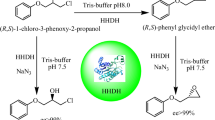
Abstract
The recombinant l-haloacid dehalogenase from the marine bacterium Psychromonas ingrahamii has been cloned and over-expressed in Escherichia coli. It shows activity towards monobromoacetic (100 %), monochloroacetic acid (62 %), S-chloropropionic acid (42 %), S-bromopropionic acid (31 %), dichloroacetic acid (28 %) and 2-chlorobutyric acid (10 %), respectively. The l-haloacid dehalogenase has highest activity towards substrates with shorter carbon chain lengths (≤C3), without preference towards a chlorine or bromine at the α-carbon position. Despite being isolated from a psychrophilic bacterium, the enzyme has mesophilic properties with an optimal temperature for activity of 45 °C. It retains above 70 % of its activity after being incubated at 65 °C for 90 min before being assayed at 25 °C. The enzyme is relatively stable in organic solvents as demonstrated by activity and thermal shift analysis. The V max and K m were calculated to be 0.6 μM min−1 mg−1 and 1.36 mM with monobromoacetic acid, respectively. This solvent-resistant and stable l-haloacid dehalogenase from P. ingrahamii has potential to be used as a biocatalyst in industrial processes.








Similar content being viewed by others
Abbreviations
- l-HAD:
-
l-Haloacid dehalogenase
- PinHAD:
-
Psychromonas ingrahamii l-haloacid dehalogenase
References
Breezee J, Cady N, Staley JT (2004) Subfreezing growth of the sea ice bacterium “Psychromonas ingrahamii” Microb. Ecol 47:300–304
Brinboim H, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Burroughs AM, Allen KN, Dunaway-Mariano D, Aravind L (2006) Evolutionary genomics of the HAD superfamily: understanding the structural adaptations and catalytic diversity in a superfamily of phosphoesterases and allied enzymes. J Mol Biol 361:1003–1034
Cabrita MT, Vale C, Rauter AP (2010) Halogenated compounds from marine algae. Marine Drugs 8:2301–2317
Cavicchioli R, Siddiqui KS, Andrew D, Sowers KR (2002) Low-temperature extremophiles and their applications. Curr Opin Biotechnol 13:253–261
Collins T, Meuwis MA, Gerday C, Feller G (2003) Activity, stability and flexibility in glycosidases adapted to extreme thermal environments. J Mol Biol 328:419–428
D'Amico S, Marx JC, Gerday C, Feller G (2003) Activity-stability relationship in extremophilic enzymes. J Biol Chem 278:7891–7896
Daniel RM, Danson MJ (2010) A new understanding of how temperature affects the catalytic activity of enzymes. Trends Biochem Sci 35:584–591
DeLano WL (2002) The PyMOL Molecular Graphics System, DeLano Scientific, Palo Alto, CA.
Feller G (2003) Molecular adaptations to cold in psychrophilic enzymes. Cell Mol Life Sci 60:648–662
Field JA, Verhagen FJM, De Jong E (1995) Natural organohalogen production by Basidiomycetes. Trends Biotechnol 13:451–456
Georlette D, Damiens B, Blaise V, Depiereux E, Uversky VN, Gerday C, Feller G (2003) Structural and functional adaptions to extreme temperatures in psychrophilic, mesophilic and thermophilic DNA ligases. J Biol Chem 278:37015–37023
Georlette D, Blaise V, Collins T, Gratia E, D'Amico S, Hoyoux A, Marx JC, Sonan G, Feller G, Gerday C (2004) Some like it cold: biocatalysis at low temperatures. FEMS Microbiol Rev 28:25–42
Gerike U, Danson MJ, Russell NJ, Hough DW (1997) Sequencing and expression of the gene encoding a cold active citrate synthase from an Antarctic bacterium, strain DS2 3R. Eur J Biochem 248:49–57
Gerike U, Danson MJ, Hough DW (2001) Cold–active citrate synthase: mutagenesis of active-site residues. Prot Eng 14:655–661
Gomes J, Steiner W (2004) The biocatalytic potential of extremophiles and extremozymes. Food Technol Biotechnol 42:223–235
Gouet P, Courcelle E, Stuart DI, Metoz F (1999) ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15:305–308
Gribble GW (1996) The diversity of natural organochlorines in living organisms. Pure Appl Chem 68:1699–1712
Gribble GW (1999) The diversity of naturally occurring organobromine compounds. Chem Soc Rev 28:335–346
Haki GD, Rakshit SK (2003) Developments in industrially important thermostable enzymes: a review. Bioresour Technol 89:17–34
Hasan AK, Takada H, Esaki N, Soda K (1991) Catalytic action of l-2-haloacid dehalogenase on long chain l-2-haloalkanoic acids in organic solvents. Biotechnol Bioeng 38:1114–1117
Holloway P, Trevors JT, Lee H (1998) A colorimetric assay for detecting haloalkane dehalogenase activity. J Microbiol Methods 32:31–36
Inatomi K, Ohba M, Oshima T (1983) Chemical properties of proteinaceous cell wall from an acido-thermophile, Sulfolobus acidocaldarius. Chemistry Letters:1191–1194
Kurata K, Taniguchi K, Agatsuma Y, Suzuki M (1998) Diterpenoid feeding-deterrents from Laurencia saitoi. Phytochemistry 47:363–369
Kurihara T, Liu JQ, Nardi-Dei V, Koshikawa H, Esaki N, Soda K (1995) Comprehensive site-directed mutagenesis of L-2-haloacid dehalogenase to probe catalytic amino acid residues. J Biochem 117:1317–1322
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Leatherbarrow R J, Horley U K (2007) GraFit Version 6, Erithacus Software Ltd.
Lee CK, Daniel RM, Shepherd C, Saul D, Cary SC, Danson MJ, Eisenthal R, Peterson ME (2007) Eurythermalism and temperature dependence of enzyme activity. FASEB J 8:1934–1941
Li YF (1999) Global technical hexachlorocyclohexane usage and its contamination consequences in the environment: from 1948 to 1997. Sci Total Environ 232:121–158
Liu JQ, Kurihara T, Hasan AK, Nardi-Dei V, Koshikawa H, Esaki N, Soda K (1994) Purification and characterization of thermostable and nonthermostable 2-haloacid dehalogenases with different stereospecificities from Pseudomonas sp. strain YL. Appl Environ Microbiol 60:2389–2393
Liu JQ, Kurihara T, Ichiyama S, Tsunasawa S, Miyagi, Kawasaki H, Soda K, Esaki (1998) Reaction mechanism of fluoroacetate dehalogenase from Moraxella sp. B J Biol Chem 273:30897–30902
MOE 2011.10, Chemical Computing Group Inc., www.chemcomp.com
Nagata Y, Endo R, Ito M, Ohtsubo Y, Tsuda M (2007) Aerobic degradation of lindane (hexachlorocyclohexane) in bacteria and its biochemical and molecular basis. Appl Microbiol Biotechnol 76:741–752
Nightingale PD, Malin G, Liss PS (1995) Production of chloroform and other low-molecular-weight halocarbons by some species of macroalgae. Limnol Oceanogr 40:680–689
Oikawa T, Kazuoka T, Soda K (2003) Paradoxical thermostable enzymes from psychrophile: molecular characterization and potentiality for biotechnological application. J Mol Catal B: Enzymatic 23:65–70
Paul NA, De Nys R, Steinberg PD (2006) Chemical defence against bacteria in the red alga Asparagopsis armata: linking structure with function. Mar Ecol Prog Ser 306:87–101
Riley M, Staley JT, Danchin A, Wang TZ, Brettin TS, Hauser LJ, Land ML, Thompson LS (2008) Genomics of an extreme psychrophile, Psychromonas ingrahamii. BMC Genomics 9:210–239
Russell RJ, Gerike U, Danson MJ, Hough DW, Taylor GL (1998) Structural adaptations of the cold-active citrate synthase from an Antarctic bacterium. Structure 6:351–361
Rye CA, Isupov MN, Lebedev AA, Littlechild JA (2009) Biochemical and structural studies of a l-haloacid dehalogenase from the thermophilic archaeon Sulfolobus tokodaii. Extremophiles 13:179–190
Schmidberger JW, Wilce JA, Tsang JS, Wilce MC (2007) Crystal structures of the substrate free-enzyme, and reaction intermediate of the HAD superfamily member, haloacid dehalogenase DehIVa from Burkholderia cepacia MBA4. J Mol Biol 368:706–717
Schoemaker HE, Mink D, Wubbolts MG (2003) Dispelling the myths-biocatalysis in industrial synthesis. Science 299:1694–1697
Scott C, Pandey G, Hartley CJ, Jackson CJ, Cheesman MJ, Taylor MC, Pandey R, Khurana JL, Teese M, Coppin CW, Weir K, Jain RK, Lal R, Russell RJ, Oakeshott JG (2008) The enzymatic basis for pesticide bioremediation. Ind J Microbiol 48:65–79
Siddiqui KS, Feller G, D'Amico S, Gerday C, Giaquinto L, Cavicchioli R (2005) The active site is the least stable structure in the unfolding pathway of a multidomain cold-adapted alpha-amylase. J Bacteriol 187:6197–6205
Siddiqui KS, Cavicchioli R (2006) Cold-adapted enzymes. Annu Rev Biochem 75:403–433
Staley JT, Gosink JJ (1999) Poles apart: biodiversity and biogeography of sea ice bacteria. Annu Rev Microbiol 53:189–215
Struvay C, Feller G (2012) Optimization to low temperature activity in psychrophilic enzyme. Int J Mol Sci 13:11643–11665
Trincone A (2011) Marine Biocatalysts: Enzymatic Features and Applications. Marine Drugs 9:478–499
Tsang JSH, Sam L (1999) Cloning and characterization of a cryptic haloacid dehalogenase from Burkholderia cepacia MBA4. J Bacteriol 181:6003–6009
Valverde C, Orozco A, Becerra A, Jeziorszi MC, Villalobos P, Solis JC (2004) Halometabolites and cellular dehalogenase systems: an evolutionary perspective. Int Rev Cytol 234:143–199
Van der Ploeg J, Van Hall G, Janssen DB (1991) Characterization of the haloacid dehalogenase from Xanthobacter autotrophicus GJ10 and sequencing of the dhlB gene. J Bacteriol 173:7925–7933
Acknowledgments
The Biotechnology and Biological Science Research Council, UK and Aquapharm Biodiscovery, Oban are acknowledged for PhD studentship funding (HRN). We thank Dr. Dorothee Gotz and Dr. Andrew Mearns Spragg for useful discussion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Novak, H.R., Sayer, C., Panning, J. et al. Characterisation of an l-Haloacid Dehalogenase from the Marine Psychrophile Psychromonas ingrahamii with Potential Industrial Application. Mar Biotechnol 15, 695–705 (2013). https://doi.org/10.1007/s10126-013-9522-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-013-9522-3