Abstract
Toxoplasma gondii infections are prevalent in humans and animals all over the world. The aim of the study was to estimate the occurrence of T. gondii oocysts in fruits and vegetables and determine the genotype of the parasites. A total number of 216 fruits and vegetables samples were taken from shops and home gardens located in the area of northern Poland. Oocysts were recovered with the flocculation method. Then, real-time polymerase chain reaction (PCR) targeting the B1 gene was used for specific T. gondii detection and quantification. Toxoplasma DNA was found in 21 samples. Genotyping at the SAG2 locus showed SAG2 type I and SAG2 type II. This is the first investigation describing T. gondii DNA identification in a large number of fruits and vegetables samples with rapid molecular detection methods. The results showed that fruits and vegetables contaminated with T. gondii may play a role in the prevalence of toxoplasmosis in Poland.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Toxoplasma gondii is a worldwide distributed protozoa that causes toxoplasmosis, one of the most prevalent parasitic infections in humans [32, 35]. The disease is usually asymptomatic in immunocompetent individuals, but may take a severe course in immunodeficient patients, i.e. pharmacologically immunosuppressed or human immunodeficiency virus (HIV)-infected [23, 28], as well as in immature foetuses and infants (if the mother suffers from primary infection during pregnancy) [8, 39].
People may acquire toxoplasmosis mainly by the consumption of raw and undercooked meat, and also by the ingestion of T. gondii oocysts present in environment (water, soil, fruits and vegetables) contaminated with the faeces of infected cats. A few outbreaks of toxoplasmosis reported worldwide were connected with water or soil contaminated with oocysts [9, 10, 12, 13]. T. gondii oocysts are resistant to unfavourable environmental conditions, as well as chemical inactivation, being an excellent reservoir of the parasite [15, 16, 29, 31].
So far, there are not many studies concerned with the detection of T. gondii oocysts in environmental samples. For years, efficient detection methods were not available. The situation is finally changing with the recent development of specific and sensitive methods to recover and identify T. gondii oocysts [17–19]. Little information has been gathered about T. gondii detection in water and soil [1, 4, 7, 13, 17, 27, 28, 30, 36, 38]. However, there is no study concerning the contamination of fresh fruits and vegetables with this parasite.
Therefore, the aim of the study was to estimate the occurrence of T. gondii oocysts on fresh fruits and vegetables (bought in shops and taken from home gardens) in Poland, as well as to determine the genotype of the detected parasites.
Materials and methods
Recovery experiments
In initial experiments prior to the evaluation of T. gondii oocysts detection in fruits and vegetables samples, recovery tests were performed by using oocysts of a ∼2-year-old PRU strain (genotype II). The oocysts were kept in 2% H2SO4 and stored at 4°C. Four oocyst-free samples of radish and strawberries bought in a greengrocers were prepared in the laboratory by washing in distilled water in order to minimise the risk of accidental contamination with T. gondii. Then, the samples were experimentally contaminated with T. gondii oocysts in amounts of 104, 103, 102 and 10, and oocysts recovery and the specific detection of T. gondii DNA was performed as described below. The recovery samples were prepared in triplicate to determine the reproducibility of the methods.
Sampling
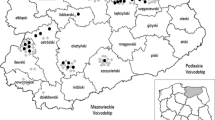
A total of 216 fruits and vegetables samples were collected between June 2006 and August 2008 at different sites located in the area of the Tri-City and Elblag and in their vicinity (northern Poland). Of these, 175 samples were bought in supermarkets, from greengrocers and in marketplaces, and 41 samples were taken from kitchen-gardens and allotments (Table 1). One sample consisted of 1 kg of strawberries, 0.5 kg of carrot, 20 radishes and one lettuce. The samples were put into disposable bags and transported to the laboratory.
Oocysts recovery
In order to concentrate and recover T. gondii oocysts from fruits and vegetables samples, the flocculation method using CaCO3 solution [37] was employed. Briefly, one fruit or vegetable sample was placed in a 3-l-volume glass vessel, with 2 l of distilled water and 20 ml of Tween20 being added, and the content was slightly mixed by a manual rotation of the vessel and then shaken for 2 h at 100 rpm on an automatic orbital shaker (Multi PSU-20, BioSan, Warren, MI, USA). Then, the fruits or vegetables were removed. After that, 20 ml of 1 M CaCl2 solution and 20 ml of 1 M NaHCO3 solution was added to the washings and shaken by several manual rotations of the vessel. Then, 1 M NaOH solution was added in order to obtain pH 10 of the mixture and it was left overnight (without shaking). The next day, the supernatant was removed with the use of an automatic pipette filler (Hirschmann Laborgeräte). The obtained pellets were suspended in 200 ml of 10% NH3O3S solution and the suspension was placed in two 200-ml tubes. The glass vessel was washed with 20 ml of 0.01% Tween80 and the washings were added to the suspended pellet. Then, this was centrifuged for 10 min at 1,700 × g at a temperature of 4°C. The supernatant was removed and the pellet was suspended in a minimal amount of 0.01% Tween80. The pellets were placed in one Falcon tube and distilled water was added up to 50 ml volume. Then, it was mixed and centrifuged for 10 min at 2,500 × g at a temperature of 4°C. The supernatant was removed and the pellet was suspended in a double amount of distilled water. The final suspension was placed in a 1.5-ml Eppendorf tube and stored at 4°C.
DNA extraction
Firstly, a few steps were used to destroy the oocysts wall and lyse T. gondii cells. To the Lysing Matrix tubes (BIO 101 Systems, USA), 1 g of 1-mm-diameter zircon beads (BioSpec Products Inc., USA) and 300 μl of L1 buffer (100 mM Tris pH 8.0, 10 mM EDTA, 2.5% Tween20) and 200–300 μl of suspension obtained after the recovery procedure was added. This matter was shaken for 2 min in the FastPrep 120 Instrument (BIO 101 Savant, USA) at an amplitude of 6.5 and then incubated in ice for 10 min. The last step was performed twice. Then, 400 μl of chloroform was added and the tubes were centrifuged for 1 min. The supernatant was placed in a 1.5-ml Eppendorf tube and 20 μl of proteinase K was added and incubated for 1 h at 50°C. Then, this was placed in a minicolumn of the commercial Genomic Mini Kit (A&A Biotechnology, Gdynia, Poland) and DNA isolation was continued according to the manufacturer’s instructions and then stored at −20°C.
Specific detection of Toxoplasma gondii by real-time PCR
For specific T. gondii detection, real-time polymerase chain reaction (PCR) was performed with the use of a pair of primers 5′ TCGAAGCTGAGATGCTCAAAGTC 3′ and 5′ AATCCACGTCTGGGAAGAACTC 3′ targeting the 129-bp fragment of 35-fold repetitive B1 gene and the fluorescently labelled TaqMan probe 5′ FAM ACCGCGAGATGCACCCGCA TAMRA 3′ [5]. Amplification was performed with an initial polymerase activation step (10 min at 95°C), followed by 40 cycles of denaturation (15 s at 95°C) and hybridisation/extension (1 min at 60°C). The amplification reaction mixture consisted of: 12.5 μl 2 × Brilliant II QPCR Master Mix (Stratagene, USA), 400 nM of each primer (Metabion, Germany), 80 nM of TaqMan probe (Metabion, Germany) and 2 μl of template DNA in a 25-μl reaction volume. Amplifications were performed in an Mx3005P thermocycler (Stratagene, USA). PCR products were analysed using MxPro QPCR Software. The cycle threshold (CT) value, determining the cycle number at which the reporter’s fluorescence exceeds the threshold value, was recorded. A sample was considered to be positive if the CT value was <40.
All PCR experiments were performed including a T. gondii-positive control (genomic DNA from the T. gondii RH strain) to ensure the correct functioning of the reaction and negative control (water template) to ensure no contamination of the PCR components. Additionally, all of the negative samples were retested for the presence of PCR inhibitors by mixing 2 μl of DNA template and 1 μl of T. gondii-positive control.
Genotyping
In order to identify T. gondii genotype, the restriction analysis method was used as described by Aspinall et al. [6]. Nested PCR reaction was performed to amplify two fragments of the T. gondii SAG2 gene (3′ and 5′ ends). The PCR reaction mixture consisted of: 2.5 μl 10 × GeneAmp PCR Buffer II (Applied Biosystems, USA), 2 mM MgCl2 (Applied Biosystems, USA), 0.2 mM of each dNTP (Fermentas, Lithuania), 0.4 μM of each primer (Metabion, Germany), 1.25 U of AmpliTaq Gold polymerase (Applied Biosystems, USA) and 2 μl of template DNA in a 25-μl reaction volume. Both amplification rounds were performed under the same conditions as described by Aspinall et al. [6], with a change in the initial polymerase activation step (15 min at 95°C).
SAG2 genotypes were determined by restriction fragment length polymorphism (RFLP) analysis using Sau3AI restriction enzyme (Promega, USA) digesting nested PCR products of the 5′ end and CfoI restriction enzyme (Promega, USA) digesting nested PCR products of the 3′ end (in separate reactions). All digestions were performed according to the manufacturer’s instructions. The digested products were analysed by 2% agarose gel electrophoresis.
Quantitative real-time PCR
In order to determine the initial copy number of the detected T. gondii DNA, each positive sample was tested with the real-time qualitative PCR (QPCR) method based on the standard curve.
Standard template was generated by cloning the insert gene (129-bp fragment of the B1 gene amplified by conventional PCR using the set of primers described above) into a plasmid. The DNA concentration of the standard was measured with the NanoDrop 1000 spectrophotometer (Thermo Scientific) according to the manufacturer’s instructions. Then, the copy number was determined with the use of the “Calculator for determining the number of copies of a template” (URI Genomics and Sequencing Center).
To generate the standard curve, two series of eight dilutions of standard DNA in the range of 5 × 108 − 0.5 DNA copies per one μl were prepared. After amplification of the standard dilution series, the standard curve was obtained by plotting the log of the initial template copy number against the CT value generated from each dilution. Amplification of the standard dilution series and the DNA isolated from fruits and vegetables samples were run on the same plate. Consequently, comparing the CT values of the unknown samples with the standard curve allowed the quantification of initial copy numbers. Then, the T. gondii oocysts number that could be present in the examined samples was estimated (including oocysts loss during the recovery and DNA extraction procedures). Our previous experiments showed that the lowest oocysts number detected with real-time PCR without taking into account the recovery procedure was 10. Considering the recovery procedure, the detectable oocysts number was 100 for vegetable samples (Table 2). If the approximate loss of oocysts was ten-fold for vegetable samples, then the oocysts charge of the sample could be estimated using the elaborated formula: OCS = N× 10/35 × 8, where: OCS = oocyst charge of the sample; N = initial copy number determined using real-time QPCR; 35 = number of copies of the B1 gene per T. gondii cell; 8 = number of T. gondii sporozoites per oocyst; 10 = approximate loss of oocysts for vegetables samples, respectively.
Determining the real-time PCR detection limit
To determine real-time PCR detection limit, standard DNA (with estimated DNA concentration) was diluted to obtain 5 × 108, 5 × 107, 5 × 106, 5 × 105, 5 × 104, 5 × 103, 5 × 102, 5 × 101, 5 and 0.5 copies, respectively, and amplification of the B1 gene fragment was performed as described above.
All molecular examinations were performed in a PCR laboratory consisting of seven rooms suited to particular functions, which prevented contamination with extraneous DNA.
Results
Results of the recovery test
The results of the recovery test showed that we were able to detect T. gondii parasites as contamination of the vegetable samples (radish) with at least 102 oocysts and only 104 oocysts in the case of fruit samples (strawberries) (Table 2, Figs. 1 and 2).
Result of the Toxoplasma gondii oocysts recovery test from experimentally contaminated fruits (strawberries) samples. Amplification plots of the T. gondii B1 gene: line 1, positive control (RH strain); line 2, sample contaminated with 104 oocysts; line 3, sample contaminated with 103 oocysts; line 4, sample contaminated with 102 oocysts; line 5, sample contaminated with 10 oocysts; line 6, negative control
Result of the T. gondii oocysts recovery test from experimentally contaminated vegetables (radish) samples. Amplification plots of the T. gondii B1 gene: line 1, positive control (RH strain); line 2, sample contaminated with 104 oocysts; line 3; sample contaminated with 103 oocysts; line 4, sample contaminated with 102 oocysts; line 5, sample contaminated with 10 oocysts; line 6, negative control
Identification of Toxoplasma gondii parasites DNA in fruits and vegetables samples
A total of 216 samples were examined with the real-time PCR detection method based on the T. gondii B1 gene. The presence of T. gondii DNA was noted in 21 samples (9.7%) (Table 3). The inhibition of real-time PCR was noted in 62 samples (Table 4).
Among the 21 tested samples, only eight that were found to be positive for both 5′ and 3′ SAG2 gene amplified fragments could be digested by Sau3AI and CfoI restriction endonucleases, respectively, and then analysed. According to the restriction patterns obtained after separation by electrophoresis (Fig. 3), we classified previously detected parasites as SAG2 type I (six samples) and SAG2 type II (two samples) (Table 1). The remaining 13 samples were found to be positive only for one either 5′ or 3′ amplified PCR products of the SAG2 gene; therefore, in these cases, we were not able to determine the SAG2 type of the parasite (Table 1).
T. gondii SAG2 typing of the vegetables samples. b Sau3AI digestions of 5′ SAG2 amplification products loaded onto a 2% gel agarose. b CfoI digestions of 3′ SAG2 amplification products loaded onto a similar gel. Line 1, molecular weight marker (pUC19 DNA/MspI (HpaII) Marker, 23, Fermentas, Lithuania). Line 2, positive control (RH strain, type I). Samples loaded in lines 3, 4, 5, 6, 7 and 8 appear to be SAG2 type I (no digestion). Samples loaded in lines 9 and 10 contain SAG2 type II parasite (digestion with CfoI endonuclease and lack of digestion with Sau3AI restriction enzyme)
The estimated contamination of examined samples was relatively low. The oocysts number per sample was less than ten, except for one sample, where approximately 20 oocysts were detected (Table 1).
Discussion
In the literature, there is no sufficient information concerning T. gondii oocysts contamination of fresh fruits and vegetables intended for consumption. However, there are suggestions that it may be one of the sources of T. gondii infection in humans [17]. Some information has been gathered on the presence of other protozoa on fruits and vegetables. For example, raspberry appeared to be a source of Cyclospora cayetanensis, which is closely related to T. gondii coccidium [17]. Cryptosporidium oocysts were also recovered from fruits and vegetables samples [11, 37, 40], as well as Giardia cysts [21]. In Poland, environmental samples of fruits and vegetables were investigated for human-virulent microsporidian spores [24].
Cool and watery fruits and vegetables may also create perfect conditions for T. gondii oocysts persistence. Experimental tests proved that they may stay viable on raspberry stored for 8 weeks at 4°C [26]. However, comprehensive environmental investigation has not yet been performed. Therefore, according to our knowledge, it is one of the first attempts to examine the contamination of a large number of fresh fruits and vegetables with T. gondii using molecular methods. A total of 216 samples of fruits and vegetables bought in shops and marketplaces, as well as taken from home gardens, were investigated. The results of the real-time PCR showed that about 10% of the samples were positive. Our findings provide evidence that fresh fruits and vegetables may play a role in toxoplasmosis epidemiology. The presence of T. gondii DNA in the examined samples clearly indicates that the consumption of raw, unwashed fruits and vegetables may be a source of T. gondii infection in humans.
The majority of all of the examined samples were bought in shops and bazaars. Among the samples from large stores, no T. gondii DNA was found. This may be due to the fact that the fruits and vegetables come from great producers and are cultivated in remote places far from inhabited areas, with a small probability of feline presence. Additionally, stored fruits and vegetables undergo refreshing preparatory processes (for example, washing). Therefore, T. gondii oocysts that could be present on the food are most likely washed away. In case of smaller greengrocers, stands at bazaars or pitches, fruits and vegetables often come from small agricultures and their contact with feline faeces is more probable. Thus, in the last mentioned group of the samples, T. gondii DNA was recorded (8%).
Among the samples taken from the farms (home gardens), seven were positive (Table 1). The farms from which the samples originated were characterised by good sanitary conditions, except for Chynow, where poor hygienic standards was noted. In all cases, the presence of domestic and/or wild felids were recorded. Analysis of the data (number of collected samples and hygienic conditions) shows that hygienic conditions are not the main factor determining the occurrence of positive samples. The negative samples are most likely due to the lack of animals excreting oocysts. However, limitations of the detection methods as well as the relation between wide sampling area to small sample size should also be taken into account. Provided with a low contamination of the environment, it is possible that effective detection is not available. Therefore, the number of oocysts present in the samples could, however, be higher than we were able to prove with the described methods.
Despite the fact that recovery methods of T. gondii oocysts are improving, the detection of this parasite at a low contamination of environmental sample still remains unavailable. It results mainly in the limitation of recovery procedures, during which a lot of oocysts are lost. Comparison of the results of real-time PCR performed with DNA extracted from T. gondii oocysts suspension in distilled water with the results of recovery tests showed that, in the case of vegetables samples, ten-fold and, in strawberries samples, thousand-fold loss of T. gondii oocysts was recorded.
Resistant oocyst wall consisting of few layers results in difficult T. gondii DNA extraction. Procedures destroying the wall developed for other closely related with T. gondii coccidia include: freezing/throwing cycles, grinding with glass beads proteinase K digestion [17]. In this study, we used mechanic disruption with zircon beads, incubation with lysis buffer containing Tween20 and proteinase K digestion. This pre-isolation step improved the efficiency of DNA isolation in comparison to the results obtained with the use of commercial DNA extraction kit only (Genomic Mini, A&A Biotechnology, Poland) (data not shown).
Moreover, PCR inhibitors such as humid acids or polysaccharides play a role in environmental samples. They may reduce the PCR sensitivity by up to 100–1,000 times [25]. Inhibition was already noted in the surface water samples (up to 50%) [7, 28, 38]. In this study, quite a large number of samples with PCR inhibition was noted (29%) (Table 3), mainly among strawberries samples (>50%). During washing, these fruits are partly destroyed. The pellet includes a large amount of the slime and fruit fragments, which creates difficulties in the oocysts recovery process and is probably a source of PCR inhibitors. This phenomenon was not noted in vegetable samples because of their tight structure.
Specific and sensitive methods are developed for other protozoa but are not yet available for T. gondii oocysts detection. Microscopy as well as bioassay are not sufficient to perform sensitive and simple detection. So, it may support rather than replace molecular investigation. So far, only a few publications concerning T. gondii oocysts detection in surface water were different PCR techniques used, PCR, nested PCR, real-time PCR, taking different T. gondii genes as a molecular target. However, most of the research involved clinical samples or animal tissues. In the case of the environmental samples, genes having many copies per cell are more efficient. At present, the most often used genes to detect T. gondii are B1 and REP [7, 19, 38]. In this study, we chose the TaqMan PCR assay described by Arkush et al. [5] to detect the T. gondii B1 gene. The assay is characterised by reproducibility and sensitivity (few DNA copies in a 25-μl reaction). Moreover, it has been tested for its specificity with the use of different species of Cryptosporidium, Sarcocystis and Neospora. The specificity of the method is important in order to make amplification of the DNA of closely related to T. gondii species impossible.
Using quantitative detection by real-time PCR, despite the improving specificity because of the molecular probes, it is also possible to estimate the approximate oocysts charge (oocyst number) of the sample. The results of the quantitative real-time PCR showed that contamination of the positive samples equalled approximately three oocysts per sample, excluding one extreme (sample no. 187, contaminated with 19 oocysts) (Table 1).
T. gondii isolates were classified into three main clonal groups (I, II and III), based on the single loci (SAG2 gene) or multilocus analysis [22, 33]. There is no doubt that the prevalence of T. gondii genotypes in the environment remains unknown. However, from the point of view of human health, it is important information. There is little information on the occurrence of Toxoplasma genotypes in Poland [14, 33]. In this study, two genotypes were determined: SAG2 type I in six samples and SAG2 type II in two samples. The data are interesting, because type I is less frequent in Europe, while type II predominates [2, 34]. However, the data correspond with our previous findings concerning T. gondii detection in soil [30]. It has been proved that the popular genotyping method based on the amplification of the SAG2 gene of T. gondii has a serious limitation. Our investigation confirmed this notion. The main problem is the unpredictable amplification of the SAG2 gene fragments. Often, only one fragment (3′ end or 5′ end) was amplified, which makes restriction analysis and, consequently, the determination of the genotype, impossible. In the literature, there is more and more information about alternative genotyping methods of T. gondii (e.g. microsatellite analysis [3]) and GRA6 gene analysis [20], that should replace or complement genotyping based on the SAG2 locus.
The performed examinations did not allow estimation of the viability of the oocysts and, consequently, the infectious ability for humans was not determined. Bioassay is mainly used to determine oocysts viability, while the methods based on RNA detection (reverse transcription PCR [RT-PCR] or fluorescence in situ hybridisation [FISH]) have not yet been developed. However, mouse bioassay is time-consuming and does not identify neither unsporulating nor damaged oocysts (they may lose their infectivity during the recovery process) [23, 38].
The results of our findings showed that real-time PCR is a specific and sensitive detection method (it was able to detect a few copies of the B1 gene, which means that it is theoretically able to detect one cell of T. gondii). However, the effectiveness of the whole procedure, including oocysts recovery, DNA isolation, as well as the presence of PCR inhibitors, is far from satisfactory.
References
Afonso E, Lemoine M, Poulle ML, Ravat MC, Romand S, Thulliez P, Villena I, Aubert D, Rabilloud M, Riche B, Gilot-Fromont E (2008) Spatial distribution of soil contamination by Toxoplasma gondii in relation to cat defecation behaviour in an urban area. Int J Parasitol 38:1017–1023
Ajzenberg D, Cogné N, Paris L, Bessières MH, Thulliez P, Filisetti D, Pelloux H, Marty P, Dardé ML (2002) Genotype of 86 Toxoplasma gondii isolates associated with human congenital toxoplasmosis, and correlation with clinical findings. J Infect Dis 186:684–689
Ajzenberg D, Bañuls AL, Su C, Dumètre A, Demar M, Carme B, Dardé ML (2004) Genetic diversity, clonality and sexuality in Toxoplasma gondii. Int J Parasitol 34:1185–1196
Aramini JJ, Stephen C, Dubey JP, Engelstoft C, Schwantje H, Ribble CS (1999) Potential contamination of drinking water with Toxoplasma gondii oocysts. Epidemiol Infect 122:305–315
Arkush KD, Miller MA, Leutenegger ChM, Gardner IA, Packham AE, Heckeroth AR, Tenter AM, Barr BC, Conrad PA (2003) Molecular and bioassay-based detection of Toxoplasma gondii oocyst uptake by mussels (Mytilus galloprovincialis). Int J Parasitol 33:1087–1097
Aspinall TV, Guy EC, Roberts KE, Joynson DHM, Hyde JE, Sims PFG (2003) Molecular evidence for multiple Toxoplasma gondii infections in individual patients in England and Wales: public health implications. Int J Parasitol 33:97–103
Aubert D, Villena I (2009) Detection of Toxoplasma gondii oocysts in water: proposition of a strategy and evaluation in Champagne-Ardenne Region, France. Mem Inst Oswaldo Cruz (Rio de Janeiro) 104:290–295
Beazley DM, Egerman RS (1998) Toxoplasmosis. Semin Perinatol 22(4):332–338
Benenson MW, Takafuji ET, Lemon SM, Greenup RL, Sulzer AJ (1982) Oocyst-transmitted toxoplasmosis associated with ingestion of contaminated water. N Engl J Med 307:666–669
Bowie WR, King AS, Werker DH, Isaac-Renton JL, Bell A, Eng SB, Marion SA (1997) Outbreak of toxoplasmosis associated with municipal drinking water. The BC Toxoplasma Investigation Team. Lancet 350:173–177
Campbell AT, Robertson LJ, Smith HV, Girdwood RW (1994) Viability of Cryptosporidium parvum oocysts concentrated by calcium carbonate flocculation. J Appl Bacteriol 76:638–639
Coutinho SG, Lobo R, Dutra G (1982) Isolation of Toxoplasma from the soil during an outbreak of toxoplasmosis in a rural area in Brazil. J Parasitol 68:866–868
de Moura L, Bahia-Oliveira LMG, Wada MY, Jones JL, Tuboi SH, Carmo EH, Ramalho WM, Camargo NJ, Trevisan R, Graça RMT, da Silva AJ, Moura I, Dubey JP, Garrett DO (2006) Waterborne toxoplasmosis, Brazil, from field to gene. Emerg Infect Dis 12:326–329
Dubey JP, Huong LTT, Lawson BWL, Subekti DT, Tassi P, Cabaj W, Sundar N, Velmurugan GV, Kwok OCK, Su C (2008) Seroprevalence and isolation of Toxoplasma gondii from free-range chickens in Ghana, Indonesia, Italy, Poland, and Vietnam. J Parasitol 94:68–71
Dubey JP, Jenkins MC, Thayer DW (1996) Irradiation killing of Toxoplasma gondii oocysts. J Eukaryot Microbiol 43:123S
Dubey JP, Thayer DW, Speer CA, Shen SK (1998) Effect of gamma irradiation on unsporulated and sporulated Toxoplasma gondii oocysts. Int J Parasitol 28:369–375
Dumètre A, Dardé ML (2003) How to detect Toxoplasma gondii oocysts in environmental samples? FEMS Microbiol Rev 27:651–661
Dumètre A, Dardé ML (2005) Immunomagnetic separation of Toxoplasma gondii oocysts using a monoclonal antibody directed against the oocyst wall. J Microbiol Methods 61:209–217
Dumètre A, Dardé ML (2007) Detection of Toxoplasma gondii in water by an immunomagnetic separation method targeting the sporocysts. Parasitol Res 101:989–996
Edvinsson B, Dardé ML, Pelloux H, Evengård B; ESCMID Study Group on Toxoplasmosis (2007) Rapid genotyping of Toxoplasma gondii by pyrosequencing. Clin Microbiol Infect 13:424–429
Ho BSW, Tam T-Y, Yam WC, Hutton P (1995) Detection and enumeration of Giardia cysts in river waters of Hong Kong by flocculation–percoll/sucrose gradient–immunofluorescence method. Water Sci Technol 31:431–434
Howe DK, Sibley LD (1995) Toxoplasma gondii comprises three clonal lineages: correlation of parasite genotype with human disease. J Infect Dis 172:1561–1566
Isaac-Renton J, Bowie WR, King A, Irwin GS, Ong CS, Fung CP, Shokeir MO, Dubey JP (1998) Detection of Toxoplasma gondii oocysts in drinking water. Appl Environ Microbiol 64:2278–2280
Jedrzejewski S, Graczyk TK, Slodkowicz-Kowalska A, Tamang L, Majewska AC (2007) Quantitative assessment of contamination of fresh food produce of various retail types by human-virulent microsporidian spores. Appl Environ Microbiol 73:4071–4073
Johnson DW, Pieniazek NJ, Griffin DW, Misener L, Rose JB (1995) Development of a PCR protocol for sensitive detection of Cryptosporidium oocysts in water samples. Appl Environ Microbiol 61:3849–3855
Kniel KE, Lindsay DS, Sumner SS, Hackney CR, Pierson MD, Dubey JP (2002) Examination of attachment and survival of Toxoplasma gondii oocysts on raspberries and blueberries. J Parasitol 88:790–793
Kourenti C, Heckeroth A, Tenter A, Karanis P (2003) Development and application of different methods for the detection of Toxoplasma gondii in water. Appl Environ Microbiol 69:102–106
Kourenti C, Karanis P (2006) Evaluation and applicability of a purification method coupled with nested PCR for the detection of Toxoplasma oocysts in water. Lett Appl Microbiol 43:475–481
Kutičič V, Wikerhauser T (1996) Studies of the effect of various treatments on the viability of Toxoplasma gondii tissue cysts and oocysts. Curr Trop Microbiol Immunol 219:261–265
Lass A, Pietkiewicz H, Modzelewska E, Dumètre A, Szostakowska B, Myjak P (2009) Detection of Toxoplasma gondii oocysts in environmental soil samples using molecular methods. Eur J Clin Microbiol Infect Dis 28:599–605
Lindsay DS, Blagburn BL, Dubey JP (2002) Survival of nonsporulated Toxoplasma gondii oocysts under refrigerator conditions. Vet Parasitol 103:309–313
Montoya JG, Remington JS (1995) Studies on the serodiagnosis of toxoplasmic lymphadenitis. Clin Infect Dis 20:781–789
Nowakowska D, Colón I, Remington JS, Grigg M, Gołąb E, Wilczyński J, Sibley LD (2006) Genotyping of Toxoplasma gondii by multiplex PCR and peptide-based serological testing of samples from infants in Poland diagnosed with congenital toxoplasmosis. J Clin Microbiol 44:1382–1389
Owen MR, Trees AJ (1999) Genotyping of Toxoplasma gondii associated with abortion in sheep. J Parasitol 85:382–384
Petersen E, Dubey JP (2001) Biology of Toxoplasma gondii. In. Joynson DHM, Wreghitt T (eds.) Toxoplasmosis: a comprehensive clinical guide. Cambridge University Press, Cambridge, pp 1–49
Sroka J, Wójcik-Fatla A, Dutkiewicz J (2006) Occurrence of Toxoplasma gondii in water from wells located on farms. Ann Agric Environ Med 13:169–175
Vesey G, Slade JS, Byrne M, Shepherd K, Fricker CR (1993) A new method for the concentration of Cryptosporidium oocysts from water. J Appl Bacteriol 75:82–86
Villena I, Aubert D, Gomis P, Ferté H, Inglard JCh, Denis-Bisiaux H, Dondon JM, Pisano E, Ortis N, Pinon JM (2004) Evaluation of a strategy for Toxoplasma gondii oocyst detection in water. Appl Environ Microbiol 70:4035–4039
Wong SY, Remington JS (1994) Toxoplasmosis in pregnancy. Clin Infect Dis 18:853–861
Zuckerman U, Gold D, Shelef G, Armon R (1997) The presence of Giardia and Cryptosporidium in surface waters and effluents in Israel. Water Sci Technol 35:381–384
Acknowledgment
This work was supported by the Ministry of Education and Science grant No. 2P05D00729 and by the European Union within the European Social Fund in the framework of the project “InnoDoktorant—Scholarships for PhD students, I edition”. The authors wish to thank Pr. Marie-Laure Dardé from the Laboratory of Parasitology-Mycology, Limoges Hospital, France, for the T. gondii sporulating oocysts that were used to perform the oocyst recovery tests. We also thank Pr. Tadeusz Dzbeński from the National Institute of Hygiene, Poland, for the T. gondii tachyzoites (RH strain) DNA, which was used as our positive control.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Lass, A., Pietkiewicz, H., Szostakowska, B. et al. The first detection of Toxoplasma gondii DNA in environmental fruits and vegetables samples. Eur J Clin Microbiol Infect Dis 31, 1101–1108 (2012). https://doi.org/10.1007/s10096-011-1414-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-011-1414-8