Abstract
Purpose
This study aimed to analyze changes in the plasma concentration of EGFR-mutated circulating tumor DNA (ctDNA) occurring immediately after the start of therapy with EGFR tyrosine kinase inhibitors (TKIs).
Methods
Serial plasma samples were collected from 30 patients with EGFR-driven non-small cell lung cancer before intake of the first tablet and at 0.5, 1, 2, 3, 6, 12, 24, 36 and 48 h after the start of the therapy. The content of EGFR alleles (exon 19 deletions or L858R) in ctDNA was measured by ddPCR.
Results
ctDNA was detected at base-line in 25/30 (83%) subjects. Twelve (50%) out of 24 informative patients showed > 25% reduction of the ctDNA content at 48 h time point; all these patients demonstrated disease control after 4 and 8–12 weeks of therapy. The remaining 12 individuals showed either stable content of EGFR-mutated ctDNA (n = 5) or the elevation of ctDNA concentration (n = 7). 10 of 12 patients with elevated or stable ctDNA level achieved an objective response at 4 weeks, but only 5 of 10 evaluable patients still demonstrated disease control at 8–12 weeks (p = 0.032, when compared to the group with ctDNA decrease). The decline of the amount of circulating EGFR mutant copies at 48 h also correlated with longer progression-free survival (14.7 months vs. 8.5 months, p = 0.013).
Conclusion
Comparison of concentration of EGFR-mutated ctDNA at base-line and at 48 h after the start of therapy is predictive for the duration of TKI efficacy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peripheral blood obtained from oncological patients often contains tumor fragments, such as circulating malignant cells, tissue-specific proteins and cancer-derived nucleic acids. The analysis of circulating tumor DNA (ctDNA) is particularly promising, given that somatic mutations are highly specific for malignant cells and that current methods are capable of detecting even single mutated molecules present within a huge excess of normal tissues [1,2,3]. There are intensive efforts to utilize so-called “liquid biopsy” for early diagnosis and monitoring of cancer disease [4, 5]. It is more or less established, that the concentration of ctDNA is generally proportional to the overall tumor mass [6,7,8,9]. Consequently, ctDNA is relatively easily detectable in patients with extensive tumor disease, and its content usually declines after successful treatment [10,11,12,13].
The reduction of tumor size, which is achieved with the use of cancer drugs, is attributed to several biological effects. Conventional chemotherapy and targeted agents have a cytostatic action, i.e. they prevent proliferation of cancer cells [14, 15]. In addition, both cytotoxic and targeted drugs may provoke tumor shrinkage by inducing programmed cell death [16,17,18]. The reduction of overall tumor volume, which is usually achieved after weeks of systemic therapy, is almost always accompanied by the decrease of the concentration of circulating markers, be it tumor-specific proteins or ctDNA [19, 20]. However, the immediate marker response to the cancer therapy is less studied, which is at least in part attributed to the difficulties in collecting multiple serial blood samples within short time intervals.
EGFR tyrosine kinase inhibitors (TKIs; gefitinib, erlotinib, afatinib, osimertinib, etc.) are highly efficient in non-small cell lung carcinomas (NSCLCs), which harbor activating mutation in exon 19 or 21 of the EGFR gene [21,22,23,24,25]. Administration of EGFR TKIs for the treatment of EGFR-mutated NSCLC is almost always accompanied by the objective tumor response or the disease stabilization. EGFR inhibitors are also characterized by the “Lazarus effect”, i.e. dramatic symptomatic relief occurring within first hours after the drug administration [26, 27]. It is well established that the reduction of tumor size, which is observed during regular patient check-ups, is paralleled by the decline of the amount of EGFR-mutated ctDNA. However, short-term effects of EGFR TKIs on the level of ctDNA have not been systematically analyzed. We aimed to investigate, how the administration of EGFR TKIs influences the concentration of plasma ctDNA within the first hours after the uptake of the drug, and whether these changes are predictive for the long-term effects of systemic therapy.
Materials and methods
Patients
The study considered consecutive treatment-naïve patients with EGFR-mutated locally advanced or metastatic NSCLC, who were referred to the St.-Petersburg City Cancer Center between August 2018 and March 2020. EGFR mutations testing in tumor tissue was performed as described in [28]. Briefly, EGFR deletions in exon 19 (19del) were analyzed using the primers 5ʹ-CTGTCATAGGGACTCTGGAT-3ʹ and 5ʹ-CAGCAAAGCAGAAACTCACAT-3ʹ; PCR products were electrophoretically separated in 10% polyacrylamide gel; a 127 bp fragment corresponded to the wild-type sequence, and an additional band of smaller size was observed in the case of deletion. Testing for EGFR L858R mutation in exon 21 was performed by allele-specific real-time PCR with the wild-type-specific primer 5ʹ-CACCCAGCAGTTTGGCCA-3’, mutation-specific primer 5ʹ-CACCCAGCAGTTTGGCCC-3ʹ, and common primer 5’-GCATGAACTACTTGGAGGAC-3.
All patients provided informed consent for the participation in the study. The study was approved by the local Ethics Committee (protocol #20 “Evaluation the clinical value of ctDNA testing in patients with EGFR-mutated NSCLC”; approval date November 23, 2017).
Thirty patients were recruited to the investigation; their characteristics are described in Table 1 [see also Tables 1S and 2S in the electronic supplementary material (ESM)] for the description of individual patient data and response to TKI treatment). Serial plasma samples were collected before intake of the first tablet (at base-line) and at 0.5, 1, 2, 3, 6, 12, 24, 36 and 48 h after the “starting point” (Fig. 1). In addition, all patients were invited to donate blood after 14 and 28 days of the treatment. EGFR TKIs were given at regular daily doses (gefitinib: 250 mg; erlotinib: 150 mg; afatinib: 40 mg; osimertinib: 80 mg). Early response evaluations were performed with spiral computed tomography (CT) on the week 4, and the routine check-ups occurred within weeks 8–12 after the beginning of the therapy. Chest scans were performed before and after the administration of the contrast agent (100 mL of non-ionic iodinated contrast with a 100 mL saline chaser at 4.5–5 ml/s). All nodules with the size of more than 10 mm were measured. The images were based on venous-phase scans. Image reconstructions were performed on a CT workstation (Vitrea). Tumor burden was evaluated using the slice-by-slice pathology volume measurement with the slice thickness of 1.0 mm and assistance of the RadiAnt DICOM Viewer V.4.5.9.18463 software. Tumor responses RECIST v.1.1 criteria and progression-free survival (PFS) were evaluated according to standard guidelines by the study investigators [29].
ctDNA analysis
Blood samples (10 mL) were collected in cf-DNA/cf-RNA Preservative Tubes (Norgen) and the plasma was separated from the rest of the specimen by the two-step centrifugation protocol (400 g for 10 min at room temperature followed by 14,400 g for 10 min at 4 °C). Cell-free DNA was extracted with the QIAamp Circulating Nucleic Acid kit from 3–5 ml of plasma according to the manufacturer's instructions and dissolved in 50 μl of water.
The fractions of EGFR mutant alleles (exon 19 deletions or L858R substitutions) were measured by droplet digital PCR (ddPCR) using the QX100 Bio-Rad System [30]. ddPCR reactions were performed in triplicate and contained 2X ddPCR Supermix for Probes (no UTP, Bio-Rad), mutation-specific oligonucleotides (see Table 3S in the ESM) and 2–3 μl of the template DNA in a total reaction volume of 22–23 μl. Data analyses were performed with the QuantaSoft Software version 1.7.4 as recommended by the manufacturer. All ddPCR reactions, which yielded 10 or more droplets with the target DNA molecule, were considered informative.
The absolute number of tumor-derived “mutated” DNA copies in 1 mL of plasma (Cmut) was calculated according to the formula:
where: Concentration—number of «mutated» droplets per 1 μL of ddPCR reaction. V template—volume of ctDNA aliquot taken into ddPCR, μL. V dilution—total volume of diluted ctDNA sample collected from the plasma, μL. V plasma—volume of processed plasma, mL.
Statistics
Quantitative data were present as a median values/range or means ± 95% confidence interval (1.960σx̄). Non-parametric Wilcoxon Signed Rank Test and Mann–Whitney U Test were utilized to compare the medians. p value of < 0.05 was considered statistically significant. All calculations were performed using IBM SPSS v.23 software package.
Results
Clinical responses to EGFR TKI therapy
All 30 included patients attended CT examination after 4 weeks of TKI treatment. 25 subjects demonstrated partial tumor response, 3 had stable disease and 2 progressed during the treatment (See Table 2 in the ESM). Twenty-nine patients continued TKI therapy (28 cases with the disease control (objective tumor response or stable disease) and 1 case beyond progression) after the first check-up. Twenty-five patients managed to attend the second CT examination, which was performed within 8–12 weeks after the beginning of the treatment; among 5 missing subjects, 3 patients underwent cytoreductive surgery, 1 subject refused examination due to COVID-19 epidemic precautions, and 1 patient died on the 6th week of treatment. The death of the patient occurred after sudden and rapid symptomatic deterioration; the cause of the death was unknown as the family of this subject refused an autopsy.
ctDNA analysis at base-line
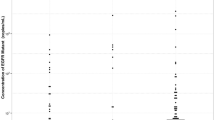
Thirty patients were subjected to the ctDNA analysis at base-line. EGFR-mutated DNA was detected in 25/30 (83%) subjects (Table 2, Fig. 1). As expected, the sum volume of the tumor lesions was evidently higher in patients with detectable mutated ctDNA level as compared with “plasma-negative” patients, but the difference did not reach the statistical significance (29,463 mm3 vs. 9963 mm3, p = 0.552, Mann–Whitney U test). The probability of detecting ctDNA at base-line did not correlate with the patient age or gender, number of metastatic sites or EGFR mutation type (Table 2). The first CT evaluation of tumor response at 4th week after the beginning of anti-EGFR therapy documented a trend towards more pronounced tumor volume decrease in the “ctDNA-positive” group as compared with “ctDNA-negative” patients (− 61% vs. − 18.5%, p = 0.208, Mann–Whitney U test). This tendency was not maintained after 8–12 weeks of treatment (Table 2). Patients with detectable EGFR-mutated DNA at base-line had shorter PFS than “ctDNA-negative” cases, however this difference was also below the level of statistical significance [11.4 months vs. 21.0 months, p = 0.238, Breslow (generalized Wilcoxon) test for comparison of Kaplan–Meier curves].
Changes of ctDNA concentration during the first hours of TKI exposure
None of 5 patients, who were negative for plasma EGFR-mutated DNA at base-line, showed the presence of ctDNA (at least 5 mutation-specific signals per reaction) in the serial samples, which were taken in the first hours after the beginning of the treatment. The remaining 25 subjects demonstrated some changes in the amount ctDNA (Table 3, Fig. 2, see also Fig. 1S in the ESM). One of these subjects, patient #Pt22, experienced in the 1st day of treatment the femur fracture at the site of the metastatic lesion; the trauma was accompanied by the increase of the concentration of EGFR mutation signals in the plasma; this patient was considered not informative for further analysis.
The analysis of changes in ctDNA concentrations occurring within the first 48 h of treatment revealed a few patterns (Table 3, Fig. 2, see also Fig. 1S in the ESM). Some patients demonstrated more or less consistent decline of the ctDNA content during first two days of therapy (#Pt7, #Pt8, #Pt10). A minority of cases showed a trend towards continuous increase in the number of circulating EGFR mutant copies (#Pt21, #Pt24). There were instances of relatively steady level of ctDNA over the first 48 h (#Pt23). The majority of patients showed less consistent variations in the ctDNA contents, with a number of spikes and drops (#Pt3, #Pt4, #Pt25, #Pt30, etc.).
We further attempted to correlate, which of the ctDNA measurements provides the best correlation with the tumor response and PFS. We have conditionally chosen 25% difference between the numbers of EGFR-mutated signals as a threshold. This cut-off was evaluated by the analysis of intrapatient differences in ctDNA concentration in 8 paired blood samples obtained at 0.5 h before the treatment start and at the time of TKI administration (see Table 4S in the ESM); none of the these pairs showed difference exceeding 25%. The samples were classified for 3 groups according to change of the ctDNA content between the base-line and a given time point; accordingly, there were groups with the increased, decreased and stable concentration of tumor DNA in plasma. The measurements made at 0.5, 1, 2, 3, 6, 12, 24 and 36 h did not produce statistically significant correlations with the disease outcome (p-values (Breslow test) for comparison of Kaplan–Meier curves in different subgroups: 0.714, 0.841, 0.206, 0.798, 0.255, 0.276, 0.161 and 0.737, respectively; p-values (Fisher exact test) for tumor response rates: 0.697, 0.697, 0.283, 0.657, 0.444, 0.978, 0.408 and 0.319, respectively). However, there were clinical correlations with the change of ctDNA level registered at 48 h after the start of the treatment (see Fig. 2S in the ESM).
Changes of ctDNA concentration at 48 h are predictive for TKI clinical efficacy
Twelve (50%) out of 24 informative patients showed > 25% reduction of the plasma ctDNA concentration (median decrease: − 85%; range: from − 100% to − 49%) at 48 h after the start of treatment. All these patients demonstrated disease control after 4 and 8–12 weeks of therapy (at 4 weeks: 11 PR and 1 SD; at 8–12 weeks: 8 PR and 2 SD; two patients underwent surgery and were not evaluable by RECIST) (Tables 2, 3). One of two patients, who underwent surgery between 1st and 2nd assessments, demonstrated complete pathologic tumor response.
The remaining 12 individuals showed either stable content of circulating EGFR-mutated DNA (n = 5) or the elevation of ctDNA concentration (n = 7) at 48 h after the start of the therapy (Table 2, 3). The median increase of the ctDNA level in the latter group was 95% (range: from 36 to 276%). 10 of 12 patients with elevated or stable ctDNA level achieved an objective response at 4 weeks, but only 5 of 10 evaluable patients still demonstrated disease control at 8–12 weeks of the treatment (Fisher exact test: p = 0.032, when compared to the group with ctDNA decrease). Progressive disease (PD) at weeks 8–12 was documented in 5/10 (50%) patients, who showed increased or stable ctDNA content at 48 h after the treatment; one additional patient died before the second assessment (Tables 2, 3).
The decline of concentration of EGFR-mutated DNA in plasma, which was observed at 48 h after the start of the TKI treatment, predicted for longer PFS as compared with patients with increasing or stable level of ctDNA (14.7 months vs. 8.5 months, p = 0.013, Kaplan–Meier method; Table 2; Fig. 3).
Discussion
EGFR TKIs are characterized by a relatively rapid absorption, with peak plasma concentrations achieved within a few hours after the uptake of the tablet [31, 32]. The therapeutic doses of the EGFR TKIs are significantly higher than the minimal drug concentrations, which exert some antitumor effect [33,34,35]. Consequently, it is explainable, that some patients experience evident symptomatic relief within the first hours after the beginning of the treatment [26, 27, 36].
Studies of EGFR inhibition in cell lines revealed, that the administration of anti-EGFR drugs results in immediate biological consequences. Exposure to TKI causes the decrease of EGFR autophosphorylation, followed by down-regulation of ERK, AKT, STAT3 and other signaling proteins; all these events are observed within 10–30 min after the addition of TKI to the cell culture medium [37,38,39]. Activation of the apoptotic signaling cascade can also be observed within first 10 min of TKI exposure [40, 41]. The reduction of the tumor mass upon EGFR TKI therapy is likely to be attributed both to the cessation of cell proliferation and to the induction of programmed cell death [17, 42, 43]. Some data indicate that immune-related mechanisms may also contribute to the tumor shrinkage [44, 45]. Use of MRI in animal experiments revealed evidence for tumor regression occurring already within 1–7 days after TKI administration [46,47,48]. Consequently, the mere fact of the existence of rapid plasma ctDNA response to TKI treatment is in agreement with preclinical observations.
The numerical data obtained in our study correspond well to the results observed in similarly designed investigations. In particular, we were able to detect EGFR mutations in plasma in 25/30 (83%) patients at base-line, which is close to the observations made in other studies [49,50,51,52,53]. Our data also validate previous findings, which suggest that the absence of detectable EGFR-mutated copies in the plasma at base-line correlates with improved PFS [13, 51, 54]. Similarly to the reports of Riediger et al. [55] and Phallen et al. [56], we observed temporary increase of the level of ctDNA during the first hours of treatment in some although not all cases. It is unclear whether these changes are attributed to the massive tumor cell shedding in response to the drug, or caused by other reasons.
Several studies monitored ctDNA concentration in the beginning of the treatment by EGFR inhibitors. Lee et al. [51] analyzed EGFR-mutated ctDNA at 8 weeks after the TKI administration, and observed the decline of its concentration in all 40 patients analyzed. Subjects with complete clearance of ctDNA at 8 weeks had significantly longer PFS as compared to patients with residual amounts of EGFR-mutated copies in the bloodstream. Molina-Vila et al. [54] examined 74 patients at 6 weeks after the start of therapy, and observed the presence of EGFR-mutated DNA only in 3 (4%) subjects. Monitoring of ctDNA at earlier time points is significantly more complicated. Riediger et al. [55] obtained serial blood samples at 1-day intervals during the first week of therapy from a patient receiving afatinib. They observed an increase of ctDNA concentration at 26 h, and subsequent continuous decline of the number of EGFR-mutated copies starting from 48 h time point. Husain et al. [57] monitored the amount of ex19del, L858R and T790M mutation in patients, who acquired resistance to erlotinib or afatinib, and started to receive osimertinib. Serial urinary samples were obtained at 1-day intervals during the first week of therapy. Some of these patients showed temporary spikes of urine tumor DNA concentration within days 1–7, however the subsequent observation revealed a consistent decline of the amount of EGFR-mutated DNA by the end of the second week. Our study has a novelty as compared to the mentioned above investigations, as it included a relatively large number of patients and involved a serial blood-takes performed within first hours after TKI administration. The justification of this effort was based on published observations describing a very rapid treatment response in a subset of patients [26, 27, 36] as well as on the data obtained in preclinical experiments [37,38,39]. Our results suggest that good responders to TKIs can be identified already at 48 h after the start of the EGFR-targeted therapy. Some NSCLCs treated by first- or second-generation EGFR TKIs demonstrate emergence of EGFR T790M mutations before clinical disease progression [58, 59]. However, treatment-naïve tumors usually do not contain EGFR T790M mutation as base-line [60] therefore our study considered only monitoring of ex19del- and L858R-mutated ctDNA.
This study has some limitations. Blood-take at 48 h after the beginning of the treatment was the latest time point in the early ctDNA response analysis. This was due to convenience reasons, as the patients started to receive TKI while been in a hospital, and the 2 days was a period between the first tablet and the hospital discharge. While 48 h was the only informative point for clinical prediction within the range 0.5–48 h, one could expect that the analysis of ctDNA at somewhat longer time intervals could have even better predictive value. The design of our study initially considered blood-takes at 2 and 4 weeks after the beginning of TKI treatment, however the compliance of patients was incomplete and the obtained data did not provide additional information (Table 3). It is also desirable to validate the obtained findings in larger studies involving serial blood-takes from NSCLCs patients undergoing EGFR TKI therapy.
Early monitoring of ctDNA after the start of treatment allows to evaluate whether the tumor will indeed consistently respond to TKI. It appears that although the majority of EGFR-mutated tumors demonstrate some initial disease control upon TKI administration, they can broadly be divided into two categories. In approximately a half of tumors the majority of cells constituting neoplastic lumps are vulnerable to TKI exposure, and these tumors demonstrate prolonged tumor response. Another half of EGFR-mutated NSCLCs is characterized by some intratumoral heterogeneity caused by compromised access to the drug for some tumor cells or by various in-built signaling mechanisms for TKI resistance. In these NSCLCs only a fraction of cells composing the tumor mass respond to TKI, while the remaining malignant clones facilitate rapid disease progression after initial short-term disease control.
Conclusions
The present study demonstrates that the clinical response to TKI can be predicted by the analysis of changes in the plasma ctDNA concentration at 48 h after the start of EGFR-targeted therapy. It is not obvious whether the results of this plasma test could call to some action. While the prediction for good response clearly supports the continuation of the treatment, it is unclear what options can be offered to potential poor responders. There are studies demonstrating promising results of the combined use of EGFR TKI inhibitors and antiangiogenic or cytotoxic drugs [61, 62]. Perhaps, lack of rapid response to a single-agent EGFR TKI may justify the addition of another antitumor compound to the front-line therapy. Current clinical trials often involve liquid biopsy; therefore testing of this concept is compatible with the available medical resources.
Availability of data and material
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.
Code availability
Not applicable.
Abbreviations
- CR:
-
Complete response;
- CT:
-
Computed tomography
- ctDNA:
-
Circulating tumor DNA
- DCR:
-
Disease control rate
- ddPCR:
-
Droplet digital PCR
- MRI:
-
Magnetic resonance imaging
- NSCLC:
-
Non-small cell lung cancer
- PD:
-
Progressive disease
- PR:
-
Partial response
- SD:
-
Stable disease
- PFS:
-
Progression free survival
- TKI:
-
Tyrosine kinase inhibitors
References
Bettegowda C, Sausen M, Leary RJ et al (2014) Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 6(224):224ra24. https://doi.org/10.1126/scitranslmed.3007094
Merker JD, Oxnard GR, Compton C et al (2018) Circulating tumor DNA analysis in patients with cancer: American society of clinical oncology and college of American pathologists joint review. J Clin Oncol 36(16):1631–1641. https://doi.org/10.1200/JCO.2017.76.8671
Gobbini E, Swalduz A, Levra MG et al (2020) Implementing ctDNA analysis in the clinic: challenges and opportunities in non-small cell lung cancer. Cancers (Basel) 12(11):3112. https://doi.org/10.3390/cancers12113112
Akhoundova D, Mosquera Martinez J et al (2020) The role of the liquid biopsy in decision-making for patients with non-small cell lung cancer. J Clin Med 9(11):3674. https://doi.org/10.3390/jcm9113674
Charo LM, Eskander RN, Okamura R et al (2021) Clinical implications of plasma circulating tumor DNA in gynecologic cancer patients. Mol Oncol 15(1):67–79. https://doi.org/10.1002/1878-0261.12791
Diehl F, Schmidt K, Choti MA et al (2008) Circulating mutant DNA to assess tumor dynamics. Nat Med 14(9):985–990. https://doi.org/10.1038/nm.1789
Tie J, Kinde I, Wang Y et al (2015) Circulating tumor DNA as an early marker of therapeutic response in patients with metastatic colorectal cancer. Ann Oncol 26(8):1715–1722. https://doi.org/10.1093/annonc/mdv177
Abbosh C, Birkbak NJ, Wilson GA et al (2017) Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545(7655):446–451. https://doi.org/10.1038/nature22364
Strijker M, Soer EC, de Pastena M et al (2020) Circulating tumor DNA quantity is related to tumor volume and both predict survival in metastatic pancreatic ductal adenocarcinoma. Int J Cancer 146(5):1445–1456. https://doi.org/10.1002/ijc.32586
Sorensen BS, Wu L, Wei W et al (2014) Monitoring of epidermal growth factor receptor tyrosine kinase inhibitor-sensitizing and resistance mutations in the plasma DNA of patients with advanced non-small cell lung cancer during treatment with erlotinib. Cancer 120(24):3896–3901. https://doi.org/10.1002/cncr.28964
Anagnostou V, Forde PM, White JR et al (2019) Dynamics of tumor and immune responses during immune checkpoint blockade in non-small cell lung cancer. Cancer Res 79(6):1214–1225. https://doi.org/10.1158/0008-5472.CAN-18-1127
Ebert EBF, McCulloch T, Hansen KH et al (2020) Clearing of circulating tumour DNA predicts clinical response to first line tyrosine kinase inhibitors in advanced epidermal growth factor receptor mutated non-small cell lung cancer. Lung Cancer 141:37–43. https://doi.org/10.1016/j.lungcan.2019.12.016
Fukuhara T, Saito H, Furuya N et al (2020) Evaluation of plasma EGFR mutation as an early predictor of response of erlotinib plus bevacizumab treatment in the NEJ026 study. EBioMedicine 57:102861. https://doi.org/10.1016/j.ebiom.2020.102861
Rixe O, Fojo T (2007) Is cell death a critical end point for anticancer therapies or is cytostasis sufficient? Clin Cancer Res 13(24):7280–7287. https://doi.org/10.1158/1078-0432.CCR-07-2141
Serkova NJ, Eckhardt SG (2016) Metabolic imaging to assess treatment response to cytotoxic and cytostatic agents. Front Oncol 6:152. https://doi.org/10.3389/fonc.2016.00152
Amaravadi RK, Thompson CB (2007) The roles of therapy-induced autophagy and necrosis in cancer treatment. Clin Cancer Res 13:7271–7279
Zhao ZQ, Yu ZY, Li J et al (2016) Gefitinib induces lung cancer cell autophagy and apoptosis via blockade of the PI3K/AKT/mTOR pathway. Oncol Lett 12(1):63–68. https://doi.org/10.3892/ol.2016.4606
Jiang MJ, Gu DN, Dai JJ et al (2020) Dark side of cytotoxic therapy: chemoradiation-induced cell death and tumor repopulation. Trends Cancer 6(5):419–431. https://doi.org/10.1016/j.trecan.2020.01.018
Osumi H, Shinozaki E, Takeda Y et al (2019) Clinical relevance of circulating tumor DNA assessed through deep sequencing in patients with metastatic colorectal cancer. Cancer Med 8(1):408–417. https://doi.org/10.1002/cam4.1913
Reece M, Saluja H, Hollington P et al (2019) The use of circulating tumor DNA to monitor and predict response to treatment in colorectal cancer. Front Genet 10:1118. https://doi.org/10.3389/fgene.2019.01118
Paez JG, Jänne PA, Lee JC et al (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304:1497–1500. https://doi.org/10.1126/science.1099314
Maemondo M, Inoue A, Kobayashi K et al (2010) Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med 362:2380–2388
Carotenuto P, Roma C, Rachiglio AM et al (2011) Optimizing response to gefitinib in the treatment of non-small-cell lung cancer. Pharmgenomics Pers Med 4:1–9. https://doi.org/10.2147/PGPM.S6626
Rosell R, Carcereny E, Gervais R et al (2012) Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 13(3):239–246. https://doi.org/10.1016/S1470-2045(11)70393-X
Sequist LV, Yang JC, Yamamoto N et al (2013) Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J Clin Oncol 31(27):3327–3334. https://doi.org/10.1200/JCO.2012.44.2806
Chien CR, Chen HJ (2016) Lazarus response to treatment of patients with lung cancer and oncogenic mutations in the intensive care unit. J Thorac Dis 8(11):E1455–E1461. https://doi.org/10.21037/jtd.2016.11.110
Conci N, Dalllio FG, Comellini V et al (2020) “Lazarus effect” in patient affected by lung adenocarcinoma carrying EGFR, CTNNB1, MET exon 11 and PIK3CA mutations treated with gefitinib. Precis Cancer Med 3:23. https://doi.org/10.21037/pcm-20-32
Moiseyenko VM, Procenko SA, Levchenko EV et al (2010) High efficacy of first-line gefitinib in non-Asian patients with EGFR-mutated lung adenocarcinoma. Onkologie 33:231–238. https://doi.org/10.1159/000302729
Eisenhauer EA, Therasse P, Bogaerts J et al (2009) New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45:228–247. https://doi.org/10.1016/j.ejca.2008.10.026
Sacher AG, Paweletz C, Dahlberg SE et al (2016) Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol 2(8):1014–1022. https://doi.org/10.1001/jamaoncol.2016.0173
Tiseo M, Bartolotti M, Gelsomino F et al (2010) Emerging role of gefitinib in the treatment of non-small-cell lung cancer (NSCLC). Drug Des Devel Ther 4:81–98. https://doi.org/10.2147/dddt.s6594
Nakamura Y, Sano K, Soda H et al (2010) Pharmacokinetics of gefitinib predicts antitumor activity for advanced non-small cell lung cancer. J Thorac Oncol 5:1404–1409
Satoh H, Inoue A, Kobayashi K et al (2011) Low-dose gefitinib treatment for patients with advanced non-small cell lung cancer harboring sensitive epidermal growth factor receptor mutations. J Thorac Oncol 6(8):1413–1417. https://doi.org/10.1097/JTO.0b013e31821d43a8
Hirano R, Uchino J, Ueno M et al (2016) Low-dose epidermal growth factor receptor (EGFR)—tyrosine kinase inhibition of EGFR mutation-positive lung cancer: therapeutic benefits and associations between dosage, efficacy and body surface area. Asian Pac J Cancer Prev 17(2):785–789. https://doi.org/10.7314/apjcp.2016.17.2.785
Cayssials E, Torregrosa-Diaz J, Gallego-Hernanz P et al (2020) Low-dose tyrosine kinase inhibitors before treatment discontinuation do not impair treatment-free remission in chronic myeloid leukemia patients: results of a retrospective study. Cancer 126(15):3438–3447. https://doi.org/10.1002/cncr.32940
Langer CJ (2009) The “lazarus response” in treatment-naive, poor performance status patients with non-small-cell lung cancer and epidermal growth factor receptor mutation. J Clin Oncol 27(9):1350–1354. https://doi.org/10.1200/JCO.2008.20.4859
Ono M, Hirata A, Kometani T et al (2004) Sensitivity to gefitinib (Iressa, ZD1839) in non-small cell lung cancer cell lines correlates with dependence on the epidermal growth factor (EGF) receptor/extracellular signal-regulated kinase 1/2 and EGF receptor/Akt pathway for proliferation. Mol Cancer Ther 3(4):465–472
Amann J, Kalyankrishna S, Massion PP et al (2005) Aberrant epidermal growth factor receptor signaling and enhanced sensitivity to EGFR inhibitors in lung cancer. Cancer Res 65(1):226–235
Wu K, Chang Q, Lu Y et al (2013) Gefitinib resistance resulted from STAT3-mediated Akt activation in lung cancer cells. Oncotarget 4(12):2430–2438. https://doi.org/10.18632/oncotarget.1431
Green DR (2005) Apoptotic pathways: ten minutes to dead. Cell 121(5):671–674. https://doi.org/10.1016/j.cell.2005.05.019
Cragg MS, Kuroda J, Puthalakath H et al (2007) Gefitinib-induced killing of NSCLC cell lines expressing mutant EGFR requires BIM and can be enhanced by BH3 mimetics. PLoS Med 4(10):1681–1689. https://doi.org/10.1371/journal.pmed.0040316 (discussion 1690)
Takeuchi K, Ito F (2010) EGF receptor in relation to tumor development: molecular basis of responsiveness of cancer cells to EGFR-targeting tyrosine kinase inhibitors. FEBS J 277(2):316–326
Fung C, Chen X, Grandis JR et al (2012) EGFR tyrosine kinase inhibition induces autophagy in cancer cells. Cancer Biol Ther 13(14):1417–1424. https://doi.org/10.4161/cbt.22002
Wang DH, Lee HS, Yoon D et al (2017) Progression of EGFR-mutant lung adenocarcinoma is driven by alveolar macrophages. Clin Cancer Res 23(3):778–788
Ayeni D, Miller B, Kuhlmann A et al (2019) Tumor regression mediated by oncogene withdrawal or erlotinib stimulates infiltration of inflammatory immune cells in EGFR mutant lung tumors. J Immunother Cancer 7(1):172. https://doi.org/10.1186/s40425-019-0643-8
Politi K, Zakowski MF, Fan PD et al (2006) Lung adenocarcinomas induced in mice by mutant EGF receptors found in human lung cancers respond to a tyrosine kinase inhibitor or to down-regulation of the receptors. Genes Dev 20(11):1496–1510. https://doi.org/10.1101/gad.1417406
Venugopalan A, Lee M, Niu G et al (2016) EGFR-targeted therapy results in dramatic early lung tumor regression accompanied by imaging response and immune infiltration in EGFR mutant transgenic mouse models. Oncotarget 7:54137–54156. https://doi.org/10.18632/oncotarget.11021
Jia Y, Li X, Jiang T et al (2019) EGFR-targeted therapy alters the tumor microenvironment in EGFR-driven lung tumors: implications for combination therapies. Int J Cancer 145(5):1432–1444. https://doi.org/10.1002/ijc.32191
Jing CW, Wang Z, Cao HX et al (2014) High resolution melting analysis for epidermal growth factor receptor mutations in formalin-fixed paraffin-embedded tissue and plasma free DNA from non-small cell lung cancer patients. Asian Pac J Cancer Prev 14(11):6619–6623. https://doi.org/10.7314/apjcp.2013.14.11.6619
Thress KS, Brant R, Carr TH et al (2015) EGFR mutation detection in ctDNA from NSCLC patient plasma: a cross-platform comparison of leading technologies to support the clinical development of AZD9291. Lung Cancer 90(3):509–515. https://doi.org/10.1016/j.lungcan.2015.10.004
Lee JY, Qing X, Xiumin W et al (2016) Longitudinal monitoring of EGFR mutations in plasma predicts outcomes of NSCLC patients treated with EGFR TKIs: Korean Lung Cancer Consortium (KLCC-12-02). Oncotarget 7(6):6984–6993. https://doi.org/10.18632/oncotarget.6874
Normanno N, Denis MG, Thress KS et al (2017) Guide to detecting epidermal growth factor receptor (EGFR) mutations in ctDNA of patients with advanced non-small-cell lung cancer. Oncotarget 8(7):12501–12516. https://doi.org/10.18632/oncotarget.13915
Kim JO, Shin JY, Kim SR et al (2020) Evaluation of two EGFR mutation tests on tumor and plasma from patients with non-small cell lung cancer. Cancers (Basel) 12(4):785. https://doi.org/10.3390/cancers12040785
Molina-Vila MA, Stahel RA, Dafni U et al (2020) Evolution and clinical impact of EGFR mutations in circulating free DNA in the BELIEF trial. J Thorac Oncol 15(3):416–425. https://doi.org/10.1016/j.jtho.2019.11.023
Riediger A, Dietz S, Schirmer U et al (2016) Mutation analysis of circulating plasma DNA to determine response to EGFR tyrosine kinase inhibitor therapy of lung adenocarcinoma patients. Sci Rep 6:33505. https://doi.org/10.1038/srep33505
Phallen J, Leal A, Woodward BD et al (2019) Early noninvasive detection of response to targeted therapy in non-small cell lung cancer. Cancer Res 79(6):1204–1213. https://doi.org/10.1158/0008-5472.CAN-18-1082
Husain H, Melnikova VO, Kosco K et al (2017) Monitoring daily dynamics of early tumor response to targeted therapy by detecting circulating tumor DNA in urine. Clin Cancer Res 23(16):4716–4723. https://doi.org/10.1158/1078-0432.CCR-17-0454
Tseng JS, Su KY, Yang TY et al (2016) The emergence of T790M mutation in EGFR-mutant lung adenocarcinoma patients having a history of acquired resistance to EGFR-TKI: focus on rebiopsy timing and long-term existence of T790M. Oncotarget 7(30):48059–48069. https://doi.org/10.18632/oncotarget.10351
Kawamura T, Kenmotsu H, Omori S et al (2018) Clinical factors predicting detection of T790M mutation in rebiopsy for EGFR-mutant non-small-cell lung cancer. Clin Lung Cancer 19(2):e247–e252. https://doi.org/10.1016/j.cllc.2017.07.002
Lavdovskaia ED, Iyevleva AG, Sokolenko AP et al (2018) EGFR T790M mutation in TKI-naïve clinical samples: frequency, tissue mosaicism, predictive value and awareness on artifacts. Oncol Res Treat 41(10):634–642. https://doi.org/10.1159/000491441
Yu HA, Schoenfeld AJ, Makhnin A et al (2020) Effect of osimertinib and bevacizumab on progression-free survival for patients with metastatic egfr-mutant lung cancers: a phase 1/2 single-group open-label trial. JAMA Oncol 6(7):1048–1054. https://doi.org/10.1001/jamaoncol.2020.1260
Nakagawa K, Garon EB, Seto T et al (2019) Ramucirumab plus erlotinib in patients with untreated, EGFR-mutated, advanced non-small-cell lung cancer (RELAY): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol 20(12):1655–1669. https://doi.org/10.1016/S1470-2045(19)30634-5
Funding
This study has been supported by the Russian Science Foundation (grant #18-75-10070).
Author information
Authors and Affiliations
Contributions
EK, FM and EI designed the study. The processing of biological materials was done by: KZ, AM. Data collection was performed by AZ, SB, ES, EA, TS, and NV. Molecular genetic analysis was carried out by: TL, TS, MK, and SC. Statistical analysis was performed by: EK, GY and EI. The first draft of the manuscript was written by FM, EK and EI. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The study design was approved by the local Ethical Committee of N.N. Petrov Institute of Oncology. All procedures performed in study were in accordance with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Moiseyenko, F.V., Kuligina, E.S., Zhabina, A.S. et al. Changes in the concentration of EGFR-mutated plasma DNA in the first hours of targeted therapy allow the prediction of tumor response in patients with EGFR-driven lung cancer. Int J Clin Oncol 27, 850–862 (2022). https://doi.org/10.1007/s10147-022-02128-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-022-02128-6