Abstract
Purpose
To investigate the role of E. coli virulence-associated genes (VAGs) in predicting urinary tract infection (UTI) as the source of bacteremia in two distinct hospital populations, one with a large general catchment area and one dominated by referrals.
Methods
E. coli bacteremias identified at Department of Clinical Microbiology (DCM), Hvidovre Hospital and DCM, Rigshospitalet in the Capital Region of Denmark from October to December 2018. Using whole genome sequencing (WGS), we identified 358 VAGs from 224 E. coli bacteremia. For predictive analysis, VAGs were paired with clinical source of UTI from local bacteremia databases.
Results
VAGs strongly predicting of UTI as primary infection source of bacteremia were primarily found within the pap gene family. papX (PPV 96%, sensitivity 54%) and papGII (PPV 93%, sensitivity 56%) were found highly predictive, but showed low sensitivities. The strength of VAG predictions of UTI as source varied significantly between the two hospital populations. VAGs had weaker predictions in the tertiary referral center (Rigshospitalet), a disparity likely stemming from differences in patient population and department specialization.
Conclusion
WGS data was used to predict the primary source of E. coli bacteremia and is an attempt on a new and different type of infection source identification. Genomic data showed potential to be utilized to predict the primary source of infection; however, discrepancy between the best performing profile of VAGs between acute care hospitals and tertiary hospitals makes it difficult to implement in clinical practice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Escherichia coli is a commensal in the human gut. Certain strains of E. coli can cause infections in humans, including urinary tract infections (UTI), intraabdominal infections, and bacteremia. E. coli is the leading cause of bacteremia, with a 30-day mortality rate of nearly 10% [1,2,3,4]. In high-income countries, more than half of E. coli bacteremias originate from the urogenital tract [5].
UTI is a common infection defined as the presence of typical symptoms from the urinary tract and bacteriuria (presence of a significant amount of uropathogenic bacteria in urine) [6]. UTIs caused by E. coli account for approximately 75% of all UTIs [7]. Important risk factors for community-acquired UTI include female sex, age, immunosuppression, diabetes, urological abnormalities, and a history of previous UTIs [8, 9].
Identification of the primary infection source of E. coli bacteremia is crucial for various reasons. Most importantly, determining the source can guide more precise and effective treatment strategies like targeted antimicrobial therapy, length of treatment, and/or surgical interventions. Misidentifying or delaying the identification of the primary infection source could increase the risk of complications and mortality [10,11,12].
Whole genome sequencing (WGS) has revolutionized the study of bacterial infections, providing a comprehensive picture of the genetic makeup of a bacterial species. Virulence-associated genes (VAGs) are genes associated with bacterial pathogenesis, and their identification is crucial in understanding the mechanisms used by bacteria [13].
We aim to assess if information on specific bacterial VAGs could help to predict the source of infection of E. coli bacteremias and improve treatment decisions. The study will contribute to a better understanding of which VAGs play a part in bacteremia and can become an important tool for the clinicians to identify the primary source of infection in E. coli bacteremia.
Materials and methods
Isolate collection
The study was conducted in the Capital Region of Denmark at the Department of Clinical Microbiology (DCM) at Hvidovre Hospital (DCM-1) and Rigshospitalet (DCM-2). E. coli bacteremias identified from October to December 2018 were consecutively included in the study. Only monomicrobial bacteremias were included and only one positive culture was included per patient.
Blood culture bottles were incubated in the Bactec blood culture system (Bactec, BD Diagnostics, NJ, USA). Identification of bacterial species was done using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) analysis (Bruker Daltonics, Germany). Included blood culture isolates were stored at − 80 °C.
Bacteremia database and hospital descriptions
Local bacteremia databases stored data on acquisition (hospital- or community-acquired bacteremia), sex, and source of bacteremia. Clinical microbiology specialists or medical doctors in clinical microbiology training collected data prospectively for each bacteremia case. Primary bacteremia infection source was determined as UTI, non-UTI, or unknown source through medical journal reviews using clinical data, radiological data, and dialogue with clinicians. Patients with a positive blood culture taken within 48 h of admission were classified as community-acquired. Positive blood cultures taken later than 48 h of admission or blood cultures from patients readmitted within 48 h of discharge were classified as hospital-acquired.
The DCM-1 provides services for five secondary acute care referral hospitals, collectively offering over 1300 beds and serving a catchment area of approximately 1.1 million inhabitants. The hospitals covered by DCM-1 do not contain departments for hematology, oncology, or urology. Consequently, they do not specialize in the care of immunocompromised patients, transplant recipients, or patients undergoing or experiencing complications from urological surgery.
Rigshospitalet is the most specialized tertiary referral hospital in Denmark and contains departments that specialize in handling immunocompromised patients as they cover departments of hematology, oncology, rheumatology, neonatology, intensive care units, in addition to transplant patients. The DCM-2 only provides services for Rigshospitalet which has a total capacity of approximately 1100 beds and has no regular catchment area.
Due to the very different patient populations and medical specialties serviced by DCM-1 and DCM-2, we decided to analyze the two isolate populations separately. This decision mitigates potential confounders introduced by pooling data and allows for more precise, context-specific insights into the VAGs of E. coli bacteremia.
Whole genome sequencing, virulence-associated genes, and bacterial typing
E. coli isolates were sequenced using short-read WGS on the Illumina system at the Department of Genomic Medicine and DCM-2. Genome libraries were prepared using the NexteraXT kit and were sequenced using an Illumina NextSeq. The raw output fastq files are stored on a High-Performance Computing (HPC) cluster service by the National Life Science Supercomputing Center – Computerome at DTU and UCPH. From here, the raw fastq files undergo a quality control and quality assurance test using fastqc. Fastqc checks for per base sequence, quality, per base GC content, N content, as well as the sequence length distribution, kmer content, and for overrepresented sequences. Per base sequence quality > 30 across > 140 bases of each read, quality scores > 36 for > 90% of reads, per base sequence content 25% for each base across positions 20–140 in each read, and a distribution of per sequence GC content with median of 52% and standard deviation of 5% as expected for E. coli strains. A strict cutoff for number of reads required to obtain > 50 × coverage was also enforced.
Paired-end reads were assembled using SPAdes (v3.11.0) annotated using PROKKA (v1.12) with the Escherichia genus setting [14, 15]. VAGs were identified by using BLASTp with the amino acid sequences of each translated open reading frames against the Virulence Factor Database (VFDB—2019) [16] and National Center for Biotechnology Information (NCBI, Bethesda, MD, USA—2023). A requirement of 80% sequence identity across at least 80% of the length of the protein sequence was enforced for positive hits of VAGs.
A system for naming the identified VAGs was imposed. Predominantly, VAGs were named according to the gene feature of the corresponding NCBI nucleotide page. As certain VAGs had no useful name findable, for ease of reading, these were dubbed an abbreviated form of their NCBI protein name and suffixed with roman numerals in case of duplicates. The self-named VAGs are kept non-cursive in the article. A list of VAGs along with VFID, NCBI accession numbers, and representative sequences are found in Table 4 in Supplementary Appendix.
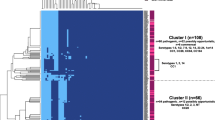
To confirm that the E. coli population was a heterogeneous group and not belong to a clonal outbreak, relatedness of E. coli was determined by MLST and core genome MLST (cgMLST). MLST and cgMLST were performed using SeqSphere + v7.2.3 (Ridom GmbH, Munster, Germany). The maximum allelic distance allowed for two samples to be considered from the same cluster was set to 10 alleles according to the clustering rules from SeqSphere + (https://www.cgmlst.org/ncs/schema/Ecoli845/). Minimum spanning trees (MSTs) were constructed to visualize the genetic relatedness among the E. coli isolates.
Data analysis
To determine specific E. coli VAGs and/or combinations of VAGs that would best predict a UTI as the primary source of the bacteremia, we compiled a list of the 358 VAGs containing the single VAGs and combined VAGs as both pairwise and triple-wise cross-pairings. The pairwise cross-pairing resulted in 63,903 pairs and the triple-wise cross-pairing resulted in 7,583,156 triplets. These VAGs or VAG combinations were then coupled with clinical data containing information on UTI status (UTI as the primary source of bacteremia or non-UTI source). We calculated the prevalence (the proportion of E. coli isolates with a given VAG or VAG combination), estimates for positive predictive value (PPV) (the proportion of patient isolates with a given VAG or VAG combination who had UTI as source), and sensitivity (the proportion of UTI source cases in the study population correctly detected with the specific VAG or combination of VAGs) (Table 5). A sorting was applied to exclude VAGs or VAG combinations having a prevalence of < 20%. The 20 VAG singles, pairs, and triplets with the highest PPVs were subjected to bootstrapping simulations of 100,000 repetitions within each DCM population. Afterwards, the high-performing 20 VAG singles, pairs, and triplets from each DCM population were tested out on the opposing DCM population with new test estimates and bootstrapping simulations calculated. No prevalence requirement was imposed here. 95% confidence intervals (CI) were calculated from nonparametric bootstrapping.
Finally, we aimed to examine to what extent our top-performing VAGs overlapped within isolate populations. To achieve this, we employed an iterative selection method to identify combinations of VAGs that would maximize the sensitivity estimate. This process involved examining all possible combinations of five individual VAGs identified in our high-PPV tables (Table 6, Table 7). The optimal combination and order were computed along with five sensitivity estimates for each combination. These sensitivity estimates were calculated as the proportion of bacteremia cases with UTI as source in which at least one of the VAGs in the combination is present. Maximizing sensitivity and minimizing isolate population overlapping allows us predictions for the vast majority of the bacteremia cases with UTI as source.
Statistical analysis and data handling were done using R (v. 4.2.2, R Foundation for Statistical Computing, Vienna, Austria).
Results
A total of 253 E. coli bacteremia cases were included in the study, 119 from DCM-1 and 105 from DCM-2. Twenty-nine cases were excluded due to unknown source of infection (n = 21) (Fig. 1). In total, there were 358 VAGs variably present across the assembled genomes.
Most cases at DCM-1 had UTI as source (81.51%), were female sex (58%), and were community-acquired (89.1%) (Table 1). This is consistent with the nature of the hospital as a secondary acute care referral hospital. Conversely, DCM-2, which contains tertiary referral hospital with specialized departments, showed an equal distribution between UTI and non-UTI as source of bacteremia (49.5% vs. 50.5%), fewer female cases (40%), and between community-acquired and hospital-acquired infections (48.6% vs. 51.4%) (Table 1). These differences likely reflect the complex patient population and specialized departments of DCM-2.
Examining the best performing single VAGs, VAG pairs, and VAG triplets in the DCM-1 bacteremia population, papX had the highest PPV of 96% (95% confidence interval (CI): [90, 100]) and a sensitivity of 54% (CI [44, 64]). intS, papE, papD, and papGII had PPVs of 93–95% and a sensitivity ranging from 34 to 56% (Table 2). Of note, papX was present in all top five-performing combinations. Multiple pap genes (papC, papF, papH) were present in the top five pairs, all predicting 100% PPV (CI [100, 100]) while still correctly predicting approximately half of the UTI source cases (sensitivity 41–46%). Adding triplets of VAGs did not improve the sensitivity.
In the DCM-2 bacteremia population, the best predicting single VAG was kpsT (PPV = 67%; CI [47, 85], sensitivity = 31%; CI [19, 44]) (Table 2). Adding pairs and triplets of VAGs increased the PPVs and sensitivity values noticeably compared to the DCM-1 population with the best ranked VAG triplet in the DCM-2 population being the combination of kpsM, z1226, and hemR (PPV = 78%; CI [61, 93], sensitivity = 40%; CI [27, 54]). No pap genes or other VAGs present in the top predicting VAGs from DCM-1 were present.
For additional single, pair, or triplet combinations with sensitivity estimates from each hospital, see Table 6 and Table 7 in the supplementary tables.
Results from the best predicting VAG combinations from the DCM-1 population (Table 2) run on the DCM-2 population data resulted in comparatively overall low sensitivity values and PPV values were also comparatively low except for papD with 71% (CI [47, 92]) and papGII with 68% (CI [46, 89]) (Table 8). Vice versa, results from the best predicting VAG combinations from the DCM-2 population (Table 2) run on DCM-1 population data performed had only VAG kpsT (PPV = 90%; CI [78, 100], sensitivity = 29%; CI [20, 38]) and in combination as kpsT and gtpp (PPV = 90%; CI [78, 100], sensitivity = 28%; CI [19, 37]) perform well with PPV value ≥ 90% (Table 8).
Iterative sensitivity optimization
Using an iterative selection approach to optimize sensitivity population coverage and minimize isolate population overlapping, we identified the optimal combinations and order of VAGs to check for in series. The combinations with the largest sensitivity values for DCM-1 population were the VAG group of papX, intS, kpsT, insN, and papC (maximum sensitivity = 92.8%) (Table 3). For the DCM-2 population, VAG group of kpsT, fimD, intS, hemR, and is3-II was found optimal in terms of individual PPV values and UTI source case sensitivity coverage (maximum sensitivity = 84.6%) (refer to Table 6 and Table 7 for individual PPV values).
Phylogenetic analyses
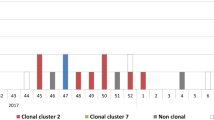
MLST and cgMLST analyses for the DCM-1 and DCM-2 populations revealed a diverse population structure which ensures that the study is performed on a diverse E. coli population and not on clonal isolates (Figs. 2 and 3). Among examined isolates, ST 131 was the most frequent, with 33 occurrences (Table 9).
Discussion
We evaluated 358 VAGs from genomes of 224 E. coli bacteremias, exploring their predictive value for UTI as the infection source of bacteremia.
We found that several VAGs predicted UTI as source in E. coli bacteremias quite well based on high PPVs. PPVs for DCM-1 and DCM-2 were respectively 93–100% and 67–78%, however with low sensitivities (DCM-1 34–54% and DCM-2 27–40%). The sensitivity increased to 85–93% by applying an “and/or” logic to various VAG combinations. Various pap genes performed well in the DCM-1 patient population in terms of both PPV and the sensitivity values and pairing these did increase slightly the PPV and sensitivities (e.g., papX, papC). In the DCM-2 patient population, kps and fim genes provide among the highest PPV values. Sensitivity values and PPV values remained low across tables for VAGs found in the DCM-2 patient population. In addition, the best performing combinations of VAGs at DCM-1 did not predict well in DCM-2 data.
Most studies examining UTI-related VAGs or virulence factors (VFs) to date have focused on E. coli in urine samples with much fewer studies focusing on E. coli bacteremia [1]. Recently, Kim et al. [17] examined the genomic difference between bacteremic UTI and non-bacteremic UTI caused by E. coli. With a study population of 80 E. coli UTI patients, of these 40 urine sample isolates and 40 blood sample isolates, they found no VFs associated with bacteremia. In contrast, Denamur et al. [18] examined E. coli bacteremia isolates in a genome-wide association study and found several pap genes (most notably the papGII operon) highly associated to the urinary tract as portal of entry, which support our findings. In the same study, a putative integrase gene, opgE, was also described and found associated to UTI as source; however, this gene was not part of the database applied to identify VAGs.
In light of previous research, it appears that certain VAGs or VFs, particularly within the pap gene cluster, may be indicators for UTI as source of E. coli bacteremia. The pap genes are a class of VFs that play a significant role in the pathogenesis of UTI. The pap operon coding for P fimbria, a chaperone-usher pathway (CUP) pilus, is located on pathogenicity islands [19]. P fimbriae are involved in adhesion to host tissues, an important step in the establishment of infection [7, 20, 21]. VFs like papC and papGII are well described as essential components of P fimbria assembly unlike papX [22]. Despite being less studied in existing literature, papX—one of the VAGs we found highly predictive of UTI as source in our study—is thought to regulate bacterial motility and expression of other E. coli fimbriae [23].
Whereas VAGs kpsT and kpsM are scarcely described in literature, the fim genes have a well-established role in UTI pathogenesis [24,25,26]. fim-genes encode the type I fimbriae, another class of CUP pili. Like P fimbriae, type I fimbriae facilitate the adherence of E. coli to host tissues, enabling initial colonization and persistent infection. The fimbriae are assembled by a conserved chaperone-usher mechanism, with fimH acting as the adhesin and other fim genes such as fimA, fimC, and fimD contributing to the complex’s assembly and transport [21, 27, 28]. Our study identified several fim-related VAGs as having strong predictive value for UTI as source as single VAGs in the DCM-2 patient population and much less so the pap genes, suggesting differences related to either foci of infections or host susceptibility factors like immunological status between the two study populations.
We selected our study populations to deliberately stem from two very different hospital setups. Having a population comprised of primarily complicated UTIs and various degrees of compromised immune systems likely makes it difficult to accurately select which patients suffer from ordinary lower UTI or pyelonephritis. The E. coli strains found in bacteremias with suspected urogenital origin from severely immunocompromised patients could also differ from those patients with normal immune system function [29]. While hospital setups such as the service area for DCM-1 appear more suited for using VAG data to predict bacteremia source origin, our findings suggest this depends heavily on patient population with no straightforward way of generalizing between hospital setups.
Study limitations include the skewed proportion of UTI as source and non-UTI as source in the DCM-1 patient population as compared to the more evenly distributed DCM-2 patient population. Not providing a specificity value is due to the VAGs being selected based on a uropathogenic profile and as such the study is not set up for examining VAGs predicting against UTI as source. While certain VAGs and combinations of VAGs emerge as significant in our data, the exact functional roles or synergistic interactions between the VAGs that might explain their predictive superiority remain outside the scope of this study. Study strengths include our large study population subjected to WGS and clinical data from our bacteremia database.
Identification of the primary source of infection will improve treatment, reduce side effects, and reduce risks associated with diagnostic procedures. We attempted to predict infection source backwards from blood culture findings using data on our landscape of local bacterial genomics. VAGs could be identified by designing a multiplex PCR targeting a list of VAGs with high individual PPV values and in unison a high cumulative sensitivity value. Long-read sequencing on the Oxford Nanopore platform could also be used to provide clinicians with fast results regarding VAGs. However, discrepancies between hospital populations require each hospital to derive its own prediction profile of VAGs.
In conclusion, genomic data showed potential to be utilized to predict the primary source of infection in E. coli bacteremia, specifically in UTIs as source of origin. However, discrepancy between best performing profile of VAGs between acute care referral hospitals (DCM-1) and a tertiary hospital (DCM-2) makes it difficult to implement in clinical practice. Comparatively, the pap genes performed the best in our analysis. Within the DCM covering acute care referral hospitals, VAGs papX and papGII were found to be both moderately sensitive and highly predictive of UTI as source of infection for E. coli bacteremia. However, no single VAGs were useful by themselves as sequentially checking a group of VAGs seems a more practical approach. The effectiveness of VAGs in predicting bacteremia source seems also to depend strongly on hospital type and patient population with no reliable ability to transfer predictions between hospital types. Future studies can test the reported predictions on external datasets. Data in larger scale will hopefully provide us more knowledge.
Data availability
The datasets analyzed in the study are available at https://www.ncbi.nlm.nih.gov/bioproject/1037322.
References
Nielsen SL, Pedersen C, Jensen TG et al (2014) Decreasing incidence rates of bacteremia: a 9-year population-based study. J Infect 69:51–59. https://doi.org/10.1016/J.JINF.2014.01.014
Luzzaro F, Viganò EF, Fossati D et al (2002) Prevalence and drug susceptibility of pathogens causing bloodstream infections in northern Italy: a two-year study in 16 hospitals. Eur J Clin Microbiol Infect Dis 21:849–855. https://doi.org/10.1007/S10096-002-0837-7
MacKinnon MC, McEwen SA, Pearl DL et al (2021) Mortality in Escherichia coli bloodstream infections: a multinational population-based cohort study. BMC Infect Dis 21:. https://doi.org/10.1186/S12879-021-06326-X
de Lastours V, Laouénan C, Royer G et al (2020) Mortality in Escherichia coli bloodstream infections: antibiotic resistance still does not make it. J Antimicrob Chemother 75:2334–2343. https://doi.org/10.1093/JAC/DKAA161
Bonten M, Johnson JR, Van Den Biggelaar AHJ et al (2021) Epidemiology of Escherichia coli bacteremia: a systematic literature review. Clin Infect Dis 72:1211–1219. https://doi.org/10.1093/CID/CIAA210
Jansåker F, Frimodt-Møller N, Bjerrum L, Dahl Knudsen J (2016) The efficacy of pivmecillinam: 3 days or 5 days t.i.d against community acquired uncomplicated lower urinary tract infections - a randomized, double-blinded, placebo-controlled clinical trial study protocol. BMC Infect Dis 16:1–7. https://doi.org/10.1186/S12879-016-2022-0/PEER-REVIEW
Flores-Mireles AL, Walker JN, Caparon M, Hultgren SJ (2015) Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat Rev Microbiol 13:269–284. https://doi.org/10.1038/NRMICRO3432
Foxman B (2014) Urinary tract infection syndromes: occurrence, recurrence, bacteriology, risk factors, and disease burden. Infect Dis Clin North Am 28:1–13. https://doi.org/10.1016/J.IDC.2013.09.003
Holm A, Cordoba G, Aabenhus R (2019) Prescription of antibiotics for urinary tract infection in general practice in Denmark. Scand J Prim Health Care 37:83–89. https://doi.org/10.1080/02813432.2019.1569425
Pedersen G, Schnheyder HC, Sørensen HT (1997) Antibiotic therapy and outcome of monomicrobial gram-negative bacteraemia: a 3-year population-based study. Scand J Infect Dis 29:601–606. https://doi.org/10.3109/00365549709035903
Kang CI, Kim SH, Wan BP et al (2005) Bloodstream infections caused by antibiotic-resistant gram-negative bacilli: risk factors for mortality and impact of inappropriate initial antimicrobial therapy on outcome. Antimicrob Agents Chemother 49:760–766. https://doi.org/10.1128/AAC.49.2.760-766.2005
Pedersen G, Schønheyder HC, Sørensen HT (2003) Source of infection and other factors associated with case fatality in community-acquired bacteremia–a Danish population-based cohort study from 1992 to 1997. Clin Microbiol Infect 9:793–802. https://doi.org/10.1046/J.1469-0691.2003.00599.X
Johnson JR (1991) Virulence factors in Escherichia coli urinary tract infection. Clin Microbiol Rev 4:80–128
Bankevich A, Nurk S, Antipov D et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/BIOINFORMATICS/BTU153
Liu B, Zheng D, Jin Q et al (2019) VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res 47:D687. https://doi.org/10.1093/NAR/GKY1080
Kim B, Kim JH, Lee Y (2022) Virulence factors associated with Escherichia coli bacteremia and urinary tract infection. Ann Lab Med 42:203–212. https://doi.org/10.3343/ALM.2022.42.2.203
Denamur E, Condamine B, Esposito-Farèse M et al (2022) Genome wide association study of Escherichia coli bloodstream infection isolates identifies genetic determinants for the portal of entry but not fatal outcome. PLoS Genet 18:e1010112. https://doi.org/10.1371/JOURNAL.PGEN.1010112
Welch RA, Burland V, Plunkett G et al (2002) Extensive mosaic structure revealed by the complete genome sequence of uropathogenic Escherichia coli. Proc Natl Acad Sci U S A 99:17020–17024. https://doi.org/10.1073/PNAS.252529799/ASSET/06E85286-A1E3-4184-8573-1107C7E853D6/ASSETS/GRAPHIC/PQ2525297003.JPEG
Klein RD, Hultgren SJ (2020) Urinary tract infections: microbial pathogenesis, host–pathogen interactions and new treatment strategies. Nat Rev Microbiol 18:211–226. https://doi.org/10.1038/s41579-020-0324-0
Waksman G, Hultgren SJ (2009) Structural biology of the chaperone–usher pathway of pilus biogenesis. https://doi.org/10.1038/nrmicro2220
Biggel M, Xavier BB, Johnson JR et al (2020) Horizontally acquired papGII-containing pathogenicity islands underlie the emergence of invasive uropathogenic Escherichia coli lineages. Nat Commun 11:1–15. https://doi.org/10.1038/s41467-020-19714-9
Simms AN, Mobley HLT (2008) PapX, a P fimbrial operon-encoded inhibitor of motility in uropathogenic Escherichia coli. Infect Immun 76:4833–4841. https://doi.org/10.1128/IAI.00630-08
Connell H, Agace W, Klemm P et al (1996) Type 1 fimbrial expression enhances Escherichia coli virulence for the urinary tract. Proc Natl Acad Sci U S A 93:9827. https://doi.org/10.1073/PNAS.93.18.9827
Wright KJ, Seed PC, Hultgren SJ (2007) Development of intracellular bacterial communities of uropathogenic Escherichia coli depends on type 1 pili. https://doi.org/10.1111/j.1462-5822.2007.00952.x
Park HK, Jung YJ, Chae HC et al (2009) Comparison of Escherichia coli uropathogenic genes (kps, usp and ireA) and enteroaggregative genes (aggR and aap) via multiplex polymerase chain reaction from suprapubic urine specimens of young children with fever. Scand J Urol Nephrol 43:51–57. https://doi.org/10.1080/00365590802299338
Sokurenko EV, Courtney HS, Maslow J et al (1995) Quantitative differences in adhesiveness of type 1 fimbriated Escherichia coli due to structural differences in fimH genes. J Bacteriol 177:3680–3686. https://doi.org/10.1128/JB.177.13.3680-3686.1995
Wurpel DJ, Beatson SA, Totsika M et al (2013) Chaperone-usher fimbriae of Escherichia coli. PLoS One 8:52835. https://doi.org/10.1371/JOURNAL.PONE.0052835
Terlizzi ME, Gribaudo G, Maffei ME (2017) UroPathogenic Escherichia coli (UPEC) infections: virulence factors, bladder responses, antibiotic, and non-antibiotic antimicrobial strategies. Front Microbiol 8:1566. https://doi.org/10.3389/FMICB.2017.01566
Acknowledgements
The authors wish to acknowledge and thank Thomas Kallemose, statistician at the Clinical Research Department, Copenhagen University Hospital Hvidovre, Hvidovre, Denmark, for helping design the statistical analysis.
Funding
Open access funding provided by Copenhagen University. Authors H.K.J., H.W., and J.M. were supported by a grant from The Novo Nordisk Foundation (Ref. nr.: NNF18SA0035306)—Pasteur21 POC. Author H.K.J. was supported by a Challenge grant (Ref. nr.: NNF19OC0056411), and authors H.K.J. and H.W. were supported by a Clinical-Academic-Group (CAG)—Greater Copenhagen Health—Science—Partners, 2020. BACINFECT.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Isolate collection and sequencing were performed by HK-J and HW. Bioinformatic analysis was performed by JM. Clinical data collection was performed by MP, KHH, and FBH. Data availability and upload were handled by PW. The data analysis and manuscript writing were performed by CSI supervised by MP and KHH. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
We have obtained all necessary permissions to access pathogen genomes and personal data from all required authorities including the Scientific Ethical Committee of the Capital Region (reference 19029688), the Danish Board for Patient Safety Authority, and the project is notified in PACTIUS at Rigshospitalet and at the Data Protection Agency.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Katrine Hartung Hansen and Mette Pinholt shared last authorship.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ilsby, C.S., Hertz, F.B., Westh, H. et al. Predicting the primary infection source of Escherichia coli bacteremia using virulence-associated genes. Eur J Clin Microbiol Infect Dis 43, 641–648 (2024). https://doi.org/10.1007/s10096-024-04754-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-024-04754-6