease.
Similar content being viewed by others
References
Mehta A, et al. Fabry disease: a review of current management strategies. QJM. 2010;103(9):641–59.
Desnick RJ. Fabry disease, an under-recognized multisystemic disorder:expert recommendations for diagnosis, management, and enzyme replacement therapy. Ann Intern Med. 2003;138:338–46.
Saito S, Ohno K, Sakuraba H. Fabry-database.org: database of the clinical phenotypes genotypes and mutant ⍺-galactosidase A sturctures in Fabry disease. J Hum Genet. 2011;56(6):467–8.
Fuller M, Meikle P, Hopwood J. Epidemiology of lysosomal storage diseases: an overview, in Fabry Disease: perspectives from 5 Years of FOS. In: Mehta A, Beck M, Sunder-Plassmann G, editors. Pharmagenesis. England: Oxford; 2006.
Sawada T, et al. Newborn screening for Fabry disease in the western region of Japan. Mol Genet Metab Rep. 2020;22:100562.
Inoue T, et al. Newborn screening for Fabry disease in Japan: prevalence and genotypes of Fabry disease in a pilot study. J Hum Genet. 2013;58(8):548–52.
Spada M, et al. High incidence of later-onset fabry disease revealed by newborn screening. Am J Hum Genet. 2006;79(1):31–40.
Deegan P. Natural history of Fabry disease in females in the Fabry Outcome Survey. J Med Genet. 2006;43(4):347–52.
Waldek S, et al. Life expectancy and cause of death in males and females with Fabry disease: findings from the Fabry Registry. Genet Med. 2009;11:790–6.
Whybra C, et al. The Main Severity Score Index: a new instrument for quantifying the Anderson–Fabry disease phenotype, and the response of patients to enzyme replacement therapy. Clin Genet. 2004;65:299–307.
Elstein D, et al. X-inactivation in Fabry disease. Gene. 2012;505:266–8.
Dobyns W, et al. Inheritance of most X-linked traits is not dominant or recessive, just X-linked. Am J Med Genet. 2004;129A:136–43.
Morey C, Avner P. Genetics and epigenetics of the X chromosome. Ann N Y Acad Sci. 2010;1214:E18-33.
Hossain MA, et al. Future clinical and biochemical predictions of Fabry disease in females by methylation studies of the GLA gene. Mol Genet Metab Rep. 2019;20:1–7.
Dobrovolny R. Relationship between X-inactivation and clinical in- volvement in Fabry heterozygotes. Eleven novel mutations in the alpha-galactosidase A gene in the Czech and Slovak population. J Mol Med (Berl). 2005;83(8):647–54.
Maier E. Disease manifestations and X inactivation in heterozygous females with Fabry disease. Acta Paediatr Suppl. 2006;95(451):30–8.
Allen RC, Zoghbi HY, Moseley A, Rosenblatt HF, Belmont J. Methylation of Hpall and Hhal sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51:1229–39.
Minamikawa S, et al. Development of ultra-deep targeted RNA sequencing for analyzing X-chromosome inactivation in female Dent disease. J Hum Genet. 2018;63:589–95.
Mignani R, et al. FAbry STabilization indEX (FASTEX): an innovative tool for the assessment of clinical stabilization in Fabry disease. Clin Kidney J. 2016;9(5):739–47.
Allen R, et al. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen receptor gene correlates With X Chromosome Inactivation. Am J Hum Genet. 1992;51(6):1229–39.
Lenders M, et al. Multicenter Female Fabry Study (MFFS)—clinical survey on current treatment of females with Fabry disease. Orphanet J Rare Dis. 2016;11:88.
Germain DP. Fabry disease. Orphanet J Rare Dis. 2010;5:1–49.
Eng CM, et al. Fabry disease: baseline medical characteristics of a cohort of 1765 males and females in the Fabry Registry. J Inherit Metab Dis. 2007;30:184–92.
Echevarria L, et al. X-chromosome inactivation in female patients with Fabry disease. Clin Genet. 2016;89:44–54.
Nowak A, et al. Plasma LysoGb3: a useful biomarker for the diagnosis and treatment of Fabry disease heterozygotes. Mol Genet Metab. 2017;120:57–61.
Juchniewicz P, et al. Female Fabry disease patients and X-chromosome inactivation. Gene. 2018;641:259–64.
Arcolino FO, et al. Human urine as a noninvasive source of kidney cells. Stem Cell Int. 2014;2015:1–7.
Botezatu I, et al. Genetic analysis of DNA excreted in urine: a new approach for detecting specific genomic DNA sequences from cells dying in an organism. Clin Chem. 2000;46:1078–84.
Lichtenstein A, et al. Circulating nucleic acids and apoptosis. Ann N Y Acad Sci. 2001;945:239–49.
Bittel D, et al. Comparison of X-chromosome inactivation patterns in multiple tissues from human females. J Med Genet. 2008;45:309–13.
Sharp A, Robinson D, Jacobs P. Age- and tissue-specific variation of X chromosome inactivation ratios in normal women. Hum Genet. 2000;107:343–9.
Gale R, et al. Tissue specificity of X-chromosome inactivation patterns. Blood. 1994;83:2899–905.
Azofeifa J, Waldherr R, Cremer M. X-chromosome methylation ratiosas indicators of chromosomal activity: evidence of intraindividual diver-gencies among tissues of different embryonal origin. Hum Genet. 1996;97:330–3.
Acknowledgements
This study was supported by Grant-in-Aid for Scientific Research (KAKENHI) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (subject ID: 19K08726 to Kandai Nozu). The authors thank Edanz (https://en-author-services.edanz.com/ac) for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Kandai Nozu has received lecture fees from Sumitomo Dainippon Pharma. Atsushi Fukunaga has received lecture fees from Sumitomo Dainippon Pharma and Sanofi K.K. Kazumoto Iijima has received lecture fees from Sanofi K.K. Hideki Fujii has received lecture fees from Sumitomo Dainippon Pharma, Sanofi K.K., and Amicus Therapeutics.
Ethical approval
All procedures performed in this study were reviewed and approved by the Institutional Review Board of Kobe University Graduate School of Medicine (IRB approval number 019-301) and performed in accordance with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Written informed consent for conducting this study was obtained from all participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Rossanti, R., Nozu, K., Fukunaga, A. et al. X-chromosome inactivation patterns in females with Fabry disease examined by both ultra-deep RNA sequencing and methylation-dependent assay. Clin Exp Nephrol 25, 1224–1230 (2021). https://doi.org/10.1007/s10157-021-02099-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10157-021-02099-4