Abstract
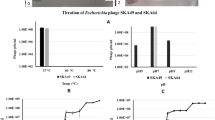
Avian pathogenic Escherichia coli (APEC) is the causative agent of avian colibacillosis, which causes significant economic losses to the poultry industry. The growing resistance of bacteria to antibiotics is a major global public health concern. However, there is limited data on the efficacy of phage therapy in effectively controlling and treating APEC infections. In this study, a novel lytic Escherichia phage, vB_EcoS_PJ16, was isolated from poultry farm wastewater and characterized in both in vitro and in vivo conditions. Transmission electron microscopy analysis revealed the presence of an icosahedral head and a long non-contractile tail, classifying the phage under the Caudoviricetes class. Host range determination showed that Escherichia phage vB_EcoS_PJ16 exhibited lytic activity against multiple strains of pathogenic E. coli, while no significant signs of lysis for Klebsiella pneumoniae, Salmonella Typhimurium, Listeria monocytogenes, and Staphylococcus aureus. Biophysical characterization revealed that the isolated phage was sturdy, as it remained viable for up to 300 days at temperatures of 30 °C, 37 °C, and 42 °C and for up to 24 h at pH 5 to 11, with only minor changes in titer. Kinetic analysis at multiplicity of infection (MOI) 0.1 showed a latency period of about 20 min and a burst size of 26.5 phage particles per infected cell for phage vB_EcoS_PJ16. Whole genome sequencing unveiled that the phage vB_EcoS_PJ16 genome consists of a double-stranded linear DNA molecule with 57,756 bp and a GC content of 43.58%. The Escherichia phage vB_EcoS_PJ16 genome consisted of 98 predicted putative ORFs, with no transfer RNA identified in the genome. Among these 98 genes, 34 genes were predicted to have known functions. A significant reduction in APEC viability was observed at MOI 100 during in vitro bacterial challenge tests conducted at different MOIs (0.01, 1, and 100). In vivo oral evaluation of the isolated phage to limit E. coli infections in day-old chicks indicated a decrease in mortality within both the therapeutic (20%) and prophylactic (30%) groups, when compared to the control group. The findings of this study contribute to our current knowledge of Escherichia phages and suggest a potentially effective role of phages in the therapeutic and prophylactic control of antibiotic-resistant APEC strains.






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this article. The complete genome sequence of novel Escherichia phage vB_EcoS_PJ16 is available in NCBI GenBank with accession number OQ955832.
References
Abdelrahman F, Rezk N, Fayez MS, Abdelmoteleb M, Atteya R, Elhadidy M, El-Shibiny A (2022) Isolation, characterization, and genomic analysis of three novel E. coli bacteriophages that effectively infect E. coli O18. Microorganisms 10(3):589. https://doi.org/10.3390/microorganisms10030589
Abedon ST, Herschler TD, Stopar D (2001) Bacteriophage latent-period evolution as a response to resource availability. Appl Environ Microbiol 67(9):4233–4241. https://doi.org/10.1128/AEM.67.9.4233-4241.2001
Adriaenssens E, Brister J (2017) How to name and classify your phage: an informal guide. Viruses 9:70. https://doi.org/10.3390/v9040070
Ahmadi H, Radford D, Kropinski AM, Lim LT, Balamurugan S (2017) Thermal-stability and reconstitution ability of Listeria phages P100 and A511. Front Microbiol 8:2375. https://doi.org/10.3389/fmicb.2017.02375
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Amarillas L, Rubi-Rangel L, Chaidez C, Gonzalez-Robles A, Lightbourn-Rojas L, Leon-Felix J (2017) Isolation and characterization of phiLLS, a novel phage with potential biocontrol agent against multidrug-resistant Escherichia coli. Front Microbiol 8:1355. https://doi.org/10.3389/fmicb.2017.01355
Bauer AW, Kirby WM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45(4):493–496. https://doi.org/10.1093/ajcp/45.4_ts.493
Becker B, de la Fuente N, Gassel M, Gunther D, Tavares P, Lurz R, Trautner TA, Alonso JC (1997) Head morphogenesis genes of the Bacillus subtilis bacteriophage SPP1. J Mol Biol 268(5):822–839. https://doi.org/10.1006/jmbi.1997.0997
Biggs KRH, Bailes CL, Scott L, Wichman HA, Schwartz EJ (2021) Ecological approach to understanding superinfection inhibition in bacteriophage. Viruses 13(7):1389. https://doi.org/10.3390/v13071389
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Buchfink B, Klaus R, Hajk-Georg D (2021) Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods 18(4):366–368. https://doi.org/10.1038/s41592-021-01101-x
Catalao MJ, Gil F, Moniz-Pereira J, Sao-Jose C, Pimentel M (2013) Diversity in bacterial lysis systems: bacteriophages show the way. FEMS Microbiol Rev 37(4):554–571. https://doi.org/10.1111/1574-6976.12006
Chakraborti S, Balakrishnan D, Trotter AJ, Gittens WH, Yang AWH, Jolma A, Paterson JR, Swiatek S, Plewka J, Curtis FA, Bowers LY, Palsson LO, Hughes TR, Taube M, Kozak M, Heddle JG, Sharples GJ (2020) A bacteriophage mimic of the bacterial nucleoid-associated protein. Fis Biochem J 477(7):1345–1362. https://doi.org/10.1042/BCJ20200146
Chen M, Xu J, Yao H, Lu C, Zhang W (2016) Isolation, genome sequencing and functional analysis of two T7-like coliphages of avian pathogenic Escherichia coli. Gene 582(1):47–58. https://doi.org/10.1016/j.gene.2016.01.049
Chibani-Chennoufi S, Sidoti J, Bruttin A, Kutter E, Sarker S, Brüssow H (2004) In vitro and in vivo bacteriolytic activities of Escherichia coli phages: implications for phage therapy. Antimicrob Agents Chemother 48(7):2558–2569. https://doi.org/10.1128/AAC.48.7.2558-2569.2004
Cuervo A, Pulido-Cid M, Chagoyen M, Arranz R, Gonzalez-Garcia VA, Garcia-Doval C, Caston JR, Valpuesta JM, van Raaij MJ, Martín-Benito J, Carrascosa JL (2013) Structural characterization of the bacteriophage T7 tail machinery. J Biol Chem 288(36):26290–26299. https://doi.org/10.1074/jbc.M113.491209
Cuervo A, Fabrega-Ferrer M, Machon C, Conesa JJ, Fernandez FJ, Perez-Luque R, Perez-Ruiz M, Pous J, Vega MC, Carrascosa JL, Coll M (2019) Structures of T7 bacteriophage portal and tail suggest a viral DNA retention and ejection mechanism. Nat Commun 10(1):3746. https://doi.org/10.1038/s41467-019-11705-9
Donlan RM (2009) Preventing biofilms of clinically relevant organisms using bacteriophage. Trends Microbiol 17(2):66–72. https://doi.org/10.1016/j.tim.2008.11.002
Doyle MP, Erickson MC (2006) Reducing the carriage of foodborne pathogens in livestock and poultry. Poult Sci 85:960–973. https://doi.org/10.1093/ps/85.6.960
Edgar R, Friedman N, Molshanski-Mor S, Qimron U (2012) Reversing bacterial resistance to antibiotics by phage-mediated delivery of dominant sensitive genes. Appl Environ Microbiol 8(3):744–751. https://doi.org/10.1128/AEM.05741-11
Elbreki M, Ross RP, Hill C, O’Mahony J, McAuliffe O, Coffey A (2014) Bacteriophages and their derivatives as biotherapeutic agents in disease prevention and treatment. J Viruses 2014:382539. https://doi.org/10.1155/2014/382539
Feng YY, Ong SL, Hu JY, Tan XL, Ng WJ (2003) Effects of pH and temperature on the survival of coliphages MS2 and Qbeta. J Ind Microbiol Biotechnol 30(9):549–552. https://doi.org/10.1007/s10295-003-0080-y
Filho RLA, Higgins JP, Higgins SE, Gaona G, Wolfenden AD, Tellez G, Hargis BM (2007) Ability of bacteriophages isolated from different sources to reduce Salmonella enterica serovar enteritidis in vitro and in vivo. Poult Sci 86(9):1904–1909. https://doi.org/10.1093/ps/86.9.1904
Fong K, Wong CWY, Wang S, Delaquis P (2021) How broad is enough: the host range of bacteriophages and its impact on the agri-food sector. PHAGE 2(2):83–91. https://doi.org/10.1089/phage.2020.0036
Fu W, Forster T, Mayer O, Curtin JJ, Lehman SM, Donlan RM (2010) Bacteriophage cocktail for the prevention of biofilm formation by Pseudomonas aeruginosa on catheters in an in vitro model system. Antimicrob Agents Chemother 54(1):397–404. https://doi.org/10.1128/AAC.00669-09
Gill JJ (2015) Phage genomic DNA extraction. https://openwetware.org/mediawiki/index.php?title=Gill:Phage_genomic_DNA_extraction&oldid=882952. Accessed Aug 2022
Guizelini D, Raittz RT, Cruz LM, Souza EM, Steffens MB, Pedrosa FO (2016) GFinisher: a new strategy to refine and finish bacterial genome assemblies. Sci Rep 6(1):34963. https://doi.org/10.1038/srep34963
Hietala V, Horsma-Heikkinen J, Carron A, Skurnik M, Kiljunen S (2019) The removal of endo- and enterotoxins from bacteriophage preparations. Front Microbiol 10:1674. https://doi.org/10.3389/fmicb.2019.01674
Huang G, Le S, Peng Y, Zhao Y, Yin S, Zhang L, Yao X, Tan Y, Li M, Hu F (2013) Characterization and genome sequencing of phage Abp1, a new phiKMV-like virus infecting multidrug-resistant Acinetobacter baumannii. Curr Microbiol 66(6):535–543. https://doi.org/10.1007/s00284-013-0308-7
Huff WE, Huff GR, Rath NC, Balog JM, Xie H, Moore PA Jr, Donoghue AM (2002) Prevention of Escherichia coli respiratory infection in broiler chickens with bacteriophage (SPR02). Poult Sci 81(4):437–441. https://doi.org/10.1093/ps/81.4.437
Huff WE, Huff GR, Rath NC, Balog JM, Donoghue AM (2005) Alternatives to antibiotics: utilization of bacteriophage to treat colibacillosis and prevent foodborne pathogens. Poult Sci 84(4):655–659. https://doi.org/10.1093/ps/84.4.655
ICTV (2011) The international committee on taxonomy of viruses. https://ictv.global/report_9th. Accessed Aug 2022
Isidro A, Henriques AO, Tavares P (2004) The portal protein plays essential roles at different steps of the SPP1 DNA packaging process. Virology 322(2):253–263. https://doi.org/10.1016/j.virol.2004.02.012
Jepson CD, March JB (2003) Bacteriophage lambda is a highly stable DNA vaccine delivery vehicle. Vaccine 22(19):2413–2419. https://doi.org/10.1016/j.vaccine.2003.11.065
Jhandai P, Kumar A, Vaishali GR (2019) Detection of Escherichia coli from food of animal origin in Hisar District of Haryana. Har Vet 58(2):245–247
Johnson TJ, Wannemuehler Y, Doetkott C, Johnson SJ, Rosenberger SC, Nolan LK (2008) Identification of minimal predictors of avian pathogenic Escherichia coli virulence for use as a rapid diagnostic tool. JCM 46(12):3987–3996. https://doi.org/10.1128/JCM.00816-08
Jonczyk E, Klak M, Miedzybrodzki R, Gorski A (2011) The influence of external factors on bacteriophages–review. Folia Microbiol 56(3):191–200. https://doi.org/10.1007/s12223-011-0039-8
Kabir SML (2010) Avian colibacillosis and salmonellosis, a closer look at epidemiology, pathogenesis, diagnosis, control and public health concerns. Int J Environ Res Public Health 7(1):89–114. https://doi.org/10.3390/ijerph7010089
Kathayat D, Lokesh D, Ranjit S, Rajashekara G (2021) Avian pathogenic Escherichia coli (APEC): an overview of virulence and pathogenesis factors, zoonotic potential, and control strategies. Pathogens 10(4):467. https://doi.org/10.3390/pathogens10040467
Kazibwe G, Katami P, Alinaitwe R, Alafi S, Nanteza A, Nakavuma JL (2020) Bacteriophage activity against and characterisation of avian pathogenic Escherichia coli isolated from colibacillosis cases in Uganda. PLoS One 15(12):e0239107. https://doi.org/10.1371/journal.pone.0239107
Kinch LN, Ginalski K, Rychlewski L, Grishin NV (2005) Identification of novel restriction endonuclease-like fold families among hypothetical proteins. Nucleic Acids Res 33(11):3598–3605. https://doi.org/10.1093/nar/gki676
Kong X, Wang H, Guo G, Li P, Tong P, Liu M, Ma X, Dong C, Li Y, Zhang H, Zhang W (2022) Duck sewage source coliphage P762 can lyse STEC and APEC. Virus Genes 58(5):436–447. https://doi.org/10.1007/s11262-022-01915-7
Kongari R, Rajaure M, Cahill J, Rasche E, Mijalis E, Berry J, Young R (2018) Phage spanins: diversity, topological dynamics and gene convergence. BMC Bioinform 19(1):326. https://doi.org/10.1186/s12859-018-2342-8
Kropinski AM, Mazzocco A, Waddell TE, Lingohr E, Johnson RP (2009) Enumeration of bacteriophages by double agar overlay plaque assay. Methods Mol Biol 501:69–76. https://doi.org/10.1007/978-1-60327-164-6_7
Kumar RN, Bhima B, Kumar PU, Ghosh S (2020) Bio-control of Salmonella spp. in carrot salad and raw chicken skin using lytic bacteriophages. LWT 122:109039. https://doi.org/10.1016/j.lwt.2020.109039
Lau GL, Sieo CC, Tan WS, Hair-Bejo M, Jalila A, Ho YW (2010) Efficacy of a bacteriophage isolated from chickens as a therapeutic agent for colibacillosis in broiler chickens. Poult Sci 89(12):2589–2596. https://doi.org/10.3382/ps.2010-00904
Levin BR, Bull JJ (2004) Population and evolutionary dynamics of phage therapy. Nat Rev Microbiol 2(2):166–173. https://doi.org/10.1038/nrmicro822
Li H, Richard D (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li P, Wang H, Li M, Qi W, Qi Z, Chen W, Dong Y, Xu Z, Zhang W (2022) Characterization and genome analysis of a broad lytic spectrum bacteriophage P479 against multidrug-resistant Escherichia coli. Virus Res 308(2022):198628. https://doi.org/10.1016/j.virusres.2021.198628
Lopes A, Amarir-Bouhram J, Faure G, Petit MA, Guerois R (2010) Detection of novel recombinases in bacteriophage genomes unveils Rad52, Rad51 and Gp2.5 remote homologs. Nucleic Acids Res. 38(12):3952–3962. https://doi.org/10.1093/nar/gkq096
Lu TK, Collins JJ (2007) Dispersing biofilms with engineered enzymatic bacteriophage. Proceedings Nat Acad Sci, USA 104(27):11197–11202. https://doi.org/10.1073/pnas.0704624104
Lu TK, Collins JJ (2009) Engineered bacteriophage targeting gene networks as adjuvants for antibiotic therapy. Proceed Nat Acad Sci, USA 106:4629–4634. https://doi.org/10.1073/pnas.0800442106
Lusiak-Szelachowska M, Zaczek M, Weber-Dąbrowska B, Miedzybrodzki R, Klak M, Fortuna W, Letkiewicz S, Rogoz P, Szufnarowski K, Jonczyk-Matysiak E, Owczarek B, Gorski A (2014) Phage neutralization by sera of patients receiving phage therapy. Viral Immunol 27(6):295–304. https://doi.org/10.1089/vim.2013.0128
Mahichi F, Synnott AJ, Yamamichi K, Osada T, Tanji Y (2009) Site-specific recombination of T2 phage using IP008 long tail fiber genes provides a targeted method for expanding host range while retaining lytic activity. FEMS Microbiol Lett 2:211–217. https://doi.org/10.1111/j.1574-6968.2009.01588.x
Mapes AC, Trautner BW, Liao KS, Ramig RF (2016) Development of expanded host range phage active on biofilms of multidrug-resistant Pseudomonas aeruginosa. Bacteriophage 6:e1096995. https://doi.org/10.1080/21597081.2015.1096995
Mozaffari P, Berizi E, Hosseinzadeh S, Derakhshan Z, Taghadosi V, Montaseri Z, Gotz F (2022) Isolation and characterization of E. coli O157:H7 novel bacteriophage for controlling this food-borne pathogen. Virus Res 315:198754. https://doi.org/10.1016/j.virusres.2022.198754
Newase S, Kapadnis BP, Shashidhar R (2019) Isolation and genome sequence characterization of bacteriophage vB_SalM_PM10, a Cba120virus, concurrently infecting Salmonella enterica serovars Typhimurium, Typhi, and Enteritidis. Curr Microbiol 76(1):86–94. https://doi.org/10.1007/s00284-018-1588-8
Ngu NT, Loc HT, Nhan NTH, Huan PKN, Anh LH, Xuan NH (2020) Isolation and characterization of bacteriophages against Escherichia coli isolates from chicken farms. Adv Anim Vet Sci 8(2):161–166. https://doi.org/10.17582/journal.aavs/2020/8.2.161.166
Niu YD, Stanford K, Kropinski AM, Ackermann HW, Johnson RP, She YM, Ahmed R, Villegas A, McAllister TA (2012) Genomic, proteomic and physiological characterization of a T5-like bacteriophage for control of Shiga toxin-producing Escherichia coli O157:H7. PLoS One 7(4):e34585. https://doi.org/10.1371/journal.pone.0034585
Nobrega F, Costa A, Santos J, Silikus M, VanLent J, Kengen S, Azeredo J, Kluskens L (2016) Genetically manipulated phages with improved pH resistance for oral administration in veterinary medicine. Sci Rep 6:39235. https://doi.org/10.1038/srep39235
Oliveira A, Sereno R, Azeredo J (2010) In vivo efficiency evaluation of a phage cocktail in controlling severe colibacillosis in confined conditions and experimental poultry houses. Vet Microbiol 146(3–4):303–308. https://doi.org/10.1016/j.vetmic.2010.05.015
Ortega ME, Catalano CE (2006) Bacteriophage lambda gpNu1 and Escherichia coli IHF proteins cooperatively bind and bend viral DNA: implications for the assembly of a genome-packaging motor. Biochem 45(16):5180–5189. https://doi.org/10.1021/bi052284b
Pereira C, Moreirinha C, Lewicka M, Almeida P, Clemente C, Romalde JL, Nunes ML, Almeida A (2017) Characterization and in vitro evaluation of new bacteriophages for the biocontrol of Escherichia coli. Virus Res 227:171–182. https://doi.org/10.1016/j.virusres.2016.09.019
Pires DP, Vilas Boas D, Sillankorva S, Azeredo J (2015) Phage therapy: a step forward in the treatment of Pseudomonas aeruginosa infections. J Virol 89(15):7449–7456. https://doi.org/10.1128/JVI.00385-15
Principi N, Silvestri E, Esposito S (2019) Advantages and limitations of bacteriophages for the treatment of bacterial infections. Front Pharmacol 10:513. https://doi.org/10.3389/fphar.2019.00513
Ryan DP, Owen-Hughes T (2011) Snf2-family proteins: chromatin remodellers for any occasion. Curr Opin Chem Biol 15(5):649–656. https://doi.org/10.1016/j.cbpa.2011.07.022
Salmond G, Fineran P (2015) A century of the phage: past, present and future. Nat Rev Microbiol 13:777–786. https://doi.org/10.1038/nrmicro3564
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Sjahriani T, Wasito EB, Tyasningsih W (2021) Isolation and identification of Escherichia coli O157:H7 lytic bacteriophage from environment sewage. Int J Food Sci 11:20217383121. https://doi.org/10.1155/2021/7383121
Soleimani-Delfan A, Bouzari M, Wang R (2021) vB_EfaS-DELF1, a novel Siphoviridae bacteriophage with highly effective lytic activity against vancomycin-resistant Enterococcus faecalis. Virus Res 298:198391. https://doi.org/10.1016/j.virusres.2021.198391
Sorensen AN, Woudstra C, Sorensen MCH, Brondsted L (2021) Subtypes of tail spike proteins predicts the host range of Ackermannviridae phages. Comput Struct Biotechnol J 19:4854–4867. https://doi.org/10.1016/j.csbj.2021.08.030
Strack HB, Freese EB, Freese E (1964) Comparison of mutation and inactivation rates induces by bacteriophage and transforming DNA by various mutagenes. Mutat Res 1(1):10–21. https://doi.org/10.1016/0027-5107(64)90048-X
Sulakvelidze A, Alavidze Z, Morris JG (2001) Bacteriophage therapy. Antimicrob Agents Chemother 45:649–659. https://doi.org/10.1128/AAC.45.3.649-659.2001
Tang F, Zhang P, Zhang Q, Xue F, Ren J, Sun J, Qu Z, Zhuge X, Li D, Wang J, Jiang M, Dai J (2019) Isolation and characterization of a broad-spectrum phage of multiple drug resistant Salmonella and its therapeutic utility in mice. Microb Pathog 126:193–198. https://doi.org/10.1016/j.micpath.2018.10.042
Vasu K, Nagaraja V (2013) Diverse functions of restriction-modification systems in addition to cellular defense. Microbiol Mol Biol Rev 77(1):53–72. https://doi.org/10.1128/MMBR.00044-12
Vinga I, Droge A, Stiege AC, Lurz R, Santos MA, Daugelavicius R, Tavares P (2006) The minor capsid protein gp7 of bacteriophage SPP1 is required for efficient infection of Bacillus subtilis. Mol Microbiol 61(6):1609–1621. https://doi.org/10.1111/j.1365-2958.2006.05327.x
Wang Z, Zheng P, Ji W, Fu Q, Wang H, Yan Y, Sun J (2016) SLPW: a virulent bacteriophage targeting methicillin-resistant Staphylococcus aureus in vitro and in vivo. Front Microbiol 7:934. https://doi.org/10.3389/fmicb.2016.00934
Wang Z, Zhang F, Liang Y, Zheng K, Gu C, Zhang W, Liu Y, Zhang X, Shao H, Jiang Y, Guo C, He H, Wang H, Sung YY, Mok WJ, Wong LL, He J, McMinn A, Wang M (2021) Genome and ecology of a novel Alteromonas podovirus, ZP6, representing a new viral genus. Mareflavirus. Microbiol Spectr. 9(2):e0046321. https://doi.org/10.1128/Spectrum.00463-21
Wang Z, Zheng X, Guo G, Hu Z, Miao J, Dong Y, Xu Z, Zhou Q, Wei X, Han X, Liu Y, Zhang W (2022) O145 may be emerging as a predominant serogroup of avian pathogenic Escherichia coli (APEC) in China. Vet Microbiol. 266:109358. https://doi.org/10.1016/j.vetmic.2022.109358
Xu J, Chen M, He L, Zhang S, Ding T, Yao H, Lu C, Zhang W (2016) Isolation and characterization of a T4-like phage with a relatively wide host range within Escherichia coli. J Basic Microbiol 56(4):405–421. https://doi.org/10.1002/jobm.201500440
Yazdi M, Bouzari M, Ghaemi EA, Shahin K (2020) Isolation, characterization and genomic analysis of a novel bacteriophage VB_EcoS-Golestan infecting multidrug-resistant Escherichia coli isolated from urinary tract infection. Sci Rep 10(1):7690. https://doi.org/10.1038/s41598-020-63048-x
Yoichi M, Abe M, Miyanaga K, Unno H, Tanji Y (2005) Alteration of tail fiber protein gp38 enables T2 phage to infect Escherichia coli O157:H7. J Biotechnol 115(1):101–107. https://doi.org/10.1016/j.jbiotec.2004.08.003
Zhao S, Maurer JJ, Hubert S, De Villena JF, McDermott PF, Meng J, Ayers S, English L, White DG (2005) Antimicrobial susceptibility and molecular characterization of avian pathogenic Escherichia coli isolates. Vet Microbiol 107(3–4):215–224. https://doi.org/10.1016/j.vetmic.2005.01.021
Funding
This work was supported by the funds provided by the Indian Council of Agricultural Research under the project Outreach Programme on Zoonotic Diseases with file/grant number AS/1/1/12017-ASR-IV.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Conceptualization: Punit Jhandai and Dinesh Mittal; methodology: Punit Jhandai; investigation: Punit Jhandai; resources: Dinesh Mittal and Rajesh Khurana; data curation: Renu Gupta; writing and preparation of original draft: Punit Jhandai; review and supervision: Dinesh Mittal, Renu Gupta, Manesh Kumar, and Rajesh Khurana. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jhandai, P., Mittal, D., Gupta, R. et al. Therapeutics and prophylactic efficacy of novel lytic Escherichia phage vB_EcoS_PJ16 against multidrug-resistant avian pathogenic E. coli using in vivo study. Int Microbiol 27, 673–687 (2024). https://doi.org/10.1007/s10123-023-00420-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-023-00420-7