Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a new virus responsible for the COVID-19 pandemic. The emergence of the new SARS-CoV-2 has been attributed to the possibility of evolutionary dynamics in the furin cleavage site (FCS) region. This study aimed to analyze the sequence of the FCS region in the spike protein of SARS-CoV-2 isolates that circulated in the Special Region of Yogyakarta and Central Java provinces in Indonesia. The RNA solution extracted from nasopharyngeal swab samples of confirmed COVID-19 patients were used and subjected to cDNA synthesis, PCR amplification, sequencing, and analysis of the FCS region. The sequence data from GISAID were also retrieved for further genome analysis. This study included 52 FCS region sequences. Several mutations were identified in the FCS region, i.e., D614G, Q675H, Q677H, S680P, and silent mutation in 235.57 C > T. The most important mutation in the FCS region is D614G. This finding indicated the G614 variant was circulating from May 2020 in those two provinces. Eventually, the G614 variant totally replaced the D614 variant from September 2020. All Indonesian SARS-CoV-2 isolates during this study and those deposited in GISAID showed the formation of five clade clusters from the FCS region, in which the D614 variant is in one specific cluster, and the G614 variant is dispersed into four clusters. The data indicated there is evolutionary advantage of the D614G mutation in the FCS region of the spike protein of SARS-CoV-2 circulating in the Special Region of Yogyakarta and Central Java provinces in Indonesia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the cause of the novel coronavirus disease 2019 (COVID-19), has become the third pathogenic zoonotic disease outbreak caused by β-coronavirus, after SARS-CoV and the Middle East respiratory syndrome coronavirus (MERS-CoV) that emerged in 2002 and 2012, respectively (Zumla et al. 2016; Graham and Baric 2010; Gorbalenya et al. 2020). COVID-19 was first reported in China in December 2019 (Zhou et al. 2020) and spread rapidly causing a pandemic around the world.
The distinct characteristics of SARS-CoV-2 have not been clearly elucidated yet in comparison to the former SARS-CoV which was responsible for the previous SARS epidemic back to 2003, especially the nature of SARS-CoV-2 that makes it more contagious. It was previously speculated that PRRAR, a unique furin-like cleavage site (FCS) in the spike (S) protein, which is absent in SARS-CoV, is responsible for the high infectivity and transmissibility of SARS-CoV-2 (Coutard et al. 2020). A study in bat CoV in China identified the presence of these residues (RmYN02), suggesting a possible natural acquisition of the FCS (Zhou et al. 2020). However, the speculation that this unique FCS contributes to the improved ability of the SARS-CoV-2 to enter the host cells was not supported with the later evidence showing that the FCS did not contribute to the high fusion capacity of SARS-CoV-2 (Xia et al. 2020; Papa et al. 2021). Indeed, it remains important to further study the critical role of the FCS region in the evolutionary dynamics of SARS-CoV-2.
One of the most important mutations in the FCS region is D614G, which is reported to spread globally and responsible for the rapid and massive transmission of SARS-CoV-2 (Korber et al. 2020). The study of the biological effect of D614G mutation was facilitated by the development of SARS-CoV-2 pseudoviruses, in which the core (backbone) virus (such as retrovirus) was manipulated to express the S protein of SARS-CoV-2 (Chen and Zhang, 2021). Pseudoviruses carrying G614 variants can enter ACE2-expressing cells more efficiently than that of D614 variants, which was correlated with less S1-domain shedding and higher S protein incorporation into the virion (Zhang et al. 2020). The G614 variant enhances viral replication in human lung epithelial cells by increasing the infectivity and stability of the virions (Plante et al. 2021). Moreover, the importance of D614G mutation in the evolutionary dynamics of SARS-CoV-2 is supported by the evidence that the increased proportion of G614 variant is consistent with an evolutionary selective advantage of the virus (Volz et al. 2021).
We analyzed the sequence of FCS region in the Spike protein of SARS-CoV-2 that circulated in the Special Region of Yogyakarta and Central Java provinces, which are two different geographic locations in Indonesia, and compared the samples to the SARS-CoV-2 sequence database deposited in GISAID. Our work showed that D614G was transmitted massively in Indonesia and might involve the founder effect of SARS-CoV-2 and be responsible for the high prevalence of COVID-19 in Indonesia.
Materials and methods
Specimens, ethical clearance, sequence data, and mutation analysis
A total of 52 RNA specimens from confirmed COVID-19 patients were obtained from RNA collection at the COVID Diagnostic Laboratory, Faculty of Medicine, Public Health and Nursing, Universitas Gadjah Mada, Yogyakarta. The samples originated from the Special Region of Yogyakarta and Central Java Provinces, collected from May to October 2020.
A total of 327 whole genome sequences of SARS-CoV-2 that originated from Indonesia were obtained from GISAID (https://www.gisaid.org/) (downloaded until February 7, 2021) (Elbe and Buckland-Merrett 2017) . The D614G Hotspot-FCS Region (amino acid position = 600–700 [spike]; nucleotide position = 23,360–23,662 [Wuhan-Hu-1 NCBI Reference Sequence: NC_045512.2]) was extracted for further analysis. Sequences containing ambiguous bases (e.g., N) in the previously mentioned region were removed from the dataset using Python Script, resulting in 323 cleaned sequences.
Overall mutations were analyzed using CoVsurver: Mutation Analysis of hCoV-19 (https://www.gisaid.org/epiflu-applications/covsurver-mutations-app/). Silent mutations were identified using BioEdit (v7.2.5) (Hall 1999). Statistical data were processed using Microsoft Excel 2016 and graphs were created using GraphPad Prism 8 (www.graphpad.com). The research protocol was approved by the Medical and Health Research Ethics Committee, Faculty of Medicine, Public Health and Nursing, Universitas Gadjah Mada/Dr. Sardjito General Hospital, Yogyakarta, Indonesia (Ref: KE/FK/0511/EC/2020).
cDNA synthesis and polymerase chain reaction
First strand cDNA was synthesized using Reverse Transcription Kit II (ExcelRT™ series, Smobio), following the manufacturer’s instructions. A total of 20 µl reverse transcription reactions, containing 9 µl of RNA samples that has been previously incubated with 1 µl primer mix (50 μM oligo (dT)20 and 100 μM random hexamers) at 70 °C for 5 min and on ice for at least 1 min, along with 4 µl RT Buffer and 1 µl RTase, were incubated at 20 °C and 50 °C for 10 and 60 min, respectively, followed by 5-min incubation at 85 °C to terminate the reaction.
One primer pair targeting D614G Hotspot-FCS Region was designed for the polymerase chain reaction (PCR): F3.CoV-2.S, 5′-CTGTCCGTGATCCACAGACACT-3′; R3.CoV-2.S, 5′- GCACCAAGTGACATAGTGTAGGC-3′. The PCR was conducted in a 25 µl-reaction mix of GoTaq ® Green Mastermix (Promega), consisting of 12.5 µl GoTaq Green Mastermix (2 ×), 0.5 µl of each primer (10 µM), and 2 µl cDNA template. Amplification steps were conducted as follows: 95 °C for 3 min; followed by 35 cycles of 95 °C, 60 °C, and 72 °C for 30 s, 45 s, and 45 s, respectively; and final extension was 2 min at 72 °C.
Sequencing
All PCR products were prepared for sequencing using Sanger methods. Sequencing procedure was conducted by DNA Sequencing Services (Genetika Science, Indonesia; 1st Base, Singapore) using BigDye® Terminator v3.1 Cycle Sequencing Chemistry Kit (Applied Biosystems).
Phylogenetic analysis
The maximum likelihood (ML) tree was constructed for both viral-specific and species-specific approaches. For the SARS-CoV-2-specific ML tree, initially the redundant sequences with 100% identity were removed from 323 Indonesian origin SARS-CoV-2 sequences obtained from GISAID. The dataset which previously consisted of 52 fresh samples, NCBI reference sequence (NC_045512.2), and the non-redundant Indonesian origin database sequences was completed by adding several sequences from important SARS-CoV-2 variants, such as the first reported G614 variant (EPI_ISL_451345), B.1.429/452R.V1 (EPI_ISL_915439), B.1.1.7/501Y.V1 (EPI_ISL_601443), B.1.351/501Y.V2 (EPI_ISL_712081), P.1/501Y.V3 (EPI_ISL_833137), and Mink Cluster (EPI_ISL_616695). A total of 92 sequences were used to construct the ML tree using RaXML v.8.2.10 with 1000 bootstraps (Silvestro and Michalak 2012). For every dataset in this study, multiple sequence alignment was performed using MAFFT (v7.471) with L-INS-i algorithm (Katoh et al. 2005; Katoh and Standley 2013). The ML tree was viewed and annotated using MEGA (v10.1.1) (Kumar et al. 2018).
Phylogenetic analysis by Bayesian Markov Chain Monte Carlo (MCMC) method was conducted using BEAST (v1.10.4) (Suchard et al. 2018). Three independent datasets, comprised of Indonesia (n = 371), Yogyakarta (n = 68), and Central Java (n = 43), were analyzed separately. Parameters for BEAST analysis were generated using Beauti (v1.10.4) (Drummond et al. 2012). The substitution model of HKY + Γ with 2 partitions ((position 1 + 2), 3) was used for MCMC analysis, in addition to the strict molecular clock and the constant coalescent tree prior parameter (Drummond et al. 2002; Kingman 1982). The MCMC estimation for Indonesia, Yogyakarta, and Central Java was independently run with the chain length 5 × 100, 1 × 50, and 1 × 30 million, respectively, and sampling every 100, 10, and 10 thousand, respectively. Burn-in 10% was used for further analysis of log files in Tracer (v1.7.1) (Rambaut et al. 2018). We ensured that every parameter had effective sample size (ESS) value that was no less than 100. The maximum clade credibility (MCC) tree was generated using TreeAnnotator (v1.10.4) (Drummond and Rambaut 2007). MCC tree visualization and annotation were done using FigTree (v1.4.4) (Rambaut 2018).
Results
The distribution of D614G mutation in Indonesia
The FCS region of the S protein derived from 52 SARS-CoV-2 isolates which were circulated in the Special Region of Yogyakarta (27 sequences) and Central Java provinces (25 sequences) Indonesia from May to October 2020 were sequenced during this study (Fig. 1).
The geographical distribution of the D614G variants reported in Indonesia from April 2020 to January 2021. The GISAID database showed that the D614 variant has been found in 25 provinces in Indonesia and it was subsequently replaced by the new G614 variant, including in the Special Region of Yogyakarta and Central Java provinces (inset). Sequence data from this study showed that during May to October 2020, the D614 variant remained circulating in Special Region of Yogyakarta in exceedingly small numbers (the last date of collected sample was August 2020), while all analyzed samples in Central Java were the G614 variant. Blue dots: D614 variants, Orange dots: G614 variants
D614G is the most frequent mutation in the FCS region of S protein of SARS-CoV-2 found in this study. There were 26 G614 variants (96.3%) and only one D614 variant (3.7%) of SARS-CoV-2 that circulated in the Special Region of Yogyakarta and all samples obtained from Central Java were the G614 variant (100%) (Fig. 1). The dominant proportions of the G614 variant in those two provinces were also observed when our data was combined with the sequence data obtained from the GISAID data base. In total, 52 SARS-CoV-2 FCS sequence data obtained from those two provinces were available from this study and GISAID database (Fig. 1). However, the frequency of G614 in those two provinces were more frequent compared to all SARS-CoV-2 isolates from Indonesia which have been deposited in GISAID (Fig. 1). The GISAID data showed that geographically the G614 variant has spread to 25 of 34 provinces in Indonesia, in contrast to the D614 variants which were reported in 11 provinces (Fig. 2).
The phylodynamics of D614G mutation in Special Region of Yogyakarta and Central Java
The time dynamics of D614G mutations among SARS-CoV-2 that circulated in the two provinces were also analyzed. The first G614 variant was detected in Yogyakarta on May 6, 2020 and in Central Java on May 12, 2020. Although we cannot exclude the possibility that G614 variants had been circulating before May 2020, the latest D614 variant was isolated in Yogyakarta in August 2020 (Fig. 3). This finding is parallel with the time dynamics of D614G in Indonesian SARS-CoV-2 isolates. The data obtained from this study (52 isolates) in combination with all Indonesian SARS-CoV-2 isolates documented in GISAID database (323) demonstrated that the D614 variant was totally replaced by the G614 variant from September 2020 (Fig. 4).
The proportion of D614G variant in Indonesia based on this study and GISAID data. The G614 variant in Indonesia was first reported since April 2020 and increasingly found to be the predominant variant. The D614 variant was no longer reported in September 2020 as indicated by this study and GISAID data analyzed from March 2020 to January 2021
The mutations and clusters distribution of FCS region in the Indonesian sample
We detected several mutations accompanying the D614G variant, i.e., Q675H, Q677H, S680P, and silent mutation in 23,557 C > T. Two sequences (3.85%) came from Yogyakarta containing Q677H mutation as of August 2020. Other variants S680P and Q675H were observed from Central Java with 1.92% of each in June and October 2020, respectively. The silent mutation was detected with 1.92% from Yogyakarta. These mutations are co-segregated with D614G (Fig. 5).
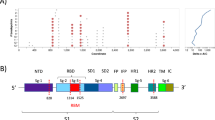
Phylogeny tree of 52 research samples. maximum likelihood (ML) tree from furin cleavage site (FCS) region of research samples (red dot) with several FCS SARS-CoV-2 strains originating from the GeneBank (black dots were originated from Indonesia and blue dots were from another country) showed 4 different mutations in the FCS region, Q675H, Q677H, one mutation near the furin cleavage PRRAR was S680P, and one silent mutation in 23,557 C > T
Analysis of the MCC Tree from the data obtained during this study and all Indonesian SARS-CoV-2 isolates deposited in GISAID showed the formation of five clade clusters from the FCS region, in which the D614 variant is in one specific cluster, and the G614 variant is dispersed into four clusters (Fig. 6).
The distribution of mutations in the FCS region in Indonesian samples. Analysis of the maximum clade credibility (MCC) tree in the research samples and the Indonesian samples obtained from GISAID data showed the formation of five clusters from the FCS region, with S680P and Q677H in the same cluster
Discussion
The G614 variant emerged in Europe since February 2020 and has spread rapidly throughout the world (Korber et al. 2020; Volz et al. 2021). This study indicated that the G614 variant was detected as the most dominant variant as early as May 2020 in the Special Region of Yogyakarta and Central Java.
The variant analysis of SARS-CoV-2 obtained from the Special Region of Yogyakarta and Central Java between May and October 2020 found only one D614 variant in Yogyakarta in August 2020, while in Central Java province there were no D614 variants identified (Fig. 1). Although these data cannot represent the overall incidence of SARS-CoV-2 infection of D614 and G614 variants in both provinces, it is clear that there was a shift in the SARS-CoV-2 variants circulating in Indonesia (Fig. 4). The same incidence was reported in London, where D614 was the most common variant at the early phase of the pandemic and the transition to G614 surpassed D614 at the end of March 2020 (Volz et al. 2021).
The data of the present study showed the existence of the G614 variant in Yogyakarta and Central Java since May 2020, earlier than the samples reported in GISAID in July 2020. This pattern means that 3 months after the G614 variant was found in Europe, this G614 variant had been circulating in Yogyakarta and Central Java, Indonesia.
The spread of G614 was very fast and massive, because the nature of the G614 variant has higher transmissibility and greater competitive fitness than the wild-type virus (Korber et al. 2020; Hou et al. 2020). This massive spread is further supported by the human travelling patterns and highly extensive transportation system all around the world. Although many countries have specific border restriction policies, the viral spread has not been effectively contained (Mallapaty 2021).
The G614 variant has completely replaced the D614 variant since September 2020 in all Indonesian regions (Fig. 4). This finding may correlate with a significant increase of confirmed COVID-19 cases in Indonesia during that particular period and the following months. The first case of COVID-19 in Indonesia was announced on March 2, 2020. In September 2020, the reported confirmed cases were less than 400,000 cases and increased rapidly to 1.4 million cases by early March 2021 (World Health Organization 2021). There are many factors that can contribute to the increase of COVID-19 cases in Indonesia. However, the role of mutations in the circulating virus characteristics should not be underestimated (Hou et al. 2020).
There are four mutations that appear to occur in the G614 variant only. Further analysis showed that G614 variants developed into four genetics clusters in Indonesia, separated from the clusters consisting of D614 variants (Fig. 6). All together with the epidemiological data showing the significant increase of COVID-19 cases in Indonesia, this pattern may support the indication that D614G variant is a founder effect of SARS-CoV-2 in Indonesia (Gunadi et al. 2020). The development of D614G variant may be the consequence of an evolutionary advantage of the D614G mutation in the FCS region of the S protein of SARS-CoV-2 circulating in Indonesia.
Currently, continuous transmission of SARS-CoV-2 leads to the emergence of novel variants with increased fitness advantage and consequently, may have a significant impact on the ongoing pandemic, including Indonesia (Perez-Gomez 2021). The SARS-CoV-2 variants of concern (VOC), including Alpha, Beta, Gamma, Delta, and the latest Omicron variant, have been associated with the surge of confirmed COVID-19 cases globally. The massive increase of COVID-19 cases in Indonesia in July 2021 was associated with the introduction of Delta variant in Indonesia. The first confirmed case of Delta variant in Central Java and Yogyakarta was on May 25, 2021 and shortly became the most dominant VOC in Indonesia (Gunadi et al. 2021). Interestingly, all these VOC harbor the D614G mutation, indicating that this mutation is positively selected and subsequently drives the dynamic evolution of SARS-CoV-2 up to this moment (He et al. 2021). Thus, D614G mutation provides a selective advantage and increases the transmissibility of SARS-CoV-2 (Harvey et al. 2021).
We found two other mutations, Q675H and Q677H, which were also reported by Begum et al. (2020) and Kim et al. (2020). These two mutations might be important because mutations near the S1/S2 cleavage site could influence the activity of the protease enzymes with respect to the cleavage sites (Begum et al. 2020). One mutation before the polybasic cleavage site within the S gene, PRRAR (681–685 amino acid) motif was identified as S680P which was previously reported by Kim et al. (Kim et al. 2020). Learning from evidence that the FCS region is important and the potential role of D614G as a founder effect of other mutations in this region, further molecular epidemiology surveillance of the FCS region mutations is important, especially in the area where transmission of COVID-19 is still high, such as Indonesia.
In conclusion, we demonstrated here that G614 variant was detected in our clinical samples since May 2020 and replaced the D614 variant massively. This finding indicated that D614G mutation is an evolutionary advantage for the SARS-CoV-2 transmission in the Special Region of Yogyakarta and Central Java provinces of Indonesia. Mutation in the FCS region is an important event which may contribute to the evolutionary process of virus in particular geographical area which eventually influence the phenotypic characteristic of the virus, including the transmissibility.
Data availability
Please contact author for data requests.
References
Begum F, Banerjee AK, Ray U (2020) Mutation hot spots in spike protein of SARS-CoV-2 virus. Preprints Available from: https://doi.org/10.20944/preprints202004.0281.v2. Accessed 15 Sept 2020
Chen M, Zhang X-E (2021) Construction and applications of SARS-CoV-2 pseudoviruses: a mini review. Int J Biol Sci 17(6):1574–1580
Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir Res 176:104742. https://doi.org/10.1016/j.antiviral.2020.104742
Drummond AJ, Nicholls GK, Rodrigo AG, Solomon W (2002) Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Genetics 161(3):1307–1320. https://doi.org/10.1016/0370-2693(87)91197-X
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7(214):1–8. https://doi.org/10.1186/1471-2148-7-214
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29(8):1969–1973. https://doi.org/10.1093/molbev/mss075
Elbe S, Buckland-Merrett G (2017) Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob Chall 1(1):33–46. https://doi.org/10.1002/gch2.1018
Gorbalenya AE, Baker SC, Baric RS, de Groot RJ, Drosten C, Gulyaeva AA, Haagmans BL, Lauber C, Leontovich AM, Neuman BW, Penzar D, Perlman S, Poon LLM, Samborskiy DV, Sidorov IA, Sola I, Ziebuhr J (2020) The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol 5:536–544. https://doi.org/10.1038/s41564-020-0695-z
Graham RL, Baric RS (2010) Recombination, reservoirs, and the modular spike: mechanism of coronavirus cross-species transmission. J Virol 84(7):3134–3146. https://doi.org/10.1128/JVI.01394-09
Gunadi WH, Marcellus HMS, Daniwijaya EW, Rizki LP, Supriyati E, Nugrahaningsih DAA, Afiahayati S, Iskandar K, Anggorowati N, Kalim AS, Puspitarani DA, Athollah K, Arguni E, Nuryastuti T, Wibawa T (2020) Full-length genome characterization and phylogenetic analysis of SARS-CoV-2 virus strains from Yogyakarta and Central Java. Indonesia Peerj 8:e10575. https://doi.org/10.7717/peerj.10575
Gunadi, Hakim MS, Wibawa H, Marcellus, Setiawaty V, Slamet, Trisnawati I, Supriyati E, El Khair R, Iskandar K, Afiahayati, Siswanto, Irene, Anggorowati N, Daniwijaya EW, Nugrahaningsih DAA, Puspadewi Y, Puspitarani DA, Tania I, Vujira KA, Ardlyamustaqim MB, Gabriela GC, Eryvinka LS, Nirmala BC, Geometri ET, Darutama AA, Kuswandani AA, Lestari, Irianingsih SH, Khoiriyah S, Lestari I, Ananda NR, Arguni E, Nuryastuti T, Wibawa T (2021) Is the infection of the SARS-CoV-2 Delta variant associated with the outcomes of COVID-19 patients? Front Med (Lausanne) 8:780611
Hall TA (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Harvey WT, Carabelli AM, Jackson B, Gupta RK, Thomson EC, Harrison EM, Ludden C, Reeve R, Rambaut A, COVID-19 Genomics UK (COG-UK) Consortium, Peacock SJ, Robertson DL (2021) SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol 19(7):409–424
He X, Hong W, Pan X, Lu G, Wei X (2021) SARS-CoV-2 Omicron variant: characteristics and prevention. MedComm 2:838–845
Hou YJ, Chiba S, Halfmann P, Ehre C, Kuroda M, Dinnon KH, Leist SR, Schäfer A, Nakajima N, Takahashi K, Lee RE, Mascenik TM, Graham R, Edwards CE, Tse LV, Okuda K, Markmann AJ, Bartelt L, de Silva A, Margolis DM, Boucher RC, Randell SH, Suzuki T, Gralinski LE, Kawaoka Y, Baric RS (2020) SARS-CoV-2 D614G variant exhibits efficient of replication ex vivo and transmission in vivo. Science 370(6523):1464–1468. https://doi.org/10.1126/science.abe8499
Katoh K, Kuma KI, Toh H, Miyata T (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33(2):511–518. https://doi.org/10.1093/nar/gki198
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kingman JFC (1982) The coalescent. Stoch Process Appl 13(3):235–248. https://doi.org/10.1016/0304-4149(82)90011-4
Kim JS, Jang JH, Kim JM, Chung YS, Yoo CK, Han MG (2020) Genome-wide identification and characterization of point mutations in the SARS-CoV-2 genome. Osong Public Health Res Perspect 11(3):101–111
Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, de Silva TI, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, Montefiori DC, Angyal A, Brown RL, Carrilero L, Green LR, Groves DC, Johnson KJ, Keeley AJ, Lindsey BB, Parsons PJ, Raza M, Rowland-Jones S, Smith N, Tucker RM, Wang D, Wyles MD (2020) Tracking changes in SARS-CoV-2 spike: evidence that D614G Increases Inactivity of the COVID virus. Cell 182(4):812–827. https://doi.org/10.1016/j.cell.2020.06.043
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Mallapaty S (2021) What the data say about border closures and COVID spread. Nature 589(185). Available from: https://www.nature.com/articles/d41586-020-03605-6. Accessed 27 Dec 2020
Papa G, Mallery DL, Albecka A, Welch LG, Cattin-Ortolá J, Luptak J, Paul D, McMahon HT, Goodfellow IG, Carter A, Munro S, James LC (2021) Furin cleavage of SARS-CoV-2 spike promotes but is not essential for infection and cell-cell fusion. PLoS Pathog 17(1):1009246. https://doi.org/10.1371/journal.ppat.1009246
Perez-Gomez R (2021) The development of SARS-CoV-2 variants: the gene makes the disease. J Dev Biol 9(4):58
Plante JA, Liu Y, Liu J, Xia H, Johnson BA, Lokugamage KG, Zhang X, Muruato AE, Zou J, Fontes-Garfias CR, Mirchandani D, Scharton D, Bilello JP, Ku Z, An Z, Kalveram B, Freiberg AN, Menachery VD, Xie X, Plante KS, Weaver SC, Shi PY (2021) Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592:116–121. https://doi.org/10.1038/s41586-020-2895-3
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67(5):901–904. https://doi.org/10.1093/sysbio/syy032
Rambaut A (2018) FigTree v. 1.4.4. Molecular evolution, phylogenetics and epidemiology: Andrew Rambaut; Available from: http://tree.bio.ed.ac.uk/software/figtree/. Accessed 30 Mar 2021
Silvestro D, Michalak I (2012) RaxmlGUI: a graphical front-end for RAxML. Org Divers Evol 12:335–337. https://doi.org/10.1007/s13127-011-0056-0
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol 4(1):1–5. https://doi.org/10.1093/ve/vey016
Volz E, Hill V, McCrone JT, Price A, Jorgensen D, O’Toole Á, Southgate J, Johnson R, Jackson B, Nascimento FF, Rey SM, Nicholls SM, Colquhoun RM, da Silva FA, Shepherd J, Pascall DJ, Shah R, Jesudason N, Li K, Jarrett R, Pacchiarini N, Bull M, Geidelberg L, Siveroni I, Goodfellow I, Loman NJ, Pybus OG, Robertson DL, Thomson EC, Rambaut A, Connor TR (2021) Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Cell 184(1):64-75.e11. https://doi.org/10.1016/j.cell.2020.11.020
World Health Organization (2021) WHO Indonesia Situation Report – 46. Available from: https://www.who.int/indonesia/news/novel-coronavirus/situation-reports. Accessed 04 Mar 2021
Xia S, Lan Q, Su S, Wang X, Xu W, Liu Z, Zhu Y, Wang Q, Lu L, Jiang S (2020) The role of furin cleavage site in SARS-CoV-2 spike protein-mediated membrane fusion in the presence or absence of trypsin. Signal Transduct Target Ther 5(1):92. https://doi.org/10.1038/s41392-020-0184-0
Zhang L, Jackson CB, Mou H, Ojha A, Peng H, Quinlan BD, Rangarajan ES, Pan A, Vanderheiden A, Suthar MS, Li W, Izard T, Rader C, Farzan M, Choe H (2020) SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat Commun 11:6013. https://doi.org/10.1038/s41467-020-19808-4
Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, Wang P, Liu D, Yang J, Holmes EC, Hughes AC, Bi Y, Shi W (2020) A novel bat coronavirus closely related to SARS-CoV-2 contains natural insertions at the S1/S2 cleavage site of the spike protein. Curr Biol 30(11):2196-2203.e3. https://doi.org/10.1016/j.cub.2020.05.023
Zumla A, Chan JFW, Azhar EI, Hui DSC, Yuen KY (2016) Coronaviruses – drug discovery and therapeutic options. Nat Rev Drug Discov 15(5):324–347. https://doi.org/10.1038/nrd.2015.37
Acknowledgements
Part of the data were extracted from the graduate thesis work of FMG. We gratefully acknowledge the authors and the originating and submitting laboratories for their sequences and metadata that they have shared through GISAID.
Funding
This work is partly supported by the COVID-19 Research Grant from the Universitas Gadjah Mada (Sofia Mubarika Haryana).
Author information
Authors and Affiliations
Contributions
Conceptual and experimental design: TW, MSH, NW, EA, TN, SMH. Data collection: FMG, MNM, NW, RN, ES, MEDW. Data analysis: FMG, NW, RN, MNM, TW. First draft of the manuscript writing: NW, TW. Paper reviewing and revising: FMG, NW, TW, SMH, MEDW, EA, MSH, ES, TN. All authors have access to the raw data. The final manuscript has been read and approved by all authors.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study protocol and informed consent form were approved by the Medical and Health Research Ethics Committee Faculty of Medicine, Public Health, and Nursing, Universitas Gadjah Mada/Dr. Sardjito General Hospital Yogyakarta, Indonesia (Ref: KE/FK/0511/EC/2020).
Consent for publication
Not applicable.
Competing interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wijayanti, N., Gazali, F.M., Supriyati, E. et al. Evolutionary dynamics of SARS-CoV-2 circulating in Yogyakarta and Central Java, Indonesia: sequence analysis covering furin cleavage site (FCS) region of the spike protein. Int Microbiol 25, 531–540 (2022). https://doi.org/10.1007/s10123-022-00239-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00239-8