Abstract
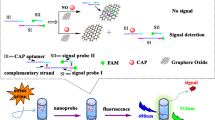
Gene detection has important applications in biology, biomedical engineering, clinical, environmental, and marine fields. Rapid, sensitive, and selective recognition of specific genes is essential in practical applications. In this study, we describe a facile and sensitive DNA sensing platform for specific and quantitative detection of Heterosigma akashiwo, which is one of the causative agents of red tides. Fast and sensitive detection is achieved by using chemically synthesized graphene oxide (GO) nanosheets. Probe DNA is designed according to the specific DNA fragments of harmful algae and labeled with fluorescent molecules FAM (fluorescein-based dye). GO nanosheet solution is made, in which the strong interaction between FAM-labeled probe and GO nanosheets keeps them in close proximity, facilitating the fluorescence quenching of the fluorophore by GO nanosheets. In the presence of a complementary target DNA, the FAM-labeled DNA probe and the target DNA hybridize and desorb from the surface of GO nanosheets, resulting in restoration of fluorophore fluorescence. The concentration of target DNA fragments is analyzed by the fluorescence intensity at ~ 520 nm with emission wavelength of 480 nm. The sensitive detecting platform achieved stable measurement of 1 pM specific genes from Heterosigma akashiwo. Our GO nanosheet-based DNA-sensing platform performs fast and sensitive detection of trace amount of DNA, and enables quantitative recognition of harmful algae, which has promising applications in real-time monitoring in the marine environment of red tide generative dynamics, allowing effective control, particularly in relation to marine aquaculture.







Similar content being viewed by others
References
Buskey EJ, Hyatt CJ (2006) Use of the FlowCAM for semi-automated recognition and enumeration of red tide cells ( Karenia brevis ) in natural plankton samples. Harmful Algae 5(6):685–692
Caracciolo AB, Dejana L, Fajardo C, Grenni P, Martin M, Mengs G, Sánchez-Fortún S, Lettieri T, Saccà ML, Medlin L (2019) A new fluorescent oligonucleotide probe for in-situ identification of Microcystis aeruginosa in freshwater. Microchem J 148:503–513
Chengke W, Yamin Z, Xiangmin M, Liansheng L (2011) A novel fluorescent biosensor for sequence-specific recognition of double-stranded DNA with the platform of graphene oxide. Analyst 136(10):2106–2110
Chou SS, Mrinmoy D, Jiayan L, Rotello VM, Jiaxing H, Dravid VP (2012) Nanoscale graphene oxide (nGO) as artificial receptors: implications for biomolecular interactions and sensing. J Am Chem Soc 134(40):16725–16733
Congcong Z, Mingxi C, Xiaoyang X, Li Z, Lei Z, Fengling X et al (2014) Graphene oxide reduced and modified by environmentally friendly glycylglycine and its excellent catalytic performance. Nanotechnology 25(13):135707
Eckford-Soper LK, Daugbjerg N (2015) Development of a multiplex real-time qPCR assay for simultaneous enumeration of up to four marine toxic bloom-forming microalgal species. Harmful Algae 48:37–43
Fan J, Yuan L, Liu Q, Tong C, Wang W, Xiao F, Liu B, Liu X (2019) An ultrasensitive and simple assay for the hepatitis C virus using a reduced graphene oxide-assisted hybridization chain reaction. Analyst 144(13):3972–3979
Ferrari AC, Robertson J (2000) Interpretation of Raman spectra of disordered and amorphous carbon. Phys Rev B 61(20):14095–14107
Fujii G, Imamura S, Hanaoka M, Tanaka K (2013) Nuclear-encoded chloroplast RNA polymerase sigma factor SIG2 activates chloroplast-encoded phycobilisome genes in a red alga, Cyanidioschyzon merolae. FEBS Lett 587(20):3354–3359
Glibert PM, Berdalet E, Burford MA, Pitcher GC, Zhou M (2018) Introduction to the Global Ecology and Oceanography of Harmful Algal Blooms (GEOHAB) synthesis. Global Ecology and Oceanography of Harmful Algal Blooms
Golubic S, Brent G, Lecampion T (1970) Scanning electron microscopy of endolithic algae and fungi using a multipurpose casting-embedding technique. Lethaia 3(2):203–209
Greenfield DI, Marin R, Jensen S, Massion E, Roman B, Feldman J, Scholin CA (2006) Application of Environmental Sample Processor (ESP) methodology for quantifying Pseudo-nitzschia australis using ribosomal RNA-targeted probes in sandwich and fluorescent in situ hybridization formats. Limnol Oceanogr Methods 4(3):426–435
Grondin PL, Ferland J, Karpboss L, Babin M (2016) High-frequency observations of phytoplankton spring bloom dynamics in Baffin Bay using imaging flow cytometry. Paper presented at the American Geophysical Union, Ocean Sciences Meeting, Abstract #HE34A-1460
Guillard RRL, Ryther JH (1962) Studies of marine planktonic diatoms: I. Cyclotella nana Hustedt, and Detonula confervacea (Cleve) Gran. Can J Microbiol 8(2):229–239
Hallegraeff GM (1993) A review of harmful algal blooms and their apparent global increase*. Phycologia 32(2):79–99
Hallegraeff GM, Anderson DM, Cembella AD, Enevoldsen H (1995) Manual on harmful marine microalgae. No.33 UNESCO:182–199
He S, Song B, Li D, Zhu C, Qi W, Wen Y, Wang L, Song S, Fang H, Fan C (2010) A graphene nanoprobe for rapid, sensitive, and multicolor fluorescent DNA analysis. Adv Funct Mater 20(3):453–459
Huang PJJ, Liu J (2012) DNA-length-dependent fluorescence signaling on graphene oxide surface. Small 8(7):977–983
Hummers WS Jr, Offeman RE (1958) Preparation of graphitic oxide. J Am Chem Soc 80(6):1339–1339
Jeffrey SW, Welschmeyer NA (1997) Spectrophotometric and fluorometric equations in common use in oceanography. Phytoplankton pigments in oceanography, pp 597–615
Kawai H, Nakamura S, Mimuro M, Furuya M, Watanabe M (1996) Microspectrofluorometry of the autofluorescent flagellum in phototactic brown algal zoids. Protoplasma 191(3–4):172–177
Kydd J, Rajakaruna H, Briski E, Bailey S (2018) Examination of a high resolution laser optical plankton counter and FlowCAM for measuring plankton concentration and size. J Sea Res 133:S1385110116302131
Lehrer S (1971) Solute perturbation of protein fluorescence. Quenching of the tryptophyl fluorescence of model compounds and of lysozyme by iodide ion. Biochemistry 10(17):3254–3263
Leqin TCTXH, Jinhui, Xiang JRXRW, Peimin YH (2010) Application and perspective of gene chip in marine microalgae research. Biotechnol Bull 11
Li D, Kaner RB (2008) Graphene-based materials. Science 320(5880):1170–1171
Li D, Müller MB, Gilje S, Kaner RB, Wallace GG (2008) Processable aqueous dispersions of graphene nanosheets. Nat Nanotechnol 3(2):101–105
Li F, Pei H, Wang L, Lu J, Gao J. Jiang B et al (2013) Nanomaterial-based fluorescent DNA analysis: a comparative study of the quenching effects of graphene oxide, carbon nanotubes, and gold nanoparticles. Adv Funct Mater 23(33):4140–4148
Li W, Jiang T, Pu Y, Jiao X, Tan W, Qin S (2019) Glucose biosensor using fluorescence quenching with chitosan-modified graphene oxide. Micro Nano Lett 14(3):344–348
Liu F, Choi JY, Seo TS (2010) Graphene oxide arrays for detecting specific DNA hybridization by fluorescence resonance energy transfer. Biosens Bioelectron 25(10):2361–2365
Liu M, Song J, Shuang S, Dong C, Brennan JD, Li Y (2014) A graphene-based biosensing platform based on the release of DNA probes and rolling circle amplification. ACS Nano 8(6):5564–5573
Liu C, Qiu S, Du P, Zhao H, Wang L (2018) Ionic liquid-graphene oxide hybrid nanomaterial: synthesis and anticorrosive applications. Nanoscale 10(17). https://doi.org/10.1039/C8NR01890A
Loh KP, Bao Q, Eda G, Chhowalla M (2010) Graphene oxide as a chemically tunable platform for optical applications. Nat Chem 2(12):1015–1024
Loo AH, Bonanni A, Pumera M (2012) Impedimetric thrombin aptasensor based on chemically modified graphenes. Nanoscale 4(1):143–147
Lu CH, Yang HH, Zhu CL, Chen X, Chen GN (2009) A graphene platform for sensing biomolecules. Angew Chem Int Ed 48(26):4785–4787
Lu C, Liu Y, Ying Y, Liu J (2017) Comparison of MoS2, WS2 and graphene oxide for DNA adsorption and sensing. Langmuir 33(2):630
Mendes CRB, Odebrecht C, Tavano VM, Abreu PC (2017) Pigment-based chemotaxonomy of phytoplankton in the Patos Lagoon estuary (Brazil) and adjacent coast. Mar Biol Res 13(1):22–35
Mirye P, Yun PS, Jinik H, Won JS, Juyun L, Man C, Taek-Kyun L (2018) Integration of the nuclease protection assay with sandwich hybridization (NPA-SH) for sensitive detection of Heterocapsa triquetra. Acta Oceanol Sin 37(5):107–112
Pehlivan ZS, Torabfam M, Kurt H, Ow-Yang C, Hildebrandt N, Yüce M (2019) Aptamer and nanomaterial based FRET biosensors: a review on recent advances (2014–2019). Microchim Acta 186(8):563
Peng Y, Zhigang Y, Chunmei D (2006) Pigment signatures of some diatoms isolated from China seas. Acta Oceanol Sin 25(1):108–118
Poulton NJ (2016) FlowCam: quantification and classification of phytoplankton by imaging flow cytometry. Methods Mol Biol 1389:237
Ravikumar CH, Gowda MI, Balakrishna RG (2019) An “OFF–ON” quantum dot–graphene oxide bioprobe for sensitive detection of micrococcal nuclease of Staphylococcus aureus. Analyst 144(13):3999–4005
Ris H, Singh RN (1961) Electron microscope studies on blue-green algae. J Biophys Biochem Cytol 9(1):63–80
Saldarriaga JF, Cavalier-Smith T, Menden-Deuer S, Keeling PJ (2004) Molecular data and the evolutionary history of dinoflagellates. Eur J Protistol 40(1):85–111
Simon N, Campbell L, Ornolfsdottir E, Groben R, Guillou L, Lange M, Medlin LK (2010) Oligonucleotide probes for the identification of three algal groups by dot blot and fluorescent whole-cell hybridization. J Eukaryot Microbiol 47(1):76–84
Singh DK, Iyer PK, Giri PK (2012) Role of molecular interactions and structural defects in the efficient fluorescence quenching by carbon nanotubes. Carbon 50(12):4495–4505
Singh DP, Herrera CE, Singh B, Singh S, Singh RK, Kumar R (2018) Graphene oxide: an efficient material and recent approach for biotechnological and biomedical applications. Mater Sci Eng C 86:173–197
Tamm M, Laas P, Freiberg R, Nõges P, Nõges T (2018) Parallel assessment of marine autotrophic picoplankton using flow cytometry and chemotaxonomy. Sci Total Environ 625:185–193
Wang Z, Zhang J, Chen P, Zhou X, Yang Y, Wu S et al (2011) Label-free, electrochemical detection of methicillin-resistant staphylococcus aureus DNA with reduced graphene oxide-modified electrodes. Biosens Bioelectron 26(9):3881–3886
Wietkamp S, Krock B, Gu H, Voß D, Klemm K, Tillmann U (2019) Occurrence and distribution of Amphidomataceae (Dinophyceae) in Danish coastal waters of the North Sea, the Limfjord and the Kattegat/Belt area. Harmful Algae 88:101637
Wietzorrek J, Stadler M, Kachel V (1994) Flow cytometric imaging-a novel tool for identification of marine organisms. Proceedings of OCEANS’94 (Vol 1, pp I-688). IEEE
Yamahara KM, Preston CM, Birch JM, Walz KR, Marin R III, Jensen S et al (2019) In-situ autonomous acquisition and preservation of marine environmental DNA using an autonomous underwater vehicle. Front Mar Sci 6:373
Zhang J, Liu S, Yang P, Sui G (2011) Rapid detection of algal toxins by microfluidic immunoassay. Lab Chip 11(20):3516–3522
Zhang Y, Chen X, Roozbahani GM, Guan X (2019) Rapid and sensitive detection of the activity of ADAM17 using a graphene oxide-based fluorescence sensor. Analyst 144(5):1825–1830
Zhen Y, Tiezhu MI, Zhigang YU (2008) Detection of Phaeocystis globosa using sandwich hybridization integrated with nuclease protection assay (NPA-SH). J Environ Sci 20(12):1481–1486
Zhen Y, Mi T, Yu Z (2009) Detection of several harmful algal species by sandwich hybridization integrated with a nuclease protection assay. Harmful Algae 8(5):651–657
Funding
This work was financially supported by the National Key Research and Development Project of China (2019YFC1407800), Natural Science Foundation for Distinguished Young Scientist of Shandong Province (Grant No. JQ201814), Natural Science Foundation for Young Scientists of China (Grant No.61701282), Qilu Young Scholar Funds (11500086963060), The Fundamental Research Funds of Shandong University (Grant No. 2017JC020 and 2017TB0020), the National Natural Science Foundation of China (41876134), and Collaborative Innovation Center of Technology and Equipment for Biological Diagnosis and Therapy in Universities of Shandong.
Author information
Authors and Affiliations
Contributions
Le Qiang: Investigation, validation, formal analysis, writing—original draft; Yu Zhang: conceptualization, supervision; Chao Wu: resources; Yingkuan Han: validation; Suchun Wang: resources; Yanyan Wang: resources; Congcong Zhang: resources; Guangzhou Liu: resources; Qi Wu: resources; Hong Liu: project administration; Ian R. Jenkinson: supervision, writing; Jun Sun: supervision, project administration; Lin Han: supervision, methodology, resources, funding acquisition.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
ESM 1
(DOCX 1787 kb)
Rights and permissions
About this article
Cite this article
Qiang, L., Zhang, Y., Wu, C. et al. A Facile and Sensitive DNA Sensing of Harmful Algal Blooms Based on Graphene Oxide Nanosheets. Mar Biotechnol 22, 498–510 (2020). https://doi.org/10.1007/s10126-020-09971-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-020-09971-6