Abstract
Aquatic organisms experience environmental hypoxia as a result of eutrophication and naturally occurring tidal cycles. Mytilus galloprovincialis, being an anoxic/hypoxic-tolerant bivalve, provides an excellent model to investigate the molecular mechanisms regulating oxygen sensing. Across the animal kingdom, inadequacy in oxygen supply is signalled predominantly by hypoxia-inducible factors (HIF) and Hif-prolyl hydroxylases (PHD). In this study, hif-α 5′-end and partial phd mRNA sequences from M. galloprovincialis were obtained. Phylogenetic and molecular characterization of both HIF-α and PHD putative proteins showed shared key features with the respective orthologues from animals strongly suggesting their crucial involvement in the highly conserved oxygen sensing pathway. Both transcripts displayed a tissue-specific distribution with prominent expression in gills. Quantitative gene expression analysis of hif-α and phd mRNAs from gills of M. galloprovincialis demonstrated that both these key sensors are transcriptionally modulated by oxygen availability during the short-time air exposure and subsequent re-oxygenation treatments proving that they are critical players of oxygen-sensing mechanisms in mussels. Remarkably, hif-α gene expression showed a prompt and transient response suggesting the precocious implication of this transcription factor in the early phase of the adaptive response to hypoxia in Mytilus. HIF-α and PHD proteins were modulated in a time-dependent manner with trends comparable to mRNA expression patterns, thus suggesting a central role of their transcriptional regulation in the hypoxia tolerance strategies in marine bivalves. These results provide molecular information about the effects of oxygen deficiency and identify hypoxia-responsive biomarker genes in mussels applicable in ecotoxicological studies of natural marine areas.






Similar content being viewed by others
References
Airaksinen S, Rabergh CMI, Sistonen L, Nikinmaa M (1998) Effects of heat shock and hypoxia on protein synthesis in rainbow trout (Oncorhynchus mykiss) cells. J Exp Biol 201:2543–2551
Anestis A, Portner HO, Michaelidis B (2010) Anaerobic metabolic patterns related to stress responses in hypoxia exposed mussels Mytilus galloprovincialis. J Exp Mar Biol Ecol 394:123–133
Aragones J, Fraisl P, Baes M, Carmeliet P (2009) Oxygen sensors at the crossroad of metabolism. Cell Metab 9:11–22
Babarro JMF, Labarta U, Reiriz MJF (2007) Energy metabolism and performance of Mytilus galloprovincialis under anaerobiosis. J Mar Biol Assoc UK 87:941–946
Benizri E, Ginouves A, Berra E (2008) The magic of the hypoxia-signaling cascade. Cell Mol Life Sci 65:1133–1149
Berra E, Benizri E, Ginouves A, Volmat V, Roux D, Pouyssegur J (2003) HIF prolyl-hydroxylase 2 is the key oxygen sensor setting low steady-state levels of HIF-1 alpha in normoxia. EMBO J 22:4082–4090
Brocato J, Chervona Y, Costa M (2014) Molecular responses to hypoxia-inducible factor 1 alpha and beyond. Mol Pharmacol 85:651–657
Brunelli E, Mauceri A, Salvatore F, Giannetto A, Maisano M, Tripepi S (2010) Localization of aquaporin 1 and 3 in the gills of the rainbow wrasse Coris julis. Acta Histochem 112:251–258
Bruning U, Cerone L, Neufeld Z, Fitzpatrick SF, Cheong A, Scholz CC, Simpson DA, Leonard MO, Tambuwala MM, Cummins EP, Taylor CT (2011) MicroRNA-155 promotes resolution of hypoxia-inducible factor 1 alpha activity during prolonged hypoxia. Mol Cell Biol 31:4087–4096
Campos C, Valente LMP, Borges P, Bizuayehu T, Fernandes JMO (2010) Dietary lipid levels have a remarkable impact on the expression of growth-related genes in Senegalese sole (Solea senegalensis Kaup). J Exp Biol 213:200–209
Chevreux B, Wetter T, Suhai S (1999) Genome sequence assembly using trace signals and additional sequence information computer science and biology. Computer Science and Biology: Proceedings of the German Conference on Bioinformatics (GCB) 99:45–56
D’Agata A, Cappello T, Maisano M, Parrino V, Giannetto A, Brundo MV, Ferrante M, Mauceri A (2014) Cellular biomarkers in the mussel Mytilus galloprovincialis (Bivalvia: Mytilidae) from Lake Faro (Sicily, Italy). Ital J Zool 81:43–54
David E, Tanguy A, Pichavant K, Moraga D (2005) Response of the Pacific oyster Crassostrea gigas to hypoxia exposure under experimental conditions. FEBS J 272:5635–5652
Diaz RJ, Rosenberg R (2008) Spreading dead zones and consequences for marine ecosystems. Science 321:926–929
Fasulo S, Iacono F, Cappello T, Corsaro C, Maisano M, D'Agata A, Giannetto A, De Domenico E, Parrino V, Lo Paro G, Mauceri A (2012) Metabolomic investigation of Mytilus galloprovincialis (Lamarck 1819) caged in aquatic environments. Ecotoxicol Environ Saf 84:139–146
Favaro E, Ramachandran A, McCormick R, Gee H, Blancher C, Crosby M, Devlin C, Blick C, Buffa F, Li JL, Vojnovic B, Pires das Neves R, Glazer P, Iborra F, Ivan M, Ragoussis J, Harris AL (2010) MicroRNA-210 regulates mitochondrial free radical response to hypoxia and krebs cycle in cancer cells by targeting iron sulfur cluster protein ISCU. PLoS One 5:e10345
Fernandes JMO, MacKenzie MG, Wright PA, Steele SL, Suzuki Y, Kinghorn JR, Johnston IA (2006) Myogenin in model pufferfish species: comparative genomic analysis and thermal plasticity of expression during early development. Comp Biochem Physiol Part D Genomics Proteomics 1:35–45
Fernandes JMO, Macqueen DJ, Lee HT, Johnston IA (2008) Genomic, evolutionary, and expression analyses of cee, an ancient gene involved in normal growth and development. Genomics 91:315–325
Fernandez CG, Roufidou C, Antonopoulou E, Sarropoulou E (2013) Expression of developmental-stage-specific genes in the gilthead sea bream Sparus aurata L. Mar Biotechnol 15:313–320
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer ELL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222–D230
Giannetto A, Nagasawa K, Fasulo S, Fernandes JMO (2013) Influence of photoperiod on expression of DNA (cytosine-5) methyltransferases in Atlantic cod. Gene 519:222–230
Giannetto A, Fernandes JMO, Nagasawa K, Mauceri A, Maisano M, De Domenico E, Cappello T, Oliva S, Fasulo S (2014) Influence of continuous light treatment on expression of stress biomarkers in Atlantic cod. Dev Comp Immunol 44:30–34
Ginouves A, Ilc K, Macias N, Pouyssegur J, Berra E (2008) PHDs overactivation during chronic hypoxia "desensitizes" HIF alpha and protects cells from necrosis. Proc Natl Acad Sci U S A 105:4745–4750
Gorr TA, Gassmann M, Wappner P (2006) Sensing and responding to hypoxia via HIF in model invertebrates. J Insect Physiol 52:349–364
Greenway SC, Storey KB (1999) The effect of prolonged anoxia on enzyme activities in oysters (Crassostrea virginica) at different seasons. J Exp Mar Biol Ecol 242:259–272
Guan L, Chi W, Xiao W, Chen L, He S (2014) Analysis of hypoxia-inducible factor alpha polyploidization reveals adaptation to Tibetan Plateau in the evolution of schizothoracine fish. BMC Evol Biol 14:192
Heise K, Puntarulo S, Nikinmaa M, Lucassen M, Portner HO, Abele D (2006) Oxidative stress and HIF-1 DNA binding during stressful cold exposure and recovery in the North Sea eelpout (Zoarces viviparus). Comp Biochem Physiol A Mol Integr Physiol 143:494–503
Henze AT, Riedel J, Diem T, Wenner J, Flamme I, Pouyseggur J, Plate KH, Acker T (2010) Prolyl Hydroxylases 2 and 3 act in gliomas as protective negative feedback regulators of hypoxia-inducible factors. Cancer Res 70:357–366
Ivan M, Harris AL, Martelli F, Kulshreshtha R (2008) Hypoxia response and microRNAs: no longer two separate worlds. J Cell Mol Med 12:1426–1431
Ivanina AV, Sokolov EP, Sokolova IM (2010) Effects of cadmium on anaerobic energy metabolism and mRNA expression during air exposure and recovery of an intertidal mollusk Crassostrea virginica. Aquat Toxicol 99:330–342
Kawabe S, Yokoyama Y (2012) Role of Hypoxia-Inducible Factor alpha in response to hypoxia and heat shock in the Pacific oyster Crassostrea gigas. Mar Biotechnol 14:106–119
Ke Q, Costa M (2006) Hypoxia-Inducible Factor-1 (HIF-1). Mol Pharmacol 70:1469–1480
Kurochkin IO, Ivanina AV, Eilers S, Downs CA, May LA, Sokolova IM (2009) Cadmium affects metabolic responses to prolonged anoxia and reoxygenation in eastern oysters (Crassostrea virginica). Am J Physiol Regul Integr Comp Physiol 297:1262–1272
Lahiri S, Roy A, Baby SM, Hoshi T, Semenza GL, Prabhakar NR (2006) Oxygen sensing in the body. Prog Biophys Mol Biol 91:249–286
Larade K, Storey KB (2009) Living without oxygen: anoxia-responsive gene expression and regulation. Curr Genomics 10:76–85
Law SHW, Wu RSS, Ng PKS, Yu RMK, Kong RYC (2006) Cloning and expression analysis of two distinct HIF-alpha isoforms-gcHIF-1alpha and gcHIF-4alpha from the hypoxia-tolerant grass carp, Ctenopharyngodon idellus. BMC Mol Biol 7:15
Lazado CC, Kumaratunga HPS, Nagasawa K, Babiak I, Giannetto A, Fernandes JMO (2014) Daily rhythmicity of clock gene transcripts in Atlantic cod fast skeletal muscle. PLoS One 9:e99172
Li T, Brouwer M (2007) Hypoxia-inducible factor, gsHIF, of the grass shrimp Palaemonetes pugio: molecular characterization and response to hypoxia. Comp Biochem Physiol B Biochem Mol Biol 147:11–19
Liu CC, Shin PKS, Cheung SG (2014) Isolation and mRNA expression of hypoxia-inducible factor α (HIF-α) in two sublittoral nassariid gastropods: Nassarius siquijorensis and Nassarius conoidalis. Mar Environ Res 99:44–51
Marchler-Bauer A, Lu SN, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke ZX, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang DC, Zhang NG, Zheng CJ, Bryant SH (2011) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229
Minamishima YA, Moslehi J, Padera RF, Bronson RT, Liao R, Kaelin WG (2009) A feedback loop involving the Phd3 Prolyl Hydroxylase tunes the mammalian hypoxic response in vivo. Mol Cell Biol 29:5729–5741
Myllyharju J, Koivunen P (2013) Hypoxia-inducible factor prolyl 4-hydroxylases: common and specific roles. Biol Chem 394:435–448
Nagasawa K, Giannetto A, Fernandes JMO (2012) Photoperiod influences growth and mll (Mixed-Lineage Leukaemia) expression in Atlantic cod. PLoS One 7:e36908
Nicastro KR, Zardi GI, McQuaid CD, Stephens L, Radloff S, Blatch GL (2010) The role of gaping behaviour in habitat partitioning between coexisting intertidal mussels. BMC Ecol 10:17
Nikinmaa M, Leveelahti L, Dahl E, Rissanen E, Rytkonen KT, Laurila A (2008) Population origin, development and temperature of development affect the amounts of HSP70, HSP90 and the putative hypoxia-inducible factor in the tadpoles of the common frog Rana temporaria. J Exp Biol 211:1999–2004
Piano A, Valbonesi P, Fabbri E (2004) Expression of cytoprotective proteins, heat shock protein 70 and metallothioneins, in tissues of Ostrea edulis exposed to heat and heavy metals. Cell Stress Chaperones 9:134–142
Piontkivska H, Chung JS, Ivanina AV, Sokolov EP, Techa S, Sokolova IM (2011) Molecular characterization and mRNA expression of two key enzymes of hypoxia-sensing pathways in eastern oysters Crassostrea virginica (Gmelin): Hypoxia-inducible factor alpha (HIF-alpha) and HIF-prolyl hydroxylase (PHD). Comp Biochem Physiol Part D Genomics Proteomics 6:103–114
Rane S, He M, Sayed D, Vashistha H, Malhotra A, Sadoshima J, Vatner DE, Vatner SF, Abdellatif M (2009) Downregulation of miR-199a derepresses hypoxia-inducible factor-1alpha and Sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ Res 104:879–886
Rissanen E, Tranberg HK, Nikinmaa M (2006) Oxygen availability regulates metabolism and gene expression in trout hepatocyte cultures. Am J Physiol Regul Integr Comp Physiol 291:R1507–R1515
Rytkonen KT, Ryynanen HJ, Nikinmaa M, Primmer CR (2008) Variable patterns in the molecular evolution of the hypoxia-inducible factor-1 alpha (HIF-1 alpha) gene in teleost fishes and mammals. Gene 420:1–10
Rytkonen KT, Williams TA, Renshaw GM, Primmer CR, Nikinmaa M (2011) Molecular evolution of the metazoan PHD-HIF oxygen-sensing system. Mol Biol Evol 28:1913–1926
Salamov AA, Nishikawa T, Swindells MB (1998) Assessing protein coding region integrity in cDNA sequencing projects. Bioinformatics 14:384–390
Sciortino MT, Medici MA, Marino-Merlo F, Zaccaria D, Giuffre-Cuculletto M, Venuti A, Grelli S, Mastino A (2008) Involvement of HVEM receptor in activation of nuclear factor kappa B by herpes simplex virus 1 glycoprotein D. Cell Microbiol 10:2297–2311
Semenza GL (2004) Hydroxylation of HIF-1: oxygen sensing at the molecular level. Physiology 19:176–182
Shen C, Nettleton D, Jiang M, Kim SK, Powell-Coffman JA (2005) Roles of the HIF-1 hypoxia-inducible factor during hypoxia response in Caenorhabditis elegans. J Biol Chem 280:20580–20588
Sollid J, Rissanen E, Tranberg HK, Thorstensen T, Vuori KA, Nikinmaa M, Nilsson GE (2006) HIF-1alpha and iNOS levels in crucian carp gills during hypoxia-induced transformation. J Comp Physiol B 176:359–369
Sonanez-Organis JG, Peregrino-Uriarte AB, Gomez-Jimenez S, Lopez-Zavala A, Forman HJ, Yepiz-Plascencia G (2009) Molecular characterization of hypoxia inducible factor-1 (HIF-1) from the white shrimp Litopenaeus vannamei and tissue-specific expression under hypoxia. Comp Biochem Physiol C Toxicol Pharmacol 150:395–405
Stiehl DP, Wirthner R, Koditz J, Spielmann P, Camenisch G, Wenger RH (2006) Increased prolyl 4-hydroxylase domain proteins compensate for decreased oxygen levels-evidence for an autoregulatory oxygen-sensing system. J Biol Chem 281:23482–23491
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony methods. Mol Biol Evol 28:2731–2739
Taylor CT, Pouyssegur J (2007) Oxygen, hypoxia, and stress. Ann N Y Acad Sci 1113:87–94
Terova G, Rimoldi S, Cora S, Bernardini G, Gornati R, Saroglia M (2008) Acute and chronic hypoxia affects HIF-1 alpha mRNA levels in sea bass (Dicentrarchus labrax). Aquaculture 279:150–159
Velázquez-Amado RM, Escamilla-Chimal EG, Fanjul-Moles ML (2012) Daily light-dark cycles influence hypoxia-inducible factor-1 and heat shock protein levels in the pacemakers of cray-fish. Photochem Photobiol 88:81–89
Wang DP, Li HG, Li YJ, Guo SC, Yang J, Qi DL, Jin C, Zhao XQ (2006) Hypoxia-inducible factor 1alpha cDNA cloning and its mRNA and protein tissue specific expression in domestic yak (Bos grunniens) from Qinghai-Tibetan plateau. Biochem Biophys Res Commun 348:310–319
Wang G, Yu Z, Zhen Y, Mi T, Shi Y, Wang J, Wang M, Sun S (2014) Molecular characterisation, evolution and expression of hypoxia-inducible factor in Aurelia sp.1. PLoS One 9:e100057
Webster KA (2003) Evolution of the coordinate regulation of glycolytic enzyme genes by hypoxia. J Exp Biol 206:2911–2922
Woo S, Jeon HY, Kim SR, Yum S (2011) Differentially displayed genes with oxygen depletion stress and transcriptional responses in the marine mussel, Mytilus galloprovincialis. Comp Biochem Physiol Part D Genomics Proteomics 6:348–356
Yao CL, Somero GN (2013) Thermal stress and cellular signaling processes in hemocytes of native (Mytilus californianus) and invasive (M. galloprovincialis) mussels: Cell cycle regulation and DNA repair. Comp Biochem Physiol A Mol Integr Physiol 165:159–168
Zhao TB, Ning HX, Zhu SS, Sun P, Xu SX, Chang ZJ, Zhao XQ (2004) Cloning of hypoxia-inducible factor 1 alpha cDNA from a high hypoxia tolerant mammal-plateau pika (Ochotona curzoniae). Biochem Bioph Res Commun 316:565–572
Acknowledgments
This research was supported by a National Interest Research Project, PRIN 2010–2011 (prot. 2010ARBLT7_001/008).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supp. 1
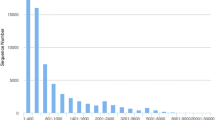
Hits from BLAST search against nucleotide collection (BLASTN) using the 308 bp hif-α cDNA (GenBank accession number JN595864) (a), the 761 bp hif-α cDNA (GenBank accession number KP185351) (b) and phd cDNA (GenBank accession number KP185352) (c) as queries. Score, query cover, E value, identity and accession numbers are showed (GIF 280 kb)
Supp. 2
Minimum Evolution (ME) phylogeny analysis of the HIF-α proteins based on the Dayhoff distance. The phylogenetic tree was constructed by MEGA.4 using the ME method. The bootstrap values from 1000 replicates are given at each branch node and only values higher than 50 % are shown. Abbreviations and sequence accession numbers are shown in Table 2 (GIF 20 kb)
Supp. 3
Minimum Evolution (ME) phylogeny analysis of the PHD proteins based on the Dayhoff distance. The phylogenetic tree was constructed by MEGA.4 using the ME method. The bootstrap values from 1000 replicates are given at each branch node and only values higher than 50 % are shown. Abbreviations and sequence accession numbers are shown in Table 2 (GIF 21 kb)
Supp. 4
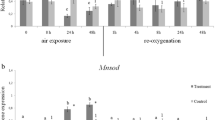
Expression level of HIF-α and PHD proteins analyzed with the image analysis Alliance LD2 87WL software (UVITEC Ltd, Cambridge, UK). Data are presented as Optical density (mean ± SD, n = 6). All densitometry values were normalized to the endogenous Actin protein. Different letters indicate significant differences between time points in the treatment group, different numbers refer to significant differences between time points in the control group. Stars represent significant differences between treatment and control group at the same time point (P < 0.05) (GIF 28 kb)
Rights and permissions
About this article
Cite this article
Giannetto, A., Maisano, M., Cappello, T. et al. Hypoxia-Inducible Factor α and Hif-prolyl Hydroxylase Characterization and Gene Expression in Short-Time Air-Exposed Mytilus galloprovincialis . Mar Biotechnol 17, 768–781 (2015). https://doi.org/10.1007/s10126-015-9655-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-015-9655-7