Abstract
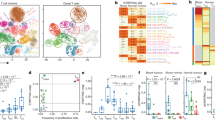
Colorectal cancer (CRC) remains a significant global health issue. In this study, the role of T-cell exhaustion-related genes (TEXs) in CRC was investigated using single-cell and bulk RNA-seq analysis. This research involved extensive data analysis using multiple databases, including the TCGA-COAD cohort, GSE14333, and GSE39582. Through single-cell analysis, distinct cell populations within CRC samples were identified and classified T-cells into four subgroups: regulatory T-cells (Tregs), conventional CD4+ T-cells (CD4+ T conv), CD8+ T, and CD8+ T exhausted cells. Intercellular communication networks and signaling pathways associated with TEXs using computational tools such as CellChat and PROGENy. Additionally, TEX-related alterations in tumor gene pathways were analyzed through Gene Ontology and Kyoto Encyclopedia of Genes and Genomes analyses. Prognostic models were developed, and their correlation with immune infiltration was assessed. The study revealed the presence of distinct cell populations within CRC, with TEXs playing a crucial role in the tumor microenvironment. CD8+ T exhausted cells exhibited expression of specific markers, indicating their involvement in tumor immune evasion. CellChat and PROGENy analyses revealed intricate communication networks and signaling pathways associated with TEXs, including RNA splicing and viral carcinogenesis. Furthermore, the prognostic risk model developed on the basis of TEXs demonstrated its efficacy in stratifying CRC patients. This risk model exhibited strong correlations with immune infiltration by various effector immune cells, highlighting the influence of TEXs on the tumor immune response. The complex interactions and signaling pathways underlying TEX-associated immune dysregulation in CRC were revealed by employing advanced analytical approaches. The development of a prognostic risk model based on TEXs offers a promising tool for prognostic stratification in patients with CRC. Furthermore, the correlations observed between TEXs and immune infiltration provide valuable insights into the potential of TEXs as therapeutic targets and highlight the need for further investigation into TEX-mediated immune evasion mechanisms. This study thus provides valuable insights into the role of TEXs in CRC.






Similar content being viewed by others
Data availability
The original contributions presented in the study are included in the article/supplementary material; further inquiries can be directed to the corresponding author.
References
Arneth B (2019) Tumor microenvironment. Medicina (Kaunas) 56(1):15
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium Nat Genet 25(1):25–29
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F et al (2016) Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 17(1):218
Benson AB, Venook AP, Al-Hawary MM, Cederquist L, Chen YJ, Ciombor KK et al (2018) NCCN guidelines insights: colon cancer, Version 2.2018. J Natl Compr Canc Netw 16(4):359–369
Brenner H, Kloor M, Pox CP (2014) Colorectal cancer. Lancet 383(9927):1490–1502
Cheng L, Xiong W, Li S, Wang G, Zhou J, Li H (2023) CRISPR-Cas9 screening identified lethal genes enriched in necroptosis pathway and of prognosis significance in osteosarcoma. J Gene Med:e3563
Collado M, Blasco MA, Serrano M (2007) Cellular senescence in cancer and aging. Cell 130(2):223–233
Darvin P, Toor SM, Sasidharan Nair V, Elkord E (2018) Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp Mol Med 50(12):1–11
Dolina JS, Van Braeckel-Budimir N, Thomas GD, Salek-Ardakani S (2021) CD8(+) T cell exhaustion in cancer. Front Immunol 12:715234
Huang H, Ke C, Zhang D, Wu J, Zhang P (2023) Molecular mechanism study and tumor heterogeneity of Chinese angelica and Fructus aurantii in the treatment of colorectal cancer through computational and molecular dynamics. Funct Integr Genomics 23(2):106
Jhunjhunwala S, Hammer C, Delamarre L (2021) Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat Rev Cancer 21(5):298–312
Jin S, Guerrero-Juarez CF, Zhang L, Chang I, Ramos R, Kuan CH et al (2021) Inference and analysis of cell-cell communication using CellChat. Nat Commun 12(1):1088
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Kobak D, Berens P (2019) The art of using t-SNE for single-cell transcriptomics. Nat Commun 10(1):5416
Li H, Yu L, Zhang X, Shang J, Duan X (2022) Exploring the molecular mechanisms and shared gene signatures between rheumatoid arthritis and diffuse large B cell lymphoma. Front Immunol 13:1036239
Li H, Zhang X, Shang J, Feng X, Yu L, Fan J et al (2023a) Identification of NETs-related biomarkers and molecular clusters in systemic lupus erythematosus. Front Immunol 14:1150828
Li H, Zhou J, Zhou L, Zhang X, Shang J, Feng X et al (2023b) Identification of the shared gene signatures and molecular pathways in systemic lupus erythematosus and diffuse large B-cell lymphoma. J Gene Med:e3558
Liu Y, Liu J, Tian Z, Zhang Z, Liu T, Chen C et al (2021) Highly expressed IFITM10 is associated with early diagnosis and T stage of gastric cancer. Transl Cancer Res 10(1):382–392
Luo G, Sun Y, Feng R, Zhao Q, Wen T (2018) ARL3 subcellular localization and its suspected role in autophagy. Biochimie 154:187–193
Morad G, Helmink BA, Sharma P, Wargo JA (2021) Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell 184(21):5309–5337
Powell L, Samarakoon YH, Ismail S, Sayer JA (2021) ARL3, a small GTPase with a functionally conserved role in primary cilia and immune synapses. Small GTPases 12(3):167–176
Rayess H, Wang MB, Srivatsan ES (2012) Cellular senescence and tumor suppressor gene p16. Int J Cancer 130(8):1715–1725
Rigatti SJ (2017) Random forest. J Insur Med 47(1):31–39
Schubert M, Klinger B, Klünemann M, Sieber A, Uhlitz F, Sauer S et al (2018) Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat Commun 9(1):20
Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S et al (2016) The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 54(1):1–30
Wherry EJ (2011) T cell exhaustion. Nat Immunol 12(6):492–499
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb) 2(3):100141
Xiao Y, Yu D (2021) Tumor microenvironment as a therapeutic target in cancer. Pharmacol Ther 221:107753
Xing X, Cai W, Shi H, Wang Y, Li M, Jiao J et al (2013) The prognostic value of CDKN2A hypermethylation in colorectal cancer: a meta-analysis. Br J Cancer 108(12):2542–2548
Zhang L, Liu C, Zhang X, Wang C, Liu D (2023) Breast cancer prognosis and immunological characteristics are predicted using the m6A/m5C/m1A/m7G-related long noncoding RNA signature. Funct Integr Genomics 23(2):117
Author information
Authors and Affiliations
Contributions
JW and HL designed the study. WT, YT, CT, HZ, SX, JW, LH, and LC performed data analysis. WT drafted the manuscript. JW and HL revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary materials

ESM 1
Figure S1. Prognostic stratification assessment of patients with CRC in the training and validation groups based on TEX risk scores. A-C. Kaplan-Meier curves, scatter plots, and dotted line plots for prognostic stratification of CRC in terms of TEX risk score in TCGA-COAD, GSE14333, and GSE39582 databases. CRC, colorectal cancer; TEX, T-cell exhaustion. (PNG 367 kb)

ESM 2
Figure S2. The Xcell method was applied to assess the immune infiltration of TEX risk scores in CRC. A. Correlation heat map evaluating the correlation of expression among multiple tumor immune cells identified by the Xcell method, B. Correlation heat map presenting the correlation of expression between multiple tumor immune cells identified by the Xcell method and the five core TEX-related prognostic genes constituting the TEX risk model, C. Box plot displaying the expression level of multiple tumor immune cells and immune pathways in the low-risk and high-risk groups, D. Scatter plot and linear correlation analysis revealing the association of risk scores with the expression of M1 macrophage, CD8 naive T-cells, Th2 T-cells, and mast cells. TEX, T-cell exhaustion, CRC, colorectal cancer. (PNG 730 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tu, W., Tu, Y., Tan, C. et al. Elucidating the role of T-cell exhaustion-related genes in colorectal cancer: a single-cell bioinformatics perspective. Funct Integr Genomics 23, 259 (2023). https://doi.org/10.1007/s10142-023-01188-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01188-9