Abstract
There is a certain belief among data science researchers and enthusiasts alike that clustering can be used to improve classification quality. Insofar as this belief is fairly uncontroversial, it is also very general and therefore produces a lot of confusion around the subject. There are many ways of using clustering in classification and it obviously cannot always improve the quality of predictions, so a question arises, in which scenarios exactly does it help? Since we were unable to find a rigorous study addressing this question, in this paper, we try to shed some light on the concept of using clustering for classification. To do so, we first put forward a framework for incorporating clustering as a method of feature extraction for classification. The framework is generic w.r.t. similarity measures, clustering algorithms, classifiers, and datasets and serves as a platform to answer ten essential questions regarding the studied subject. Each answer is formulated based on a separate experiment on 16 publicly available datasets, followed by an appropriate statistical analysis. After performing the experiments and analyzing the results separately, we discuss them from a global perspective and form general conclusions regarding using clustering as feature extraction for classification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
There seems to exist a certain belief among the data science community members, which says that data clustering can be used to improve the quality of classification [15, 32, 33]. The main idea of this theory is very straightforward: (1) cluster the training examples, (2) encode the clusters as new features, (3) train the model and use it for prediction. The intuition behind this approach is also quite appealing and can be summarized as follows. In classification, we assume that classes in the dataset reflect its structure (which is an assumption necessary for any classifier to be able to learn) all the while knowing that this reflection may not be perfect. In clustering, the goal is to discover the underlying structure of the data (by partitioning). Clearly, on some level these two methods can be considered as two sides of the same coin. It is therefore tempting to check whether discovering this inherent, “true” structure of the dataset using clustering could aid the process of classification. If the two structures (i.e., the one revealed by clustering and the one implied by classes) map perfectly to one another, this means that the dataset is trivial to classify. If not, then the attributes conveying this “hidden” structure of the data can lead to better generalization of the classification model.
This simple idea seems so attractive that it even crops up in popular science outside academia, e.g., in the form of blog posts written by data science practitioners [15]. However, we were unable to find any serious study that would consider this hypothesis in a methodical manner or even formalize this concept, even though many questions come to mind when considering generating clustering features for the purpose of classification.
-
How should this process exactly work?
-
How to encode clusters as features?
-
Does it really improve the classification quality and if so, under what conditions?
-
Do clustering-generated features work well with all classifiers?
-
What impact does the clustering algorithm have on the quality of predictions?
These are only some of the unanswered questions surrounding this idea which we aim to address in this study.
In this paper, we examine the idea of using data clustering as a way of extracting new features for the purpose of classification. To achieve this goal, we propose a framework which works by augmenting the dataset with new features encoding the clusters. The framework is generic w.r.t. similarity measures, clustering algorithms, classifiers, and datasets, and requires only that a similarity measure between data points exists. The proposed framework is then used to verify various hypotheses regarding the discussed process. We experiment with several cluster representations, clustering algorithms, classifiers, and distance measures, to see what impact each of these components has on classification quality. Delving deeper into clustering, we discuss two possible clustering scenarios: global, i.e., clustering regardless of classes, and local, i.e., clustering separately in each class. We also discuss the issue of overfitting by performing a sensitivity test with respect to the number of clusters. Furthermore, we examine how the discriminative power of each cluster impacts the classification performance. We dedicate a separate set of experiments to analyze the impact of various isolated dataset characteristics on the quality of predictions. Naturally, we also check how new features compare against the original ones in terms of their predictive capabilities. Finally, we test the discussed scenario in a semi-supervised setting to observe if it benefits from the additional information. All experiments are carried out on 16 publicly available datasets with appropriate statistical tests to firmly verify our findings.
In conclusion, the main contributions of this paper can be summarized as follows.
-
1.
We define, both formally and conceptually, a framework for using data clustering as a feature extraction method.
-
2.
We carry out an empirical study on using clustering-generated features for classification in which we consider the following issues:
-
(a)
which cluster representation works best,
-
(b)
what influence does the clustering algorithm have on classification quality,
-
(c)
what is the impact of using global/local clustering,
-
(d)
sensitivity w.r.t. the number of clusters,
-
(e)
does cluster purity w.r.t. class influence classification quality,
-
(f)
what is the impact of distance measures on the process,
-
(g)
how well does the process work in a semi-supervised setting,
-
(h)
can the obtained features replace the original ones or should they be used together,
-
(i)
does the described methodology work with every classifier,
-
(j)
what is the impact of various dataset characteristics on the discussed process.
-
(a)
-
3.
We perform various statistical tests and discuss the results in detail to form general conclusions regarding using clustering in classification problems.
The remainder of this paper is organized as follows. Section 2 gives the background of related work. In Sect. 3, we describe and define the framework on which our study is built. Section 4 explains the experimental setup. In Sect. 5, we experimentally evaluate each of the described issues and provide answers to the stated questions. In Sect. 6, we discuss our findings along with the broad view of using clustering-generated features in classification tasks. Finally, in Sect. 7, we conclude the paper and draw lines of future research.
2 Related work
As one of the key ingredients to a successfully trained classifier, feature engineering has been extensively studied over the recent decades [6, 30]. Although vast literature on this topic is available, the direct inspiration for the research conducted in this paper comes from a study on pattern-based classification by Cheng et al. [4, 5]. The authors propose the following classification framework: (a) mine discriminative patterns, (b) represent the data in the feature space of these patterns, (c) build a classifier on the new representation. Based on this framework, the authors then show that discriminative patterns are good candidates for new features, especially when dealing with complex decision boundaries, and that there is high correlation between a pattern’s discriminative power and its frequency. The link between this study and ours is that, similarly to patterns, clusters also represent global information, i.e., information available only through analyzing the dataset as a whole. So the initial question which sparked our research was whether the global information obtained through clustering can improve classification just as the global information obtained through frequent patterns.
The idea of using clustering in classification is not new. In fact, over the years it has been studied in 3 main contexts related to our work: prototype-based, semi-supervised, and feature extraction. Prototype-based classifiers can use it to select the prototypes according to which the predictions will be made. Hastie et al. [13] show how clusters can be used both directly for predictions and as an initializing step for more advanced methods. In both scenarios, they propose to cluster the training dataset separately for each class using the k-means algorithm. Then, in the first scenario, they propose to use the cluster centers together with their corresponding classes as the prototypes. When new examples arrive, they are assigned to the class indicated by the nearest prototype. In the second scenario, the authors propose to use the cluster centers as initial prototypes for the LVQ method proposed by Kohonen [16].
In the semi-supervised context, an approach has been proposed by Gan et al. [12] which aims at mapping the hidden structure of the data. This is achieved by clustering both training and test sets using the fuzzy c-means algorithm—a semi-supervised method. First, the combined training and test datasets are clustered, and a classifier is trained using only the training data. The clustering produces a membership degree for each unlabeled example to each of the classes. The unlabeled examples which have high certainty of belonging to a given class are then classified by the trained classifier. Subsequently, the most confidently predicted examples, together with their predicted labels, are added to the training set and the classifier is re-trained. This process is repeated until all unlabeled examples are labeled. This approach is very similar in its goal to our proposal, as both aim at mapping the underlying structure of the dataset; however, our method is fully unsupervised, while the proposal of Gan et al. is semi-supervised. Moreover, our method encodes the information in new features, while the authors’ approach encodes this information directly into a trained classifier.
The feature extraction context, which is the one most closely related to this study, can be further divided into two sub-categories: feature clustering and data clustering. The feature clustering approach is extensively used, e.g., to facilitate face recognition [7] or improve classification of proteins by clustering the amino acids which they are composed of [2]. This approach is similar to ours in a sense that it also relies on clustering and also generates features based on the representatives of the obtained clusters. However, it relies on manipulating the feature space, while our approach aims at mapping the structure hidden in the examples themselves. The data clustering approach relies on clustering of the training examples and encoding the clusters as new features, which is the exact approach we explore in this research. Alapati et al. [1] do this by adding a single new feature encoding the cluster to which each example belongs, while Srinivasa et al. [28] do this by selecting top k numerical features generated using fuzzy c-means algorithm. However, the scope of these studies is very limited and the former [1] contains a serious factual error, since the new feature is treated as a numerical attribute even though it contains categorical values.
Several other scenarios involving clustering with classification have been considered, e.g., clustering for relabeling of training examples [26], clustering for data partitioning in ensemble classification [3], clustering to generate weights for training examples [18], or clustering to address the problem of class imbalance [8, 14]. However, the only common factor between these methods and the one studied in this paper is the sole fact of using clustering together with classification, while all other details are different.
Despite the fact that the concept of using clustering together with classification is clearly not new, to the best of our knowledge, a rigorous study exploring its properties and soundly answering the major questions related to it has never been attempted. Filling this gap is the aim of this paper.
3 A framework for feature extraction using data clustering
The study conducted in this paper lies on the intersection between clustering, feature extraction, and classification, and its goal is to check whether using data clustering can enhance the quality of classification. In order to achieve this task and meaningfully answer the questions stated in Sect. 1, we put forward a generic framework which will allow us to alter the components of the process to test various hypotheses. In this section, we will describe this framework in detail, first, conceptually, and later, formally.
3.1 Conceptual description
The first step of the framework is to cluster the whole training dataset (without the class attribute) using a chosen clustering algorithm. We assume that the number of clusters is either determined by the algorithm automatically or provided explicitly as a parameter. For each resulting cluster, a representative is selected, e.g., a centroid. These representatives together with the original data points are then used to compute new features using a selected distance measure, one new feature per each cluster. Both old and new features constitute an augmented dataset which is then used to train a classifier, while the set of representatives is stored for transforming new, unlabeled examples into the new feature space. When a new unlabeled example arrives, its distance from each cluster representative is evaluated and used to compute the values of new features. Afterward, the unlabeled example is in the same feature space as the training examples, so the trained classifier can be used without any modifications. The whole process is summarized in Fig. 1.
Workflow illustrating the proposed procedure: (1) training examples are grouped into k clusters; (2a) new features based on cluster representatives are created and added to the existing features in the training examples; (2b) cluster representatives are saved for later; (3) the classifier undergoes training using the new training set; (4) a classification model is produced; (5) a new example arrives for classification; (6) distances to each cluster representative are computed and encoded as new features; (7) the modified example is passed to the classification model; (8) the model makes a prediction
3.2 Formal description
Let us now formalize the framework according to the conceptual description from previous section.
3.2.1 Preliminaries
Given a dataset of training examples \({\mathcal {X}}=\{\pmb {x_1}, \pmb {x_2}, \ldots , \pmb {x_n}\}\) and their corresponding classes (labels) \({\mathcal {Y}}=\{y_1, y_2, \ldots , y_n\}\), the task of a classifier is to predict the class \({\hat{y}}\) of each unlabeled example \(\pmb {{\hat{x}}}\) (example, for which the class is unknown). Every example \(\pmb {x_i}\) is a vector \((x_{i1}, x_{i2},\ldots ,x_{im})\) of the same length m, where each position j holds a value a given example has for a particular feature (attribute) \(F_j\in {\mathcal {F}}\).
The aim of feature extraction is to create a new set of features \(\mathcal {F'}\) based on the original feature set \({\mathcal {F}}\). Because feature extraction is applied on the training dataset, it has to produce a mapping function capable of transforming any future examples \(\pmb {x}\) to the same feature space. This is due to the fact that after the classifier has been trained, the training examples should not be changed (or else it would require a retraining of the classifier). In other words, new, unlabeled examples should not require the recalculation of the features in the training data. Although this constraint seems subtle, it has important consequences for our proposal since it is based on clustering.
Given a dataset \({\mathcal {X}}\), the aim of clustering is to identify groups (clusters) of examples, so that the examples within a particular cluster are more similar to one another than to the examples from other clusters. It is common for clustering algorithms to produce a representative for each cluster of equal dimensionality as the examples (e.g., centroid), which summarizes the cluster’s characteristics.
3.2.2 Framework definition
Given the above, the process examined in this paper is defined as follows. Given a dataset of training examples \({\mathcal {X}}=\{\pmb {x_1}, \pmb {x_2}, \ldots , \pmb {x_n}\}\) with features \({\mathcal {F}}=\{F_1,F_2,\ldots ,F_m\}\) and a distance measure \(\delta (\pmb {x_i}, \pmb {x_j})\) in the feature space, the examples are grouped together into k clusters using any clustering algorithm. If the algorithm does not produce cluster representatives, they are selected in post-processing as centroids (i.e., examples with the smallest average distance to all other examples in a given cluster). The number of clusters k can be either user-defined or computed by the algorithm. Then, k new features \(F_{j+1},F_{j+2},\ldots ,F_{j+k}\), one for each cluster representative \(\pmb {c_l}\in {\mathcal {C}}\), \(l=1..k\), are added to the training dataset, so that each training example \(\pmb {x_i}=(x_{i1},\ldots ,x_{im})\) becomes \(\pmb {x_i'}=(x_{i1},\ldots ,x_{im}, x_{im+1},\ldots ,x_{im+k})\), where \(x_{im+l}\) is calculated based on the distance \(\delta \) between example \(\pmb {x_i}\) and cluster representative \(\pmb {c_l}\), \(l=1..k\). The transformed set of training examples \({\mathcal {X}}'\) is then passed to the classifier for training. Afterward, each new unlabeled example \(\pmb {{\hat{x}}}\) has to be transformed into the same feature space as the training examples prior to being classified. This is done using the previously obtained cluster representatives, analogously to how training examples were transformed, so \(\pmb {{\hat{x}}}=\{{\hat{x}}_1,\ldots ,{\hat{x}}_m\}\) becomes \(\pmb {{\hat{x}}'}=\{{\hat{x}}_1,\ldots ,{\hat{x}}_m,{\hat{x}}_{m+1},\ldots ,{\hat{x}}_{m+k}\}\), where \({\hat{x}}_{m+l}\) is calculated based on \(\delta (\pmb {{\hat{x}}},\pmb {c_l})\), \(l=1..k\).
3.3 Example
The idea of putting forward a framework is to establish a flexible platform allowing to easily swap its parts to test various hypotheses. Given the above description and the illustration in Fig. 1, it is clear which parts of the process are generic, namely: the clustering algorithm, the distance measure, the encoding of new features, and the classifier. To better illustrate how all of these parts come together, let us analyze a sample instantiation of the framework.
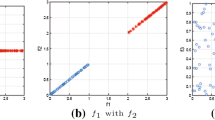
Consider a training dataset shown in Fig. 2a. The first step of the framework involves clustering of the training examples. For this purpose we will use the k-means algorithm with the Euclidean distance and the number of clusters \(k=2\). The result of this step is illustrated in Fig. 2b. In the second step, we encode the obtained clusters as new features. In this example, each new feature encodes the distance from a given training example to the cluster representative of a cluster corresponding with the feature. To keep the example clear we remove the original features and represent each example only in the new feature space. The result of this step is visualized in Fig. 2c. Afterward, a classifier can be trained on the transformed training dataset. For this purpose let us use simple logistic regression. The trained model is visualized in Fig. 2d. After training, we are ready to predict new examples. Let us consider an unlabeled example \(\pmb {{\hat{x}}}=(-1, 0.5)\). The example is presented in the original feature space together with cluster representatives in Fig. 2e. To use the trained model from Fig. 2d, first, we need to transform the new example into the new feature space by calculating its distance from each cluster representative and encoding this information into new features. This step is illustrated in Fig. 2f. Afterward, we can make a prediction which in this case is \({\hat{y}}=0\).
The above-described example instantiates the proposed framework in the following way: clusters are encoded as a distance from each example to each centroid; clustering is performed using the k-means algorithm; Euclidean metric is used as a distance measure; and logistic regression is used as a classifier. However, it is clear that each of these components could be easily swapped for a different one independently from other components, allowing us to experiment with various options in isolation from other factors.
4 Experimental setup
To test the hypothesis put forward in this study, i.e., whether clustering improves the quality of classification, we methodically answer the questions stated in the introduction:
-
1.
Which cluster representation works best? (Sect. 5.1)
-
2.
What influence does the clustering algorithm have on classification quality? (Sect. 5.2)
-
3.
What is the impact of using global/local clustering? (Sect. 5.3)
-
4.
How sensitive is this method w.r.t. the number of clusters? (Sect. 5.4)
-
5.
Does cluster purity w.r.t. class influence classification quality? (Sect. 5.5)
-
6.
What is the impact of distance measures on the process? (Sect. 5.6)
-
7.
Does the additional information stemming from semi-supervised learning help? (Sect. 5.7)
-
8.
Can the obtained features replace the original ones or should they be used together? (Sect. 5.8)
-
9.
Does the described methodology work with any classifier? (Sect. 5.9)
-
10.
Which dataset characteristics influence the described process and to what extent? (Sect. 5.10)
Each answer is formulated based on a dedicated experiment, followed by appropriate statistical tests (described in Sect. 4.5) and a discussion. All experiments are carried out using a common procedure described in Sect. 4.4 on 16 datasets described in Sect. 4.6. After addressing all of the stated questions, we summarize our findings and discuss the broad view of using clustering in classification in Sect. 6.
4.1 Clustering
In the majority of our experiments, we use one of two clustering algorithms: k-means [21] or affinity propagation [10]. The choice of these methods was dictated by two main reasons. The first reason follows directly from the requirements of our methodology, i.e., the algorithms have to produce cluster representatives. The second reason is that we wanted to test different approaches to determining the number of clusters. Affinity propagation detects the number of clusters automatically, whereas k-means requires this parameter to be given as an input. This way, one version makes the process conveniently parameterless, while the second one allows us to experiment with the number of clusters and check how this value influences the proposed method. The use of any other clustering algorithm will be reported directly in the description of a given experiment (see Sect. 5.2).
4.2 Classification
The main hypothesis of this study, i.e., does adding clustering-generated features improve the quality of classification, is tested on 8 different classifiers, namely: penalized multinomial regression, penalized discriminant analysis, Bayesian generalized linear model, decision trees, k-nearest neighbors, SVM with linear and RBF kernel, and random forest. The main idea behind selecting these classifiers was to test a diverse, representative group of methods which would cover a wide spectrum of possible options. This way, we can test whether clustering works in general or only with some particular families of classifiers. However, multiple classifiers are not needed when experimenting with the properties of the discussed process, since in such cases we are no longer comparing our results against the baseline, but rather comparing different versions of the same approach against each other. The results should therefore be general and independent of the classifier being used. That is why, in the majority of the experiments we rely on the SVM classifier with a linear kernel for its overall good performance and robustness, leaving all 8 classifiers only for testing the main hypothesis (see Sect. 5.9).
4.3 Distance measure
Another required component of our method is the distance measure \(\delta (x_i, x_j)\) between the examples. For this purpose, in the majority of our experiments we rely on the Euclidean distance. However, since the distance measure is yet another component of the proposed framework with a potential impact on the whole process, we isolate it in a separate experiment, in which we compare the Euclidean distance against the Mahalanobis distance (see Sect. 5.6).
4.4 Experimental procedure
The experimental procedure used to assess the validity of our hypotheses is defined as follows. Each dataset is divided into a training set and a holdout test set, each containing 50% of the data. The samples are stratified to ensure equal proportions of classes in each set. Each classifier is tuned and trained on the training set using fivefold cross-validation repeated 2 times and later tested on the holdout set with predictions evaluated using accuracy defined as:
The procedure covers both the baseline and our hypothesized approaches. When testing our approaches, we first cluster the training set and generate new features in training and test sets based on the cluster representatives according to the framework described in Sect. 3. Afterward, the augmented sets undergo the same train–test–evaluate procedure as the baseline. The whole procedure, summarized in Fig. 3, is repeated 10 times for two main reasons. The first one is obvious, as it allows us to average the results over random train/test splits to achieve a more reliable score for each approach. The second reason stems from the fact that some clustering algorithms (like k-means which we heavily rely on) are randomized and can therefore produce unstable results for the same data. Given the above, the results could in fact be averaged in another way, namely by additionally repeating the described process for each train/test split. This double-averaging scheme would address each randomization separately (train/test split and clustering). However, our preliminary experiments showed that the employed averaging scheme does not influence the results, so we decided to use the simpler single-averaging.
4.5 Statistical tests
After gathering the results, we perform a series of statistical tests which allow us to draw firm conclusions about our findings. To compare two approaches on each dataset, we use the t-test [29], and to compare multiple approaches on each dataset we rely on the ANOVA statistical test [9], and if it indicates significant differences between the classifiers, we follow it by a pairwise t-test [29] to assess the hierarchy among them. In every case, we also make sure the assumptions for the ANOVA test [9] are satisfied, namely we check if the values are normally distributed using Shapiro and Wilk test [27] and if the variances across groups are homogeneous using Levene test [19]. To compare two classifiers across all datasets, we use Wilcoxon signed ranks test [34], and to compare multiple classifiers across all datasets, we use the Friedman test [11], and if it reveals significant differences between the results, we follow it with a post hoc Nemenyi test [23] to establish a hierarchy between the classifiers. In all tests, we assume the significance level \(\alpha =0.05\).
4.6 Datasets
We used 16 numeric datasets with various numbers of classes, examples, and attributes. All of the datasets are publicly available through the UCI machine learning repository [20]. Table 1 presents the main characteristics of each dataset. The datasets were chosen so that they display a wide range of properties: the size varies between 150 and 20,000 examples; the number of features varies between 5 and 102; the number of classes varies between 2 and 48. Since in most of our experiments we used the number of clusters determined by affinity propagation, we include the average number of clusters into the dataset characteristics table. We report the average because the analyzed process was repeated several times and for each repeat, clustering was performed on a different train/test data split; therefore, the outcomes were slightly different.
All experiments were written in the R programming language [25] using the caret package [17] to unify the experimental procedure across different classifiers and datasets. The code used during the experiments along with the raw experimental results and the detailed results of statistical tests are publicly available at: https://github.com/MaciejPiernik/clustering-generated-features.
5 Experiments
5.1 Cluster representation
Before beginning to assess how various clustering algorithms, distance measures, classifiers, etc., influence the discussed process, first, we need to decide on the one component which has not yet been completely defined, namely cluster representation, in other words, how to encode the discovered clusters as new features. So far, we have only stated that this encoding is done based on the distances between the examples and cluster representatives. This, however, can be done in many different ways. In this experiment, we are going to test 5 methods of encoding clusters as features, which we refer to as: binary, binary distance, distance, inverse distance squared, and probability.
5.1.1 Binary representation
According to our framework, each feature corresponds to a certain cluster, so the most straightforward encoding is a one in which each feature encodes whether a given example \(\pmb {x_i}\) belongs to a cluster with a given representative \(\pmb {c_l}\) or not:
5.1.2 Binary distance representation
Since not all examples are in the same distance from the centers of their clusters, a natural extension of the binary representation is encoding the distance from a given example \(\pmb {x_i}\) to a given cluster center \(\pmb {c_l}\) if the example belongs to this cluster, and 0 otherwise:
5.1.3 Distance representation
Since for each example we can evaluate its distance from all cluster representatives, a further extension of the binary distance representation is to let go of the cluster membership altogether and simply encode each cluster as a distance from its representative \(\pmb {c_l}\) to each example \(\pmb {x_i}\): \(\pmb {x_{im+l}} = \delta (\pmb {x_i},\pmb {c_l})\).
5.1.4 Inverse distance squared representation
When assessing the importance of objects, a common approach is to use the inverse square of distance (e.g., in knn classifier). Although measuring the importance of examples is not our direct goal, it is tempting to check what would be the effect of using this method in our problem. Therefore, the fourth examined way of encoding a cluster is to record the squared inverse of the distance between its representative \(\pmb {c_l}\) from each example \(\pmb {x_i}\): \(\pmb {x_{im+l}} = 1/\delta (\pmb {x_i},\pmb {c_l})^2\).
5.1.5 Probability representation
Some clustering algorithms, e.g., expectation-maximization or fuzzy clustering methods, estimate a cluster membership probability for each example. Although based on distance, these estimates represent something different and are therefore interesting to include in the analysis, especially since they can be naturally used as new features without any additional processing. For any given example \(\pmb {x_i}\) and cluster representative \(\pmb {c_l}\), each new feature is encoded as: \(\pmb {x_{im+l}} = 1/\sum _{i=1}^{k}{\left( \frac{\delta (\pmb {x_i},\pmb {c_l})}{\delta (\pmb {x_i},\pmb {c_i})}\right) ^2}\), where k is the number of clusters.
The results of this experiment are presented in Fig. 4. The distance representation stands out as a clear winner, although the probability representation is not far behind, in one case (ionosphere) even prevailing over distance. However, at individual level, not all results are significant. The ANOVA test was unable to distinguish between the results of each approach for breast-cancer-wisconsin dataset. Moreover, the further pairwise t-test was also inconclusive for ecoli, iris, and image-segmentation dataset. For glass, pima-indians-diabetes, pendigits, spectrometer, and statlog-vehicle, the distance representation was significantly better than all other representations, while on the remaining datasets, it was only rivaled by the probability representation (with one exception of statlog-satimage, where it was also indistinguishable from the inverse distance squared representation).
Analyzing the results collectively, the superiority of the distance and probability representations is confirmed. The Friedman test and the post hoc Nemenyi test, results of which are illustrated in Fig. 5, reveal that the measured accuracies were not produced by chance and that, indeed, the distance and probability representations are both significantly better than the other representations. Although distance was also on average better than probability, the test revealed that this difference is not significant.
All of the above test results allow us to draw a clear conclusion that the distance representation is a better choice than the alternatives. The reason for its superiority is in most cases also rather straightforward. The binary, binary distance, and distance representations convey different amounts of information, with the last one conveying the most, and the first one—the least. It is very easy to convert the distance representation into binary distance and the binary distance into binary, but the reverse is impossible. This asymmetry of conversion can be viewed as a lossy data compression, and the experiments simply show that the information lost in this process is valuable. Regarding the inverse distance squared representation, the aim was to check whether looking at the features differently, namely as the importance of a given example w.r.t. a given cluster rather than its simple spatial orientation, would benefit the process. This hypothesis turned out to be wrong; however, we suspect that the importance of examples could be potentially useful as a way of weighting training examples. The probability, although collectively statistically indistinguishable from the distance representation, was ultimately significantly outperformed by it on several datasets, while the reverse was never true. Although we do not see a straightforward reason for this outcome, our educated guess is that, since probability by definition puts an emphasis on cluster membership, it focuses on single clusters, so the information is not as well distributed across all features as in the case of the distance representation. Given the above, we recommend relying on the distance representation as the default way of encoding clusters as new features and use this representation in the remainder of the experiments.
5.2 Comparison of clustering algorithms
The first major component of the analyzed framework is the clustering algorithm used to form groups of training examples. Although there are many options to choose from, we wanted to focus on two main characteristics: a) how it deals with the number of clusters and b) what is the shape of the produced clusters. As a result, we decided to compare three approaches along those axes. The first two are k-means and affinity propagation, which allow us to deal with the first characteristic (as already described in Sect. 4.1). Since these approaches tend to produce spherically shaped clusters, we selected spectral clustering as the third candidate, as it allows for discovering clusters of irregular shapes. Conveniently, all of these approaches produce cluster representatives, so there is no need for determining them separately.
Since the selected algorithms deal differently with the number of clusters, we had to make a small adjustment in the experimental procedure. Affinity propagation determines the number of clusters automatically and our experiments have shown that it produces significantly more clusters than we would normally select for k-means or spectral clustering using other methods like the gap statistic [31]. Since each cluster is encoded as a new feature, this poses a problem, namely how to isolate the influence of the clustering algorithm from the influence of the number of features on the classification quality. Here, we only want to measure the impact of the former, while the impact of the latter will be analyzed in Sect. 5.4. Therefore, for each dataset we first perform the experiment using affinity propagation and then use the detected number of clusters as an input for k-means and spectral clustering. The results of this experiment are presented in Fig. 6.
Looking at the results, there is not a single case in which one algorithm would produce noticeably better results than the others. This is only further confirmed through the ANOVA tests which were unable to find significant differences between the algorithms on any dataset. Analyzing the results collectively, we were still unable to find any regularities. The Friedman statistical test confirms this observation as it was unable to reject the null hypothesis which states that the differences between the results are the product of random chance.
The above analysis allows us to draw a firm conclusion that the clustering algorithm does not impact the outcome of the discussed process in a significant way, if at all. On the one hand, this is a positive outcome as it allows us to select any clustering algorithm (e.g., the most efficient) rather than the one which produces the best clusters. On the other hand, it begs the question whether the composition of clusters has any impact on the process or does not matter at all. We will address this question with a dedicated experiment in Sect. 5.5.
5.3 Clustering per class or global clustering?
The intuition behind the hypothesis stated in this study is that clustering of training examples regardless of their class could help generalization through the use of global information. We refer to this approach as global as it requires a “global look” at the dataset, without artificially dividing it into predefined categories. However, one could argue for an alternative approach in which clustering is performed per class. We are still adding some global information about distant objects’ similarity, however, with the additional potential benefit of modeling the space occupied by each class. We refer to this approach as local as it analyzes the dataset “locally” in each class. To verify which of these approaches is better, we compared them empirically.
To make the comparison meaningful, we have to ensure an equal number of clusters in both approaches, to make sure that the results solely rely on the generated clusters and not their quantity. This is the same issue we faced when comparing clustering algorithms in Sect. 5.2. To achieve this goal, the experiment is performed as follows. First, we cluster the dataset separately in each class using affinity propagation to automatically determine the number of clusters. Next, we perform the same experiment using the global approach using k-means with the number of clusters equal to the total number of clusters found in all classes by affinity propagation. This way both approaches generate the same number of clusters for each dataset but using different approach. Since in the previous experiment we showed that the selected clustering algorithm does not influence the outcome of the process we can be confident in using different algorithms (affinity propagation and k-means) for each approach. The results of this experiment are presented in Fig. 7.
At first glance, the results do not reveal a clear winner, although the global approach seems to work better in more cases than the local. Looking at individual cases, the differences become more apparent with the global approach dominating significantly in several cases. A t-test confirms this observation indicating a significant difference between the means of the approaches for wine, glass, vowel-context, iris, sonar.all, image-segmentation, ionosphere, pendigits, and statlog-satimage datasets, among which only one (pendigits) is in favor of the local approach.
Analyzing the results collectively, the Wilcoxon signed ranks test was unable to find a significant difference between these two approaches. We therefore cannot state with all certainty that clustering the whole dataset produces better results than clustering for each class separately. That being said, global clustering did significantly outperform local on eight datasets, while the opposite was only true in one case. Moreover, global clustering is also the more reasonable choice, as it ensures a more even spread of the cluster representatives, while local clustering can produce highly overlapping clusters and, as a result, redundant features. Finally, the global approach is also the more intuitive one, as it allows the algorithm to access the hidden structure of the dataset which may be invisible through the lens of the classes. Given the above, even though we cannot statistically state that global clustering works on average better than local clustering, we still deem it a superior choice.
5.4 Sensitivity test
So far, in each experiment the number of clusters was determined by affinity propagation and remained unchanged for all tested approaches to isolate the influence of a given component. In this experiment, in turn, we isolate the number of clusters parameter to measure what impact it has on the quality of classification. We do this by executing the experimental procedure with the k-means clustering algorithm for increasing numbers of clusters from 1 to 200. Notice that 200 clusters is an unreasonably large number for certain datasets as some of them do not even have this many examples. This number was picked on purpose to pinpoint a moment at which the added features start leading to model overfitting. To fully achieve this goal, in addition to measuring the test set accuracy we also report training set accuracy to check when exactly the model starts overfitting the data. The results of the experiment are presented in Fig. 8.
It is easy to observe a point of saturation for most datasets at which training accuracy starts to diverge from test accuracy when it reaches a plateau. This is particularly well exemplified by the plots for glass, pima-indians-diabetes, sonar.all, and spectrometer datasets. On the other hand, for datasets image-segmentation, optdigits, pendigits, and statlog-satimage this effect is fairly insignificant, although still observable. Since SVM is a reasonably robust algorithm, this effect is not as dramatic as one could expect; however, the interpretation of these results is still very clear—there is a certain number of clusters after which the model starts to get needlessly complex. Interestingly, judging by the vertical lines in each plot, affinity propagation seems to produce a reasonable number of clusters, usually around the point at which the model starts overfitting. The only significant exceptions from this case are optdigits and pendigits datasets; however, coincidentally they also exhibit the effect of overfitting in the smallest amount, so, apart from the models being needlessly complicated, training accuracy still reasonably approximates test accuracy in these two cases.
The main conclusion from this experiment is that the number of clusters has a significant impact on the quality of predictions. Furthermore, most of the datasets show a predictable link between the number of clusters and classification quality. Up until a certain point, the more clusters we create the better. Afterward, the model starts to overfit the data and gets needlessly complicated without further improving the quality of predictions. The experiments also demonstrate that the number of clusters produced by affinity propagation is usually a good compromise between how complicated the model is and how well is it able to predict. From our point of view, this is a very important finding as it confirms that our choice of affinity propagation as the source of the number of clusters for all clustering algorithms was appropriate.
5.5 Taking into account cluster purity
Since we have already established in our sensitivity experiment that the number of clusters has a clear influence on the quality of classification, let us now check whether the purity w.r.t. the class of the clusters themselves makes any difference. In order to do so, we will cluster the datasets into the number of clusters indicated by affinity propagation, encode the clusters as new features, and evaluate the quality of each new feature using Fisher score defined as:
where \(n_k\) is the number of examples in k-th class, \(\mu ^j_k\) and \(\sigma ^j_k\) are the mean and standard deviation of k-th class corresponding to j-th feature, while \(\mu ^j\) and \(\sigma ^j\) are the mean and standard deviation of the whole dataset corresponding to j-th feature. Next, we will add new features one by one in order of their increasing and decreasing quality to observe the effect they have on classification accuracy. The results of this experiment are presented in Fig. 9.
The results of measuring the impact of cluster purity w.r.t. class on classification. The plots illustrate how adding new features in order of their quality influences the accuracy. The “top” (light yellow) line represents adding new features from best to worst, the “bottom” (dark blue) line—from worst to best
Ultimately, the approach discussed in this study is based on selecting points in m-dimensional space (where m is the number of features), calculating the distances between all data points and the selected points, and encoding these distances as new features. The analyzed plots illustrate that the choice of these points matters and has a clear impact on the quality of classification. On the diagram, the yellow (lighter) line represents classification quality when adding new features according to their descending Fisher score, while the blue (darker) line represents the same in an ascending order. The lines necessarily meet at the end, since in both cases in the end all features are used for classification. A direct observation from these plots is that adding new features in order of descending Fisher score usually produces better results than adding them in reversed order. This, however, leads to a more general and more important observation that some points (clusters) hold more information than others from the classification perspective which, in turn, seems to indicate that the choice of these points should matter. This effect is displayed most prominently in the plots for breast-cancer-wisconsin, ionosphere, iris, pima-indians-diabetes, sonar.all, and wine datasets. An odd case is the statlog-satimage dataset, where adding features in the ascending order of their discriminative power produces better results most of the time. However, this can be easily explained by, e.g., a single feature which achieved a low Fisher score but was actually crucial from the classification point of view.
Another obvious observation from the plots is that the maximum accuracy is almost always at the end of each plot, i.e., when all features are being used. This suggests that all of the generated features are important (albeit to a varied extent) and contribute valuable information from the classification perspective. This observation also confirms the results from the sensitivity experiment, hinting that the number of clusters selected by affinity propagation indeed strikes a good balance from the under-/overfitting perspective.
In conclusion, the experiment suggests that not all clusters are equally valuable. The results clearly indicate that some clusters yield features of higher purity, and this difference has a noticeable impact on the quality of predictions.
5.6 The impact of distance measure
In Sect. 5.1, we have established that encoding clusters as a distance from training examples to cluster representatives is the best among the proposed representations. Still, there are various distance measures to choose from. So far, in all of the tested approaches we have been relying on the Euclidean distance which by its nature produces spherically shaped clusters. However, one often encounters correlations between features which produce non-spherical clusters, to which the Euclidean distance is completely insensitive. Therefore, in this experiment we would like to test just how big of an impact this particular characteristic has on the classification quality by comparing the Euclidean distance against the Mahalanobis distance. Mahalanobis distance effectively measures the distance from a given point to a group of points and is capable of tracking correlations of features within that group. Therefore, in our case its values will be adjusted according to the data distribution in each cluster allowing us to form non-spherically shaped clusters.
Both measures will be tested using linear SVM classifier and k-means clustering algorithm with equal number of clusters for each dataset according to the experimental procedure from Fig. 3. However, since k-means inherently relies on the Euclidean distance, we have to alter it slightly for the purpose of this experiment. The modification for the Mahalanobis distance is as follows. The first iteration of the algorithm is carried out in a regular fashion using the Euclidean measure. This allows us to form first, candidate groups of objects. Each next iteration is carried out using the Mahalanobis distance, evaluated for each object based on its distance from each cluster representative, taking into account the covariance matrix calculated based on the objects within a given cluster. Just as in regular k-means, cluster centers are recalculated after each iteration along with the covariance matrix of the objects in each cluster. The results of this experiment are presented in Fig. 10.
The results of the experiment do not reveal a superiority of one measure over the other; however, noticeable differences are observable in many cases. For datasets yeast, vowel-context, glass, iris, sonar.all, optdigits, pendigits, spectrometer, statlog-satimage, and statlog-vehicle, the plots clearly indicate significant differences between the applied measures. The Euclidean distance came out on top in 7 out of 10 of these cases, while the Mahalanobis distance prevailed in the remaining 3. This observation is statistically confirmed for each of these datasets using a t-test. Analyzing the results collectively, the Wilcoxon test was unable to differentiate between the two measures (by a large margin with \(p\text {-value}=0.86\)).
Although from a global perspective the results of this test seem inconclusive, the individual results clearly indicate that the applied measure can have a significant impact on the outcome of the process. The experiment therefore hints that the effectiveness of the measure depends on the dataset. This observation could be somewhat expected, as each dataset can have a different data distribution and correlations between features. We can therefore speculate that in some cases the Euclidean distance sufficiently maps the analyzed space, while the Mahalanobis distance makes it excessively complicated, and in other cases, the Euclidean distance is unable to track irregularly shaped data patterns which are detectable by Mahalanobis measure. In any case, obtaining the information about the shapes appearing within a dataset is the exact task of clustering, so we naturally cannot know this information in advance. Consequently, since the Euclidean distance proved to be sufficient more often than not, we use it in all of our remaining experiments.
5.7 Semi-supervised learning
Since the methodology discussed in this study creates new features regardless of the decision attribute, it is tempting to try it in a semi-supervised setting and compare it against a fully supervised one, since the transition from one to the other is very straightforward. We achieve this by checking whether clustering on both training and testing data produces better features than clustering only on the former. Apart from using both training and testing data for clustering, the methodology and experimental procedure remains unchanged. The only extra modification concerns the number of clusters for each approach as the additional data may lead to more clusters being detected. Since in Sect. 5.4 we have established that the number of clusters has a significant impact on the outcome of the process, we wanted to eliminate this factor from the experiment. Analogous to Sect. 5.3, we achieve this by first clustering each dataset on both training and testing sets for the semi-supervised approach using affinity propagation and then using the detected number of clusters as an input for the k-means algorithm in the supervised approach. Using two different clustering algorithms is not a problem, since in Sect. 5.2 we have established that the effect of this choice is insignificant. The results of the experiment are presented in Fig. 11.
The boxplots clearly show that there is virtually no difference in the quality of predictions between the supervised and semi-supervised scenario. A t-test for every dataset detected significant differences only in two cases—vowel-context and iris—one in favor of the supervised, while the other in favor of the semi-supervised approach. A further Wilcoxon test only confirms this lack of difference on a global scale as it was unable to differentiate between the approaches by a large margin (\(p\text {-value}=0.9\)).
One would expect that the additional information stemming from clustering both training and testing datasets would benefit the quality of predictions; however, the results clearly indicate that this is simply not the case. On the one hand, this result seems surprising as one would expect the additional information provided by the test set to be beneficial for the classifier. After all, more information allows us to model the structure of the data more accurately and form better clusters which are later used to produce new features. On the other hand, the comparison of clustering algorithms from Sect. 5.2 has already shown us that the clusters themselves are not that important. Why is it then that more information does not benefit our method? The answer to this paradox could lie in the way we encode clusters as new features, namely as distance from each data point to each cluster representative. What this result seems to suggest is that it is not about how accurately the cluster representatives are selected but how well they cover the dataset space. If true, this observation would be consistent with the observations from the feature quality and sensitivity experiments as they clearly suggested that the right number of clusters is more important than their exact positions.
5.8 Can clustering-generated features improve classification quality?
In the next two experiments, the main premise of this study is put to the test, namely do clustering-generated features improve classification quality or not. In this first experiment, we compare the original features against two alternative approaches: relying only the new features or combining them with the original ones. This comparison is carried out using the linear SVM classifier. The better of the two tested alternatives will be further evaluated in Sect. 5.9 on 7 other classifiers, allowing us to form conclusions regarding the utility of using clustering-generated features in classification in general. The results of the first comparison are presented in Fig. 12.
The boxplots reveal many notable differences in performance between the compared approaches. Only in 5 cases all three feature sets performed in a comparable way, namely for breast-cancer-wisconsin, ecoli, sonar.all, wine, and yeast datasets. Comparing the original and new features, one can notice a substantial variation in the results and none of the two seems to be working better than the other. However, when combined they produce an outcome which is more often than not equal or better than the two feature sets used separately.
The ANOVA statistical tests reveal significant differences for glass, vowel-context, iris, image-segmentation, ionosphere, optdigits, pendigits, spectrometer, statlog-satimage, and statlog-vehicle datasets. The pairwise t-tests show that in 6 out of these 10 cases using both features was significantly better than relying only on the original ones, while the opposite was not true in any case. In the same test using new features significantly improved the result exactly as many times as it has diminished it, i.e., in 5 cases.
Analyzing the results collectively, the Friedman statistic was unable to find significant differences between the results. We are therefore forced to conclude that clustering-generated features do not improve classification quality in a general sense. However, given the above discussion, we can clearly see that they can significantly improve classification accuracy in some cases while not diminishing the quality in other cases. We can also infer that clustering-generated features can be used as an alternative to the original features, albeit with some reservation as the results vary significantly. Consequently, we recommend using clustering-generated features in conjunction with the original ones as this way they can improve classification quality without adding unnecessary risk. The only obvious downside to this is the extended dimensionality of the dataset which can result in longer training.
5.9 Classifiers
So far we have been only testing our hypotheses on a single, linear SVM classifier. In this final experiment, we want to observe what impact does the analyzed process have on other classifiers. Since in the previous experiment we have established that augmenting original features with new ones works favorably to relying solely on the new ones, in this experiment we are only going to compare the original features against the augmented features. We evaluate the process on several different, linear and non-linear classifiers, namely: penalized multinomial regression (multinom), penalized discriminant analysis (pda), Bayesian generalized linear model (bayesglm), decision trees (rpart), k-nearest neighbors (knn), SVM with RBF kernel (svmRadial), random forest (rf). The results are presented in Figs. 13, 14, 15, 16, 17, 18, and 19.
The first thing that one notices when analyzing the results is that the differences are very subtle, with the exception of only a few cases. However, some of these differences appear consistently across all repetitions and are therefore significant. In the case of penalized multinomial regression, one can easily identify several datasets for which adding clustering-generated features was significantly beneficial to the quality of predictions without degrading the quality on other datasets. The same observations hold true for penalized discriminant analysis classifier. Boxplots of Bayesian generalized linear model classifier paint a similar picture, although in the case of image-segmentation dataset the accuracy is noticeably degraded after adding new features. Decision trees produced similar results, although noticeable quality degradation can now be observed in two datasets: glass and image-segmentation. Adding the clustering-generated features proved to be detrimental to the accuracy in the case of the k-nearest neighbors classifier, where it seems to have helped only in two cases (ionosphere and vowel-context) but harmed the quality on at least 7 datasets. Interestingly, swapping the kernel in SVM from linear to RBF dramatically changed the effect of the analyzed process on classification quality. While adding clustering-generated features generally helped with the linear kernel, it seems to have no particular effect in case of the RBF kernel, except for two datasets (optdigits and pendigits), where it significantly degraded the quality of predictions. The worst result recorded was on the random forest classifier, where new features did not improve the quality of predictions on any dataset, but degraded the accuracy in at least 7 cases.
The t-tests for each dataset on each classifier only confirm the above observations but do not provide any additional insight. The clustering-generated features significantly improved the quality on: 6 datasets for multinom classifier, 7 datasets for pda classifier, 5 datasets for bayesglm classifier, 6 datasets for rpart classifier, and 2 datasets for knn classifier. However, they also significantly degraded the quality on: 1 dataset for rpart classifier, 7 datasets for knn classifier, 2 datasets for svmRadial classifier, and 4 datasets for rf classifier. The additional Wilcoxon signed ranks test was unable to distinguish between the two sets of features for any of the discussed classifiers, which forces us to conclude that from a global perspective, the discussed process on average does not influence the quality of predictions. However, given the above discussion, we can see that it can have a huge and consistent impact on the quality of predictions, albeit, depending on the classifier. For instance, it is uncontroversial to say that clustering-generated features seem to be a safe option when dealing with linear models, as adding them did not degrade classification quality on any dataset and was significantly beneficial in many cases. It is also safe to say that the discussed methodology should not be used in conjunction with the knn classifier as it can significantly harm the quality of predictions.
The results also lead to an interesting observation in terms of datasets as they seem to indicate that some datasets are more easily impacted by adding new features than others. In particular, the vowel-context, optdigits, pendigits, and statlog-satimage datasets produce significantly different results in nearly every case. However, and this is the most intriguing part, this effect swings between significant improvement and significant degradation depending on the classifier. The general trend remains unchanged, i.e., for linear models the datasets benefit when adding new features, while the nonlinear models are more often than not harmed by the addition of new features, but the reason for this pattern remains unclear.
In conclusion, we can state that clustering-generated features can improve classification quality, although not with every classifier. In general, the discussed methodology is beneficial for linear models, however, should be used with caution for nonlinear learners. Moreover, clustering-generated features are particularly harmful when used with k-nearest neighbors and random forest classifiers.
5.10 Dataset characteristics
Accuracies for original features and clustering-generated features for datasets with varying characteristics. Each point represents a different dataset generated with the parameter value given on the x axis, the regression lines represent the LOESS curves, while the bands represent their corresponding \(95\%\) confidence intervals
Many of the experiments conducted so far indicate that the results are highly dependent on the characteristics of the analyzed data. That is why, in this section, we aim at exploring how each dataset characteristic influences the performance of clustering-generated features compared to the original features. Although in Sect. 5.8 we have established that augmenting the original features produces in general better results than relying solely on the new ones, in this experiment, we compare the original features only against the new features to be able to focus on their properties in isolation. To facilitate a meaningful comparison, we use a dataset generator (from scikit-learn package [24]) which allows us to compare both approaches in various scenarios by changing several distinct data characteristics in a precisely controlled manner and separation from each other. We chose 5 common characteristics to experiment with: number of features, number of classes, number of clusters per class (i.e., number of distributions the data points are drawn from for each class), class distribution, and dataset difficulty (measured by class separation—the higher the separation the easier the dataset). To better capture the influence of each characteristic, we generate a range of datasets for each parameter by gradually changing its value. We rely on the experimental procedure presented in Fig. 3, but the whole process is executed for each parameter value in a specified range and each repeat generates a new dataset. Also, due to more randomness, we increased the number of repeats to 20. For each characteristic, we selected a wide range of parameter values to better illustrate the whole spectrum of possible outcomes. The parameters in each experiment change as follows, with the remaining parameters set to default:
-
number of features: 2, 3, ..., 100 [default: 10]
-
number of classes: 2, 3, ..., 50 [default: 2]
-
number of clusters per class: 1, 2, ..., 50 [default: 1]
-
class distribution: 1:39, 2:38, ..., 19:21, 1:1 [default: 1:1]
-
difficulty (class separation): 1.5, 1.4, ..., 0.1, 0 [default: 0.7].
The maximum value of class separation (1.5) was picked experimentally, as higher values produced trivial datasets which did not add any further insight into the analysis. In most of the experiments, we used affinity propagation with automatically determined number of clusters, with one exception discussed below. The results of this experiment are presented in Fig. 20.
The results reveal some very interesting properties of the analyzed method. Figure 20a clearly shows that the dimensionality of the dataset has a major impact on the quality of clustering-generated features. It also influences the quality of predictions on original features, but to a much lesser extent. However, this result does not paint the whole picture. Since we chose affinity propagation as the clustering method, the number of clusters was selected automatically. Interestingly, regardless of the number of features, the number of clusters was relatively stable between 15 and 25 for each dataset. This gave the clustering-generated features an advantage up until 20 features, while in higher dimensions the advantage was reversed. To address this issue, we re-ran the experiment with k-means and the number of clusters equal to the dimensionality of the dataset. This way, the dimensionality of the clustering-generated features is equal to the dimensionality of the original data. The result presented in Fig. 20b paints a very different picture—the outcome is to some extent reversed. These two results further amplify our main finding that the number of clusters is the key to the performance of the clustering-generated features.
Figure 20c, d shows that varying the number of classes and the number of distributions within each class has a major impact on the quality of predictions, albeit not much different than on original features. These results only indicate that the clustering-generated features do not introduce any additional robustness against these factors, nor do they hinge the performance with regard to these two class characteristics. A similar conclusion can be drawn with respect to class distribution, as illustrated in Fig. 20e, with one additional observation. On average, clustering-generated features seem to produce slightly more stable and better results with the increased balance of the class distribution.
The true highlight of the dataset characteristics analysis is the result of the dataset difficulty experiment presented in Fig. 20f. The plot clearly illustrates that clustering-generated features are much better at predicting difficult to distinguish classes than the original features, without hinging the performance for easier datasets.
In conclusion, the dataset characteristics analysis reveals two main findings. Firstly, it amplifies the importance of the number of clusters as the key to the performance of clustering-generated features. Secondly, it reveals that clustering-generated features are well suited for dealing with difficult problems, i.e., datasets with difficult to distinguish classes. However, it is very important to note that the second point holds only given that the first point is fulfilled, i.e., there is a sufficient number of clusters.
6 Discussion of the results
Let us now summarize the main findings of our study and discuss them in a broader context. In this paper, we formulated ten questions concerning using clustering as a means for feature extraction which we deemed important and consequential. The results of our experiments allow us to answer these questions, provoke other questions, and also shed some light on the general idea of using clustering for classification purposes.
We start the discussion by answering the fundamental question: does using clustering in conjunction with classification always improve the quality predictions? The answer to this question is unambiguously negative. Sometimes, clustering a dataset prior to classification improves the accuracy, and sometimes it does not. This outcome is dependent both on the dataset and the classification algorithm being used. The results of this study show that the improvement in classification accuracy concerns mainly linear classifiers, namely support vector machine, penalized multinomial regression, penalized discriminant analysis, and Bayesian generalized linear model. A possible explanation of this observation is that clustering-generated features give linear classifiers an access to more variance in the data (by introducing nonlinear feature transformations), thus improving the bias-variance balance of a given model. In case of nonlinear classifiers, clustering-generated features are much less beneficial and in some cases even degrade the quality of predictions, most prominently for k-nearest neighbors and random forest classifiers. This difference between linear and nonlinear models is very clearly illustrated on support vector machine, where going from a linear kernel to RBF kernel drastically hurts the outcome, from mostly improving the quality in the case of the former, to no improvement and sometimes degrading in the case of the latter. This can be explained by the fact that nonlinear models already have access to higher-dimensional feature spaces (infinitely dimensional in the case of the RBF kernel), so in these instances clustering-generated features do not introduce any new information. These observations confirm the findings of Trivedi et al. [32], where the authors also observed that clustering improved the quality of logistic regression, but was much less beneficial for random forest.
Another unambiguous and intuitive conclusion that follows from our study concerns cluster representation. The results of the experiment in Sect. 5.1 clearly demonstrate that constructing clustering-generated features as distances from cluster centers or cluster membership probabilities provide significantly better results than the analyzed alternative representations, most importantly, the binary representation, which only encodes cluster membership of each example. This observation can be easily explained by the fact that distance representation contains exactly the same information as binary representation and adds some additional information on top of it, which is clearly important from the classification perspective. A similar case can be made for cluster membership probability as it is based on distance but does not introduce any obvious information loss. This is an interesting finding in the context of using frequent patterns for classification, studied by Cheng et al. [5], which was the inspiration for our study. In this work, the authors show that encoding the information about frequent patterns as new (binary) features significantly improved the quality of classification. Our results regarding representations suggest that adding the information about the distance between frequent patterns and the examples could improve the quality of predictions even further. Of course, one would first have to devise such a measure; however, given the potential gain in quality, we consider it to be a very promising area of new research.
The final conclusion which we find uncontroversial concerns the clustering methodology, i.e., whether clustering should be done separately in each class (local approach) or for the whole dataset at once (global approach). It turns out that clustering globally often leads to much better results than local clustering. This result aligns with our intuition that looking at the dataset as a whole allows global clustering to access more information by unveiling the hidden structure of the dataset rather than being confined to predefined classes.
In addition to the above-described intuitive conclusions, our study also reveals many unexpected outcomes, some of which even standing in contradiction to one another. Firstly, the experiment in Sect. 5.2 suggests that there is no difference in classification quality when using different clustering algorithms. This surprising result leads to a bizarre conclusion that clusters themselves (their contents and shapes) have no impact on the quality of predictions. This observation is only amplified by the results from Sect. 5.7, where using semi-supervised learning to generate clustering features did not improve the classification accuracy whatsoever. So not only is the clustering algorithm irrelevant, but adding more relevant information into the process also does not help. What is even more intriguing is the fact that both of these results seem to stand in contradiction to the results from Sect. 5.5 which clearly indicate that there is a difference in the quality of features produced by the clusters which is then reflected in the quality of predictions. We think that the answer to this paradox is hinted by the results of the sensitivity test from Sect. 5.4 and the dataset characteristics analysis from Sect. 5.10. The sensitivity analysis shows that adding new clustering-generated features improves the accuracy to a certain point, but once enough clusters have been generated, adding more does not improve the quality further. Similarly, the dataset characteristics analysis reveals that clustering-generated features are well suited even for datasets with difficult to distinguish classes, however, only granted that the right number of clusters is selected—if not, the new features can even be detrimental to the quality of predictions. What all of these contradictory results seem to suggest is that the clusters themselves do not matter. What really matters is the number of clusters. This observation somewhat explains the work of Trivedi et al. [32], which, given the fact that they rely on k-means with a relatively low value of k, we suspect could yield much better results by simply using more clusters.
The conclusion that the character of clusters is irrelevant but the number of clusters matters is rather difficult to explain, unless we entertain the possibility that it is not clustering that drives the improvement and that in the discussed process clustering is not even necessary. After all, given the fact that the process relies on adding new features as distances from cluster representatives, all that the clustering algorithm really does is finding points in space. The process then proceeds to encode new features as distances from these points, but the points themselves could in principle be generated by any other method. This readily explains the fact that neither the clustering algorithm nor the clustering of both training and testing datasets had any impact on the quality of later predictions. Ultimately, the features generated by the process are simply nonlinear combinations of the existing features. In our experiments, this nonlinear combination was done according to the Euclidean distance; however, different measures would naturally produce different feature combinations. This is consistent with the observations taken from the experiment involving the Mahalanobis distance from Sect. 5.6, where the results varied significantly in comparison with the Euclidean distance from one dataset to another.
This line of thought leads to a very interesting question, namely is there any merit to using clustering as a way of creating new features or could the same result be achieved by simply selecting random points in space? Answering this question constitutes a fascinating new area of research, one which could potentially change the way we think about using clustering in supervised learning problems.
7 Conclusions
In this paper, we explored the idea of using data clustering as a way of extracting new features with the aim of improving the quality of classification. To achieve this goal, we proposed a framework which augments every dataset with clustering-generated features. The framework is generic w.r.t. similarity measures, clustering algorithms, classifiers, and datasets, and requires only that a similarity measure between data points exists. The framework was then used to answer several pressing questions regarding using clustering as a means of extracting new features. Each answer was supported with a separate experiment on 16 publicly available datasets, followed by a series of appropriate statistical tests.
The results of the study show that adding clustering-generated features can improve the quality of classification, albeit only when done in a specific way and not for all classifiers. Encoding clusters as features using distance from the examples to cluster representatives or cluster membership probability is significantly better than merely relying on their cluster membership, and the distance measure used produces significantly different results for different datasets. Moreover, adding the generated features to the existing ones rather than replacing them altogether is a safer option which yields more improvements without degrading the quality in other cases. As for the classifiers, the methodology seems to work better with linear classifiers, sometimes improving but never degrading the quality of predictions, but should not be used with nearest neighbors and random forest approaches. Finally, clusters generate features of varying quality which predictably impacts the quality of predictions; however, the character of the clusters themselves does not matter (even in a semi-supervised setting when clustering both training and testing sets at once), as long as there is enough of them.
Finally, this study opens several new doors of potential scientific inquiry. As adding clustering-generated features in a way described by our methodology is a generic approach, it would be interesting to see what results would this method yield with complex data types, e.g., sequential data, graphs, etc., and compare them with features generated using domain-specific methods. Furthermore, since we focused on the classification task, it would also be interesting to test the discussed approach in other machine learning and data analysis scenarios, both supervised and unsupervised. Although using clustering-generated features for clustering may seem redundant, it could prove useful in complex data types, in particular when dealing with multiview clustering [35] if each view is of complex, potentially different data type. In this scenario, relying on the proposed methodology for each view separately would allow for using existing multiview clustering approaches working on regular features [22]. On another front, the discussion of the results provokes many new questions related to the nature of encoding clusters as new features. Since in the discussed methodology, it is achieved based on the distances from cluster representatives to examples, it inherently reduces the role of clustering to an elaborate way of selecting points. The main question here is, are clustering-generated points in any way better than randomly or uniformly distributed points, and if so—why?
References
Alapati YK, Sindhu K (2016) Combining clustering with classification a technique to improve classification accuracy bibtex. Int J Comput Sci Eng 5(6):336–338
Bonet I, Saeys Y, Ábalo RG, García MM, Sanchez R, Van de Peer Y (2006) Feature extraction using clustering of protein. In: Martínez-Trinidad JF, Carrasco Ochoa JA, Kittler J (eds) Progress in pattern recognition. Image analysis and applications, Springer, Berlin, pp 614–623
Chakraborty T (2017) Ec3: Combining clustering and classification for ensemble learning. In: 2017 IEEE international conference on data mining (ICDM), pp 781–786
Cheng H, Yan X, Han J, Hsu CW (2007) Discriminative frequent pattern analysis for effective classification. In: Proceedings of the 2007 IEEE 23rd international conference on data engineering (ICDE ’07). IEEE Computer Society, Washington, DC, USA, pp 716–725
Cheng H, Yan X, Han J, Yu PS (2008) Direct discriminative pattern mining for effective classification. In: Proceedings of the 2008 IEEE 24th international conference on data engineering (ICDE ’08). IEEE Computer Society, Washington, DC, USA, pp 169–178
Dash M, Liu H (1997) Feature selection for classification. Intell Data Anal 1:131–156
El Ferchichi S, Zidi S, Laabidi K, Ksouri M, Maouche S (2011) A new feature extraction method based on clustering for face recognition. In: Iliadis L, Jayne C (eds) Engineering applications of neural networks. Springer, Berlin, pp 247–253
Fernández A, García S, Herrera F, Chawla NV (2018) Smote for learning from imbalanced data: progress and challenges, marking the 15-year anniversary. J Artif Int Res 61(1):863–905
Fisher RA (1921) On the probable error of a coefficient of correlation deduced from a small sample. Metron 1:3–32
Frey BJ, Dueck D (2007) Clustering by passing messages between data points. Science 315:2007
Friedman M (1937) The use of ranks to avoid the assumption of normality implicit in the analysis of variance. J Am Stat Assoc 32(200):675–701
Gan H, Sang N, Huang R, Tong X, Dan Z (2013) Using clustering analysis to improve semi-supervised classification. Neurocomputing 101:290–298
Hastie T, Tibshirani R, Friedman J (2001) The elements of statistical learning. Springer series in statistics, Springer, New York
Jo T, Japkowicz N (2004) Class imbalances versus small disjuncts. ACM SIGKDD Explor Newslett 6(1):40–49
Khan M (2017) Kmeans clustering for classification. https://towardsdatascience.com/kmeans-clustering-for-classification-74b992405d0a
Kohonen T (1989) Self-organization and associative memory, 3rd edn. Springer, Berlin
Kuhn M (2009) The caret package. http://topepo.github.io/caret/
Lango M, Brzezinski D, , Stefanowski J (2018) Imweights: Classifying imbalance data using local and neighborhood information. In: Proceedings of the 2nd international workshop on learning with imbalanced domains: theory and applications (LIDTA 2018), Dublin, Ireland, September 10–14, 2018
Levene H (1960) Robust tests for equality of variances. Stanford University Press, Stanford, pp 278–292
Lichman M (2013) UCI machine learning repository. http://archive.ics.uci.edu/ml
Lloyd SP (1982) Least squares quantization in PCM. IEEE Trans Inf Theory 28:129–137
Ma J, Zhang Y, Zhang L (2021) Discriminative subspace matrix factorization for multiview data clustering. Pattern Recogn 111:107676
Nemenyi P (1963) Distribution-free multiple comparisons. Ph.D. thesis, Princeton University, New Jersey, USA
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V, Vanderplas J, Passos A, Cournapeau D, Brucher M, Perrot M, Duchesnay E (2011) Scikit-learn: machine learning in Python. J Mach Learn Res 12:2825–2830
R Development Core Team (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org, ISBN 3-900051-07-0
Sa A, Sulaiman N, Mustapha A, Mustapha N (2009) K-means clustering to improve the accuracy of decision tree response classification. Inf Technol J 8:1256–1262
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality (complete samples). Biometrika 52(3/4):591–611
Srinivasa KG, Venugopal KR, Patnaik LM (2006) Feature extraction using fuzzy c-means clustering for data mining systems. Int J Comput Sci Netw Secur 6(3A):230–236
STUDENT (1908) The probable error of a mean. Biometrika 6(1):1–25
Tang J, Alelyani S, Liu H (2014) Feature selection for classification: a review. CRC Press, pp 37–64
Tibshirani R, Walther G, Hastie T (2001) Estimating the number of clusters in a dataset via the gap statistic. J R Stat Soc Ser B (Stat Methodol) 63(2):411–423
Trivedi S, Pardos ZA, Heffernan NT (2015) The utility of clustering in prediction tasks. CoRR arXiv:1509.06163
Tsai CF, Lin WY, Hong ZF, Hsieh CY (2011) Distance-based features in pattern classification. EURASIP J Adv Signal Process 1:62
Wilcoxon F (1945) Individual comparisons by ranking methods. Biom Bull 1(6):80–83
Yang Y, Wang H (2018) Multi-view clustering: a survey. Big Data Mini Anal 1(2):83–107
Acknowledgements
The authors would like to thank Dariusz Brzezinski for insightful comments on a draft of this paper. The authors are also grateful to the reviewers whose suggestions greatly improved the quality of the paper. This research is partly funded by the Polish National Science Center under Grant No. DEC-2015/19/B/ST6/02637.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Piernik, M., Morzy, T. A study on using data clustering for feature extraction to improve the quality of classification. Knowl Inf Syst 63, 1771–1805 (2021). https://doi.org/10.1007/s10115-021-01572-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10115-021-01572-6