Abstract
Background
Gastric cancer risk can be accurately predicted by measuring the methylation level of a single marker gene in gastric mucosa. However, the mechanism is still uncertain. We hypothesized that the methylation level measured reflects methylation alterations in the entire genome (methylation burden), induced by Helicobacter pylori (H. pylori) infection, and thus cancer risk.
Methods
Gastric mucosa of 15 healthy volunteers without H. pylori infection (G1), 98 people with atrophic gastritis (G2), and 133 patients with gastric cancer (G3) after H. pylori eradication were collected. Methylation burden of an individual was obtained by microarray analysis as an inverse of the correlation coefficient between the methylation levels of 265,552 genomic regions in the person’s gastric mucosa and those in an entirely healthy mucosa.
Results
The methylation burden significantly increased in the order of G1 (n = 4), G2 (n = 18), and G3 (n = 19) and was well correlated with the methylation level of a single marker gene (r = 0.91 for miR124a-3). The average methylation levels of nine driver genes tended to increase according to the risk levels (P = 0.08 between G2 vs G3) and was also correlated with the methylation level of a single marker gene (r = 0.94). Analysis of more samples (14 G1, 97 G2, and 131 G3 samples) yielded significant increases of the average methylation levels between risk groups.
Conclusions
The methylation level of a single marker gene reflects the methylation burden, which includes driver gene methylation, and thus accurately predicts cancer risk.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prediction of gastric cancer risk is very important to implement endoscopic screening for those who need it and to remove unnecessary concerns for those with negligible risk, even after eradication of Helicobacter pylori (H. pylori) [1]. To this end, multiple modalities, such as endoscopic findings and pepsinogen I/II ratio, have been developed [2,3,4]. At the same time, DNA methylation in gastric mucosa was shown to be highly useful, and multiple marker genes have been isolated after extensive selection in a number of cross-sectional studies [5,6,7,8]. Even compared with point mutation accumulations, DNA methylation accumulation in gastric mucosa had a greater impact on cancer risk [9]. Importantly, a multicenter prospective cohort study demonstrated that the methylation level of a single marker gene, such as miR124a-3, EMX1, and NKX6-1, could predict gastric cancer risk accurately [10, 11]. However, it is still unclear why the methylation level of a single marker gene can predict gastric cancer risk accurately.
Biologically, DNA methylation of a promoter CpG island (CGI) consistently represses transcription of its downstream gene, and this mechanism can inactivate tumor-suppressor genes, such as CDH1, CDKN2A and MLH1 [12, 13]. Indeed, in gastric cancer cells, cancer-related pathways were more frequently inactivated by aberrant DNA methylation than by mutations [14]. The deep involvement of aberrant DNA methylation in gastric cancer is explained by the fact that H. pylori infection, the most important inducer of gastric cancer, potently induces aberrant DNA methylation via induction of chronic inflammation [15,16,17,18]. DNA methylation is induced in a large number of genomic regions, which include i) promoter CGIs of driver genes, ii) those of passenger genes, and iii) CGIs in exons [19]. These lead to a hypothesis that methylation alterations in the entire genome (methylation burden), the sum of deviations in the methylation levels of genome-wide loci from an entirely healthy gastric mucosa, is critical for gastric cancer risk, and that a marker gene methylation level, if carefully selected, is correlated with the methylation burden.
In this study, we aimed to demonstrate that the methylation burden and the accumulation of driver gene methylation in gastric mucosae were correlated with the methylation level of a single marker gene.
Materials and methods
Tissue samples and DNA extraction
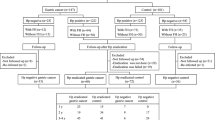
Gastric mucosa samples, regardless of the presence of atrophy or intestinal metaplasia, were collected from 246 people, who were classified into three groups (Groups 1, 2 and 3). Group 1 (G1) consisted of 15 healthy volunteers who had never been infected with H. pylori, who were considered to have a low risk of developing gastric cancer [20]. Group 2 (G2) consisted of 98 healthy individuals who received successful eradication of H. pylori and had gastric atrophy, who were considered to have an intermediate risk [21]. Group 3 (G3) consisted of 133 gastric cancer patients who underwent curative endoscopic submucosal dissection and received successful eradication of H. pylori, who were considered to have a high risk [22]. Endoscopic gastric mucosal atrophy was determined by Kimura-Takemoto classification [23]. Among all the participants, seven G1 and 27 G2 people were newly recruited, and the others were enrolled in our previous studies [11, 19]. The previous studies and this study were approved by the National Cancer Center Ethics Committee (approval nos. 2008–104, 2015–139) and Hoshi University Research Ethics Committee (approval nos. 2022–10, 2022–12), and written informed consent was obtained from all participants. Characteristics of all the samples are shown in Supplementary Table 1.
All the samples were endoscopically biopsied from antral region and were stored in RNA later (Life Technologies, Carlsbad, CA, USA) or ISOGEN (Nippon Gene, Tokyo, Japan). For H. pylori-eradicated individuals, the samples were collected six months or more after eradication. Genomic DNA was extracted by the phenol/chloroform method, and was quantified using a Quant-iT PicoGreen dsDNA Assay Kit (Life Technologies).
Selection of driver and marker genes
Driver genes were selected based on previous papers in which those genes were reported as driver genes of gastric cancer [14, 24, 25] (Supplementary Table 2). As marker genes, we selected two marker genes, miR124a-3 and EMX1, which were previously identified by a genome-wide analysis and were shown to be useful as a cancer risk marker in a prospective clinical study [10, 11].
Genome-wide and region-specific DNA methylation analysis
Genome-wide DNA methylation analysis was performed by an Infinium HumanMethylation450 BeadChip microarray (Illumina, CA, USA) using 500 or 750 ng genomic DNA modified by sodium bisulfite using an EZ DNA Methylation Kit (Zymo Research, CA, USA). A DNA methylation level was obtained as a β value in the range from 0 (unmethylated) to 1 (fully methylated), as previously described [26]. 485,512 probes were assembled into 276,831 genomic blocks using MACON web tool (https://rias.rhelixa.com/macon/human_methylation/new) [27]. A genomic block was defined as a collection of probes within 500 bp, and annotated by its location against a transcription start site (TSS) and a CpG island (CGI). A single genomic block in a CGI and within 200 bp from a TSS of a driver gene, except for CDKN2A (p16), was assessed for its methylation level. For CDKN2A, a probe immediately downstream of its TSS was analyzed because no probes were located in 200 bp from its TSS although it had a CGI spanning from its promoter region to exon 1.
Region-specific DNA methylation analysis was performed by bisulfite-pyrosequencing using PyroMark Q24 Advanced (Qiagen, CA, USA). Genomic DNA (1 μg) was modified by sodium bisulfite using an EZ DNA Methylation Kit (Zymo Research). A target CpG site of a driver gene was selected in the same CGI analyzed in the genome-wide DNA methylation analysis, except for CDKN2A. For CDKN2A, its target CpG site was located in 200 bp upstream region from its TSS. DNA methylation levels of the target CpG site were calculated using PSQ Assay Design software (Qiagen). The primer sequences for the target CpG site and PCR conditions are listed in Supplementary Table 3 [28]. Fully unmethylated DNA was prepared by amplifying genomic DNA using an Illustra™ GenomiPhi™ DNA amplification kit (Cytiva, Tokyo, Japan) twice, and fully methylated DNA was prepared by methylating the fully unmethylated DNA using CpG methyltransferase (M. SssI) (New England Bio Labs, Ipswich, MA, USA).
Calculation of methylation burden
The methylation burden was defined as the sum of deviations in the methylation levels in the entire genome from an entirely healthy gastric mucosa. From the 276,831 genomic blocks, 6,582 blocks on sex chromosome and 4,697 blocks whose methylation levels failed to be analyzed in any of the samples were excluded, and 265,552 genomic blocks were used for calculation. The methylation burden of an individual was calculated as an inverse of the correlation coefficient between the methylation levels of the 265,552 genomic blocks in the person’s gastric mucosa and those in an entirely healthy gastric mucosa, namely the average of four young healthy individuals (G1_Y1-Y4).
Statistical analysis
Scatter plots were drawn by R (version 4.2.0.). Pearson’s correlation coefficients were calculated also by R. The methylation burden and the average methylation levels of driver and marker genes were compared between two of the three groups (G1, G2, and G3) by Student’s t test. P values < 0.05 were considered as significant.
Results
Increased methylation burden according to the risk groups
To investigate methylation burden in the three risk groups, we performed genome-wide DNA methylation analysis of four entirely healthy gastric mucosa and four old G1, 18 G2 and 19 G3 samples (Supplementary Table 4). The methylation burdens were calculated by comparing between the average methylation level of four entirely healthy gastric mucosa and the methylation level of an individual sample in three risk groups (Fig. 1a, and Supplementary Fig. 1). The methylation burden significantly increased according to the risk levels of the samples (P < 0.05 for G1 vs G2 and G2 vs G3) (Fig. 1b). In addition, the methylation burden was well correlated with the methylation level of a single marker gene (r = 0.910 for miR124a-3, and 0.931 for EMX1) (Fig. 1c). Together, the methylation burden was considered to be highly correlated with gastric cancer risk and could be measured by the methylation level of a single marker gene in gastric mucosa.
Methylation burden in the entire genome, which was considered as a deviation of the methylation levels from an entirely healthy gastric mucosa. a Representative analysis of the methylation burden in old G1, G2, and G3 samples. Each scatter plot shows the differences of the methylation levels (β values) of 265,552 genomic blocks in the entire genome between an individual sample and an entirely healthy gastric mucosa. b The methylation burden in the three risk groups. The methylation burden significantly increased according to the risk levels of samples. c A correlation between the methylation burden and the methylation level of a single marker gene. The methylation burden was well correlated with the methylation level of a single marker gene
Accumulation of driver gene methylation in the three risk groups
The methylation burden was composed of hypermethylation of many genomic blocks and hypomethylation, also of many blocks. In general, hypermethylation of promotor CGI of tumor suppressor genes can silence these genes, and global hypomethylation can lead to chromosomal instability [29]. Hence, the increased methylation burden was expected to lead to methylation of multiple driver genes and increased gastric cancer risk. Therefore, we selected 59 driver genes based on the literature [14, 24, 25] (Supplementary Table 2), and analyzed the accumulation of methylation of the 59 driver genes using their average methylation level. The average level increased according to the risk levels of the samples, but not significantly (P < 0.05 for G1 vs G2, and P = 0.56 for G2 vs G3) (Fig. 2a). The average level was correlated with the methylation level of a single marker gene (r = 0.89 for miR124a-3, and 0.85 for EMX1) (Fig. 2b). The lack of significant difference between G2 and G3 suggested that the selection of driver genes needed improvement.
Accumulation tendency of driver gene methylation. a Accumulation levels of 59 driver genes in the three risk groups were measured as their average methylation levels. The average level increased according to the risk levels of the samples, but not significantly. b A high correlation between the average methylation levels of the 59 driver genes and the methylation level of a single marker gene, miR124a-3 or EMX1. c The average methylation levels of nine driver genes in the three risk groups. The average level tended to increase according to the risk levels of the samples (P < 0.05 for G1 vs G2, and P = 0.08 for G2 vs G3). d A high correlation between the average methylation levels of the nine driver genes and the methylation level of a single marker gene
Therefore, we conducted additional literature searches to select genes extensively analyzed by multiple investigators in gastric cancer using the following search terms [(gene name) AND (methylation) AND (gastric cancer) AND ((silencing) OR (silenced) OR (inactivated) OR (inactivating))] in the PubMed database. We selected two genes that had at least three papers from eight major pathways of gastric cancer. Finally, nine driver genes were selected (CDKN2A, CHFR, CDH1, BNIP3, miR34B, RPRM, MLH1, SFRP1, and SFRP5) (Supplementary Table 5). The average methylation levels of the nine driver genes tended to increase according to the risk levels of the samples (P < 0.05 for G1 vs G2, and P = 0.08 for G2 vs G3) (Fig. 2c). The average level was also well correlated with the methylation level of a single marker gene (r = 0.944 for miR124a-3, and 0.92 for EMX1) (Fig. 2d). The result indicated that DNA methylation of driver genes was accumulated in gastric mucosa, composing gastric cancer risk.
Driver gene methylation in a large-scale sample
To confirm the accumulation of driver gene methylation according to gastric cancer risk, we increased the number of samples by adopting a cost-effective method, namely pyrosequencing (Supplementary Table 6). We successfully designed high-quality primers for pyrosequencing of eight of the nine driver genes (CDKN2A, CHFR, CDH1, miR34B, RPRM, MLH1, SFRP1, and SFRP5). The high quality of the primers was confirmed from two aspects. First, linearity was evaluated by analyzing the mixture of fully unmethylated DNA and fully methylated DNA. The primers for the eight driver genes and the two marker genes had good linearity, especially at low DNA methylation levels (Supplementary Fig. 2a). Second, we analyzed the correlation between the result obtained by DNA methylation microarray and that by pyrosequencing using the same samples analyzed by the DNA methylation microarray. There was a high correlation between the two measurement methods for the eight driver genes (Supplementary Fig. 2b). CDKN2A did not show such correlation due to different locations of target CpG sites (Supplementary Fig. 2c; see discussion).
The methylation levels of the eight driver genes were analyzed in 14 G1, 97 G2, and 131 G3 samples (Supplementary Table 6). CHFR, miR34B, RPRM and SFRP1 were methylated and their methylation levels increased according to the risk levels of the samples. CDKN2A and MLH1 were not methylated at detectable levels in almost any of the samples. CDH1 and SFRP5 were more methylated in G2 and G3 samples than in G1 samples, but their methylation levels did not increase more in G3 than in G2 samples. The two marker genes showed increased methylation levels according to the risk levels of the samples (Fig. 3a). The average methylation levels of the eight driver genes significantly increased according to the risk levels of the samples (P < 0.05 for G1 vs G2 and G2 vs G3) (Fig. 3b). The average level was correlated with the methylation level of a single marker gene (Fig. 3c).
Significant increases of the accumulation levels of driver gene methylation using a large number of samples. a The methylation levels of eight driver genes and two marker genes in the three risk groups. In CHFR, miR34B, RPRM and SFRP1, their methylation levels increased according to the risk levels. CDKN2A and MLH1 were not methylated at detectable levels in almost any of the samples. CDH1 and SFRP5 were more methylated in G2 and G3 samples than in G1 samples, but their methylation levels were lower in G3 samples than in G2 samples. b The average methylation levels of the eight driver genes. The average level significantly increased according to the risk levels of the samples. c A high correlation between the average methylation levels of the eight driver genes and the methylation level of a single marker gene
Subgroup analysis by age, gender, and gastric atrophy
Finally, the independence of methylation level from known risk factors of gastric cancer, namely age [30, 31], gender [32], and the extent of gastric atrophy [33], was assessed by subgroup analyses. In the age subgroups with 65 years old as a cutoff point, the methylation level of miR124a-3 was significantly higher in G3 than in G2. Similarly, in males and females, the methylation level of miR124a-3 was significantly higher in G3 than in G2. In the subgroups with closed-type and open-type atrophy, the methylation level of miR124a-3 was significantly higher in G3 than in G2. On the other hand, within the same risk group, there were no significant differences in the methylation level of miR124a-3 by age, gender, and the extent of gastric atrophy (Fig. 4, Supplementary Fig. 3 for EMX1 and the eight driver genes). These findings showed that the methylation level of a single marker gene and the average methylation levels of the eight driver genes were associated with gastric cancer risk, even after stratifying for age, gender, and gastric atrophy.
Subgroup analyses by age, gender and gastric atrophy. There were significant differences in the methylation levels of miR124a-3 between G2 and G3 in all the three subgroup analyses. At the same time, within one risk group, there were no significant differences in the methylation levels of miR124a-3 by age, gender, or gastric atrophy. yr, year; Closed, Closed-type atrophy; Open, Open-type atrophy; *P < 0.05
Discussion
In the current study, we demonstrated that methylation burden, along with the accumulation of driver gene methylation, increased according to the risk levels of gastric mucosa samples, and that they were strongly correlated with the methylation level of a single marker gene. This result explained the reason why the methylation level of a single marker gene can accurately predict cancer risk. It provides a rationale for the clinical use of measuring the methylation level of a single marker gene for cancer risk prediction. The use of a single marker gene is convenient and inexpensive, and will bring great benefits to patients and healthcare burden. Our finding is expected to be applicable to other cancers such as colitic, liver, and uterine cervical cancer, which are closely related to chronic inflammation. Indeed, several marker genes have been identified for these cancers [34,35,36].
The methylation burden was defined as the sum of deviations of methylation levels of genomic loci throughout the genome in an individual sample from those in an entirely healthy gastric mucosa, including hypermethylation and hypomethylation. As for the hypermethylation, DNA methylation of a promoter CGI is known to inactivate tumor-suppressor genes and promote cancer development, and the methylation accumulation levels of eight driver genes significantly increased according to the risk levels of the samples. However, its weaker association with cancer risk than methylation burden indicated that methylation burden is more directly involved in cancer risk. As for the hypomethylation, global hypomethylation is considered to be mainly due to hypomethylation of repetitive elements, and can lead to chromosomal instability, which are involved in carcinogenesis [29, 37]. The probes in the DNA methylation microarray used here are distributed mainly in gene promoters and CpG islands [38], and may be insufficient to analyze global hypomethylation. However, hypomethylation of repetitive elements has been shown to be reflected in methylation statuses of probes in a DNA methylation microarray [39].
The CDKN2A methylation level analyzed by pyrosequencing did not show a high correlation with that analyzed by DNA methylation microarray. This was because target CpG sites analyzed by pyrosequencing were located in 200 bp upstream region from its TSS and those by DNA methylation microarray were located in the 1st exon. The CpG sites analyzed by pyrosequencing for CDKN2A were located in 200 bp region upstream its TSS overlapping a CpG island, and were reported to be a valid target in terms of methylation-silencing [40]. Another issue that needs explanation is that CDH1 and SFRP5 methylation levels were lower in G3 than in G2. This could be due to the larger fraction of cells, for example, metaplastic cells, that need CDH1 and SFRP5 expression in G3 samples.
The accumulation level of driver gene methylation was calculated as an average methylation level of selected driver genes. This method at least enabled us to obtain a correlation between the accumulation level of driver gene methylation and the methylation level of a single marker gene. However, the impacts of individual driver genes on carcinogenesis are considered to be variable, and it might be better to take their relative importance into account. In addition, some driver genes may work in parallel and need to coexist, and others may work in upstream or downstream and be alternative. Since our current knowledge is far less than complete, we adopted here the simplest equation to avoid overfitting due to multiple hypotheses.
The limitation includes the small number of samples, especially G1 samples for genome-wide analysis (n = 4). Fortunately, variation within the G1 samples was small (r ≥ 0.981 for any two of the four samples), and we considered that the number is tolerable. At the same time, even within the same risk level, some samples may show extraordinary values, such as a few G2 samples with high methylation burden and high average methylation levels of the nine driver genes. These patients may develop gastric cancer in the future, and to minimize the influence of such samples, it would be better to have a large number of samples. In addition, in the subgroup analyses, we assessed the effects of age, gender, and gastric atrophy, but were unable to assess that of intestinal metaplasia (IM), an established risk factor of gastric cancer [41] because we did not have information on IM status for all the samples.
In conclusion, we revealed the mechanism of why the methylation level of a single marker gene can accurately predict cancer risk.
Data availability
DNA methylation data are available at the Gene Expression Omnibus (GEO) database with the accession number GSE226818.
References
Sakitani K, Nishizawa T, Arita M, et al. Early detection of gastric cancer after helicobacter pylori eradication due to endoscopic surveillance. Helicobacter. 2018;23:e12503.
In H, Sarkar S, Ward J, et al. Serum Pepsinogen as a Biomarker for Gastric Cancer in the United States: A Nested Case-Control Study Using the PLCO Cancer Screening Trial Data. Cancer Epidemiol Biomark Prev. 2022;31:1426–32.
Yanaoka K, Oka M, Yoshimura N, et al. Risk of gastric cancer in asymptomatic, middle-aged Japanese subjects based on serum pepsinogen and Helicobacter pylori antibody levels. Int J Cancer. 2008;123:917–26.
Yoshida T, Kato J, Inoue I, et al. Cancer development based on chronic active gastritis and resulting gastric atrophy as assessed by serum levels of pepsinogen and Helicobacter pylori antibody titer. Int J Cancer. 2014;134:1445–57.
Ando T, Yoshida T, Enomoto S, et al. DNA methylation of microRNA genes in gastric mucosae of gastric cancer patients: its possible involvement in the formation of epigenetic field defect. Int J Cancer. 2009;124:2367–74.
Maekita T, Nakazawa K, Mihara M, et al. High levels of aberrant DNA methylation in Helicobacter pylori-infected gastric mucosae and its possible association with gastric cancer risk. Clin Cancer Res. 2006;12:989–95.
Nanjo S, Asada K, Yamashita S, et al. Identification of gastric cancer risk markers that are informative in individuals with past H. pylori infection. Gastric Cancer. 2012;15:382–8.
Shin CM, Kim N, Park JH, et al. Prediction of the risk for gastric cancer using candidate methylation markers in the non-neoplastic gastric mucosae. J Pathol. 2012;226:654–65.
Yamashita S, Kishino T, Takahashi T, et al. Genetic and epigenetic alterations in normal tissues have differential impacts on cancer risk among tissues. Proc Natl Acad Sci USA. 2018;115:1328–33.
Asada K, Nakajima T, Shimazu T, et al. Demonstration of the usefulness of epigenetic cancer risk prediction by a multicentre prospective cohort study. Gut. 2015;64:388–96.
Maeda M, Nakajima T, Oda I, et al. High impact of methylation accumulation on metachronous gastric cancer: 5-year follow-up of a multicentre prospective cohort study. Gut. 2017;66:1721–3.
Baylin SB, Jones PA. A decade of exploring the cancer epigenome - biological and translational implications. Nat Rev Cancer. 2011;11:726–34.
Usui G, Matsusaka K, Mano Y, et al. DNA methylation and genetic aberrations in gastric cancer. Digestion. 2021;102:25–32.
Yoda Y, Takeshima H, Niwa T, et al. Integrated analysis of cancer-related pathways affected by genetic and epigenetic alterations in gastric cancer. Gastric Cancer. 2015;18:65–76.
Lee YC, Chiang TH, Chou CK, et al. Association between helicobacter pylori eradication and gastric cancer incidence: a systematic review and meta-analysis. Gastroenterology. 2016;150(1124):e5.
Niwa T, Tsukamoto T, Toyoda T, et al. Inflammatory processes triggered by Helicobacter pylori infection cause aberrant DNA methylation in gastric epithelial cells. Cancer Res. 2010;70:1430–40.
Sugimoto K, Ito T, Hulbert A, et al. DNA methylation genome-wide analysis in remnant and primary gastric cancers. Gastric Cancer. 2019;22:1109–20.
Takeshima H, Niwa T, Yamashita S, et al. TET repression and increased DNMT activity synergistically induce aberrant DNA methylation. J Clin Invest. 2020;130:5370–9.
Yamashita S, Nanjo S, Rehnberg E, et al. Distinct DNA methylation targets by aging and chronic inflammation: a pilot study using gastric mucosa infected with Helicobacter pylori. Clin Epig. 2019;11:191.
Wang F, Meng W, Wang B, et al. Helicobacter pylori-induced gastric inflammation and gastric cancer. Cancer Lett. 2014;345:196–202.
Take S, Mizuno M, Ishiki K, et al. The long-term risk of gastric cancer after the successful eradication of Helicobacter pylori. J Gastroenterol. 2011;46:318–24.
Mori G, Nakajima T, Asada K, et al. Incidence of and risk factors for metachronous gastric cancer after endoscopic resection and successful Helicobacter pylori eradication: results of a large-scale, multicenter cohort study in Japan. Gastric Cancer. 2016;19:911–8.
Kimura K, Takemoto T. An endoscopic recognition of the atrophic border and its significance in chronic gastritis. Endoscopy. 1969;1:87–97.
Ebrahimi V, Soleimanian A, Ebrahimi T, et al. Epigenetic modifications in gastric cancer: focus on DNA methylation. Gene. 2020;742:144577.
Padmanabhan N, Ushijima T, Tan P. How to stomach an epigenetic insult: the gastric cancer epigenome. Nat Rev Gastroenterol Hepatol. 2017;14:467–78.
Shigematsu Y, Niwa T, Yamashita S, et al. Identification of a DNA methylation marker that detects the presence of lymph node metastases of gastric cancers. Oncol Lett. 2012;4:268–74.
Iida N, Okuda Y, Ogasawara O, et al. MACON: a web tool for computing DNA methylation data obtained by the Illumina Infinium Human DNA methylation BeadArray. Epigenomics. 2018;10:249–58.
Sugai T, Yoshida M, Eizuka M, et al. Analysis of the DNA methylation level of cancer-related genes in colorectal cancer and the surrounding normal mucosa. Clin Epigenetics. 2017;9:55.
Eden A, Gaudet F, Waghmare A, et al. Chromosomal instability and tumors promoted by DNA hypomethylation. Science. 2003;300:455.
Wang K, Liu H, Hu Q, et al. Epigenetic regulation of aging: implications for interventions of aging and diseases. Sig Transduct Target Ther. 2022;7:374.
Ahuja N, Li Q, Mohan AL, et al. Aging and DNA methylation in colorectal mucosa and cancer. Cancer Res. 1998;58:5489–94.
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Ohata H, Kitauchi S, Yoshimura N, et al. Progression of chronic atrophic gastritis associated with Helicobacter pylori infection increases risk of gastric cancer. Int J Cancer. 2004;109:138–43.
Braghini MR, Lo Re O, Romito I, et al. Epigenetic remodelling in human hepatocellular carcinoma. J Exp Clin Cancer Res. 2022;41:107.
Cardoso MFS, Castelletti CHM, Lima-Filho JL, et al. Putative biomarkers for cervical cancer: SNVs, methylation and expression profiles. Mutat Res Rev Mutat Res. 2017;773:161–73.
Papadia C, Louwagie J, Del Rio P, et al. FOXE1 and SYNE1 genes hypermethylation panel as promising biomarker in colitis-associated colorectal neoplasia. Inflamm Bowel Dis. 2014;20:271–7.
Bae JM, Shin SH, Kwon HJ, et al. ALU and LINE-1 hypomethylations in multistep gastric carcinogenesis and their prognostic implications. Int J Cancer. 2012;131:1323–31.
Bibikova M, Barnes B, Tsan C, et al. High density DNA methylation array with single CpG site resolution. Genomics. 2011;98:288–95.
Zheng Y, Joyce BT, Liu L, et al. Prediction of genome-wide DNA methylation in repetitive elements. Nucleic Acids Res. 2017;45:8697–711.
Tramontano A, Boffo FL, Russo G, et al. Methylation of the Suppressor Gene p16INK4a: Mechanism and Consequences. Biomolecules. 2020;10:446.
Li D, Bautista MC, Jiang SF, et al. Risks and predictors of gastric adenocarcinoma in patients with gastric intestinal metaplasia and dysplasia: a population-based study. Am J Gastroenterol. 2016;111:1104–13.
Acknowledgements
The authors are grateful to Ms. Mika Wakabayashi for her technical assistance.
Funding
This study is funded by AMED (grant number 22ck0106552h9903).
Author information
Authors and Affiliations
Contributions
Conception and design: TI, HY, CT, and TU. Development of methodology: TI, HY, and CT. Acquisition of materials: TS, TA, TM, SA, IO, MK, and KM. Analysis and interpretation of data: TI, HY, CT, YL, HC, TS, and TU. Writing, review, and/or revision of the manuscript: TI, KS (Kiichi Sugimoto), KS (Kazuhiro Sakamoto), and TU.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1964 and later versions.
Consent to participate
Informed consent was obtained from all patients for being included in the study.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Irie, T., Yamada, H., Takeuchi, C. et al. The methylation level of a single cancer risk marker gene reflects methylation burden in gastric mucosa. Gastric Cancer 26, 667–676 (2023). https://doi.org/10.1007/s10120-023-01399-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-023-01399-w