Abstract
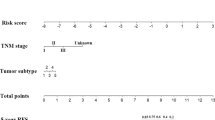
According to statistics, breast cancer (BC) has replaced lung cancer as the most common cancer in the world. Therefore, specific detection markers and therapeutic targets need to be explored as a way to improve the survival rate of BC patients. We first identified m6A/m5C/m1A/m7G-related long noncoding RNAs (MRlncRNAs) and developed a model of 16 MRlncRNAs. Kaplan–Meier survival analysis was applied to assess the prognostic power of the model, while univariate Cox analysis and multivariate Cox analysis were used to assess the prognostic value of the constructed model. Then, we constructed a nomogram to illustrate whether the predicted results were in good agreement with the actual outcomes. We tried to use the model to distinguish the difference in sensitivity to immunotherapy between the two groups and performed some analyses such as immune infiltration analysis, ssGSEA and IC50 prediction. To explore the novel anti-tumor drug response, we reclassified the patients into two clusters. Next, we assessed their response to clinical treatment by the R package pRRophetic, which is determined by the IC50 of each BC patient. We finally identified 11 MRlncRNAs and based on them, a risk model was constructed. In this model, we found good agreement between calibration plots and prognosis prediction. The AUC of ROC curves was 0.751, 0.734, and 0.769 for 1-year, 2-year, and 3-year overall survival (OS), respectively. The results showed that the IC50 was significantly different between the risk groups, suggesting that the risk groups can be used as a guide for systemic treatment. We regrouped patients into two clusters based on 11 MRlncRNAs expression. Next, we conducted immune scores for 2 clusters, which showed that cluster 1 had higher stromal scores, immune scores and higher estimated (microenvironment) scores, demonstrating that TME of cluster 1 was different from cluster 2. The results of this study support that MRlncRNAs can predict tumor prognosis and help differentiate patients with different sensitivities to immunotherapy as a basis for individualized treatment for BC patients.









Similar content being viewed by others
Data availability
The original contributions presented in the study are included in the article/Supplementary Material. Further inquiries can be directed to the corresponding author.
References
An Y, Duan H (2022) The role of m6A RNA methylation in cancer metabolism. Mol Cancer 21(1):14. https://doi.org/10.1186/s12943-022-01500-4
Barzaman K, Karami J, Zarei Z, Hosseinzadeh A, Kazemi MH, Moradi-Kalbolandi S et al (2020) Breast cancer: biology, biomarkers, and treatments. Int Immunopharmacol 84:106535. https://doi.org/10.1016/j.intimp.2020.106535
Bhan A, Soleimani M, Mandal SS (2017) Long noncoding RNA and cancer: a new paradigm. Cancer Res 77(15):3965–3981. https://doi.org/10.1158/0008-5472.Can-16-2634
Coughlin SS (2019) Epidemiology of breast cancer in women. Adv Exp Med Biol 1152:9–29. https://doi.org/10.1007/978-3-030-20301-6_2
Das S, Camphausen K, Shankavaram U (2020) Cancer-specific immune prognostic signature in solid tumors and its relation to immune checkpoint therapies. Cancers (Basel) 12(9):2476. https://doi.org/10.3390/cancers12092476
DeBerardinis RJ (2020) Tumor microenvironment, metabolism, and immunotherapy. N Engl J Med 382(9):869–871. https://doi.org/10.1056/NEJMcibr1914890
Denaro N, Merlano MC, Lo NC (2019) Long noncoding RNAs as regulators of cancer immunity. Mol Oncol 13(1):61–73. https://doi.org/10.1002/1878-0261.12413
Galván Morales MA, Barrera Rodríguez R, Santiago Cruz JR, Teran LM (2020) Overview of new treatments with immunotherapy for breast cancer and a proposal of a combination therapy. Molecules 25(23):5686. https://doi.org/10.3390/molecules25235686
Harjunpää H, Guillerey C (2020) TIGIT as an emerging immune checkpoint. Clin Exp Immunol 200(2):108–119. https://doi.org/10.1111/cei.13407
Huang W, Kong F, Li R, Chen X, Wang K (2022) Emerging Roles of m(6)A RNA Methylation regulators in gynecological cancer. Front Oncol 12:827956. https://doi.org/10.3389/fonc.2022.827956
Jang BS, Han W, Kim IA (2020) Tumor mutation burden, immune checkpoint crosstalk and radiosensitivity in single-cell RNA sequencing data of breast cancer. Radiother Oncol 142:202–09. https://doi.org/10.1016/j.radonc.2019.11.003
JingSong H, Hong G, Yang J, Duo Z, Li F, WeiCai C et al (2017) siRNA-mediated suppression of collagen type iv alpha 2 (COL4A2) mRNA inhibits triple-negative breast cancer cell proliferation and migration. Oncotarget 8(2):2585–2593. https://doi.org/10.18632/oncotarget.13716
Katsura C, Ogunmwonyi I, Kankam HK, Saha S (2022) Breast cancer: presentation, investigation and management. Br J Hosp Med (lond) 83(2):1–7. https://doi.org/10.12968/hmed.2021.0459
Li D, Li K, Zhang W, Yang KW, Mu DA, Jiang GJ et al (2022) The m6A/m5C/m1A regulated gene signature predicts the prognosis and correlates with the immune status of hepatocellular carcinoma. Front Immunol 13:918140. https://doi.org/10.3389/fimmu.2022.918140
Liu Q (2021) Current advances in N6-Methyladenosine methylation modification during bladder cancer. Front Genet 12:825109. https://doi.org/10.3389/fgene.2021.825109
Pan J, Huang Z, Xu Y (2021) m5C RNA Methylation regulators predict prognosis and regulate the immune microenvironment in lung squamous cell carcinoma. Front Oncol 11:657466. https://doi.org/10.3389/fonc.2021.657466
Shao D, Li Y, Wu J, Zhang B, Xie S, Zheng X et al (2022) An m6A/m5C/m1A/m7G-related long non-coding rna signature to predict prognosis and immune features of glioma. Front Genet 13:903117. https://doi.org/10.3389/fgene.2022.903117
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer Statistics, 2021. CA Cancer J Clin 71(1):7–33. https://doi.org/10.3322/caac.21654
Sun T, Wu R, Ming L (2019) The role of m6A RNA methylation in cancer. Biomed Pharmacother 112:108613. https://doi.org/10.1016/j.biopha.2019.108613
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A et al (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Tang K, Wu YH, Song Y, Yu B (2021) Indoleamine 2,3-dioxygenase 1 (IDO1) inhibitors in clinical trials for cancer immunotherapy. J Hematol Oncol 14(1):68. https://doi.org/10.1186/s13045-021-01080-8
Tong C, Wang W, He C (2022) m1A methylation modification patterns and metabolic characteristics in hepatocellular carcinoma. BMC Gastroenterol 22(1):93. https://doi.org/10.1186/s12876-022-02160-w
Wang X, Wang C, Guan J, Chen B, Xu L, Chen C (2021) Progress of Breast Cancer basic research in China. Int J Biol Sci 17(8):2069–2079. https://doi.org/10.7150/ijbs.60631
Wei JL, Wu SY, Yang YS, Xiao Y, Jin X, Xu XE et al (2021) GCH1 induces immunosuppression through metabolic reprogramming and IDO1 upregulation in triple-negative breast cancer. J Immunother Cancer 9(7):e002383. https://doi.org/10.1136/jitc-2021-002383
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26(12):1572–1573. https://doi.org/10.1093/bioinformatics/btq170
Wu M, Fu P, Qu L, Liu J, Lin A (2020) Long noncoding RNAs, new critical regulators in cancer immunity. Front Oncol 10:550987. https://doi.org/10.3389/fonc.2020.550987
Xu Y, Zhang M, Zhang Q, Yu X, Sun Z, He Y et al (2021) Role of main RNA methylation in hepatocellular carcinoma: N6-Methyladenosine, 5-Methylcytosine, and N1-Methyladenosine. Front Cell Dev Biol 9:767668. https://doi.org/10.3389/fcell.2021.767668
Yan J, Liu Z, Du S, Li J, Ma L, Li L (2020) Diagnosis and treatment of breast cancer in the precision medicine era. Methods Mol Biol 2204:53–61. https://doi.org/10.1007/978-1-0716-0904-0_5
Yu Z, Wang Y, Deng J, Liu D, Zhang L, Shao H et al (2021) Long non-coding RNA COL4A2-AS1 facilitates cell proliferation and glycolysis of colorectal cancer cells via miR-20b-5p/hypoxia inducible factor 1 alpha subunit axis. Bioengineered 12(1):6251–6263. https://doi.org/10.1080/21655979.2021.196983
Zhang J, Pang Y, Xie T, Zhu L (2019) CXCR4 antagonism in combination with IDO1 inhibition weakens immune suppression and inhibits tumor growth in mouse breast cancer bone metastases. Onco Targets Ther 12:4985–92. https://doi.org/10.2147/ott.S200643
Zhang Q, Gao C, Shao J, Wang Z (2021) TIGIT-related transcriptome profile and its association with tumor immune microenvironment in breast cancer. Biosci Rep 41(3). https://doi.org/10.1042/bsr20204340
Author information
Authors and Affiliations
Contributions
DL contributed to the study design and critical revision of the manuscript. LZ carried out the study and drafted the manuscript. LZ, CL, XZ, and CW analyzed the data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Fig. S1
The IC50 prediction of 24 commonly used anti-tumor drugs in risk groups. (PNG 1255 kb)

Fig. S2
The heat map, cumulative distribution function (CDF) plot, and the consensus CDF plots of consensus clustering matrix. (PNG 1052 kb)

Fig. S3
40 novel targeted drugs showing significant IC50 difference in 2 clusters. (PNG 1442 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Liu, C., Zhang, X. et al. Breast cancer prognosis and immunological characteristics are predicted using the m6A/m5C/m1A/m7G-related long noncoding RNA signature. Funct Integr Genomics 23, 117 (2023). https://doi.org/10.1007/s10142-023-01026-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01026-y