Abstract
The aim of this study was to evaluate the prevalence and molecular characteristics of methicillin-resistant Staphylococcus aureus (MRSA) among pigs and estimate the impact of this animal reservoir on human healthcare. Nasal swabs were derived from 1,600 pigs at 40 German farms. The MRSA were characterized using S. aureus protein A (spa) typing, multilocus sequence typing (MLST) and detection of toxin genes. In a retrospective case control study, we compared risk factors for the carriage of MRSA between patients carrying spa types found among regional pigs and patients with other MRSA molecular types. Pigs carrying MRSA were identified on 70% of the farms (spa types t011, t034, t108, t1451 and t2510, all associated with MLST sequence type ST398). Contact to pigs and cattle were independent risk factors for the carriage of these spa types in patients at hospital admission. Our results indicate that livestock represents a relevant reservoir for the import of MRSA into regional German hospitals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During recent decades methicillin-resistant Staphylococcus aureus (MRSA) have emerged as human nosocomial pathogens throughout the world [1]. More recently, several reports described their importance in veterinary medicine. MRSA were found to colonize and infect various animal species comprising horses [2–4], cows [5], dogs and cats [6, 7], rabbits [7] and other companion animals [8]. A special focus was put on the isolation of MRSA from pigs in several countries including The Netherlands, Denmark, France, Canada, the USA and Singapore [9–15]. Although in the majority of reports pigs were colonized, but not infected by MRSA, some studies also described the development of mastitis or urinary tract infections in the animals [16]. Molecular typing showed that the porcine MRSA isolates were mostly associated with clonal complex (CC) 398 by multilocus sequence typing (MLST), corresponding to S. aureus protein A (spa) types t011, t034, t108, t1451 and relatives [9, 13].
Consecutively, several reports suggested transmission between pigs and humans causing MRSA colonization in 23–45% of pig-farmers [14, 17, 18] and 4.6% of pig-care veterinarians [19]. A porcine MRSA reservoir therefore might exert manifold influence on human healthcare. Since MRSA CC398 will potentially cause severe infections in humans [20, 21], contact to pig-farming might represent a risk-factor for the development of MRSA-associated illness. Moreover, as studies indicated the entrance of MRSA in the food chain, e.g. via contamination of carcasses during the slaughtering process [22] or via secondary contamination by slaughterhouse staff [23], the potential of these MRSA to cause foodborne disease has to be evaluated. Apart from a direct pathogenic impact for carriers, patients colonized with MRSA originating from the livestock-reservoir might introduce antibiotic-resistant pathogens into the hospitals. As MRSA screening strategies in hospitals are usually based upon on-admission swabbing of patients who are attached to classical nosocomial risk factors for MRSA carriage, the established “search” policies might fail to detect patients with a livestock-related MRSA-acquisition.
Currently, there is growing concern about MRSA colonization of livestock in Germany, but, besides a first pilot study [24], no further data are available. In order to elucidate this issue, we performed a prevalence study on MRSA colonization among pigs at randomly selected farms in Germany in an area with a high density of pig production. The MRSA isolates were characterized using spa typing, MLST, SCCmec typing and detection of important S. aureus toxin-encoding genes.
Furthermore, the importance of an import of livestock-related MRSA into hospitals in countries with medium or high MRSA prevalence, as described for Germany [25], is currently unknown. In consequence, we performed a retrospective case control study among patients admitted to a tertiary-care hospital situated in the region in which the prevalence study was carried out. In this case control study, we included patients carrying such MRSA spa types identified among regional pigs at admission to the hospital. As a control group, we chose patients colonized with other MRSA spa types hitherto not associated with livestock. For both groups we assessed classical nosocomial risk factors for MRSA carriage [26] and asked for a status of contact to pigs, cattle, horses, sheep or other animals. The comparison of risk factors among patients in the case group and the control group shall clarify whether patients carrying presumptive livestock-related MRSA types are likely to be colonized via transmission from animals.
Methods
MRSA screening on pig farms
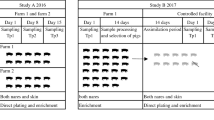
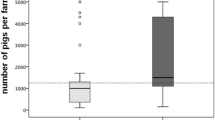
From April 2007 to November 2007 we collected nasal swabs from weaned pigs living on 40 different randomly chosen pig farms (25 farrowing farms, 4 rearing farms and 11 farrow-to-finishing farms) on the German side of the Euregio, situated in the central Dutch-German border area. The farms were situated in the following districts according to the nomenclature of territorial units for statistics (NUTS) (http://ec.europa.eu/comm/eurostat/ramon/nuts): DEA34, DEA37, DEA35, DEA29, DEA1B, DEA14, DEA15, DEA1D, DEA1E, DEA1F. The average animal density was 1,277 pigs (range 375–3400 pigs) in the farrowing and farrow-to-finish units and 1,900 pigs (range 1800–2000 pigs) on the rearing farms.
On each farm we derived 40 swabs from weaned piglets. For microbiological analysis the 40 swabs were pooled into eight culture samples. After 24-h enrichment culture in selective broth [27], all cultures were streaked onto Columbia-blood agar and a chromogenic medium for the detection of MRSA (bioMérieux, Marcy l’Etoile, France). Presumptive S. aureus colonies were confirmed by coagulase tube test, by use of VITEK2 automated systems (bioMérieux) and by S. aureus-specific PCR [28]. Strains with ambiguous results in phenotypic tests were analyzed by 16 S rRNA gene sequencing. Methicillin-resistance was tested by cefoxitin disc agar diffusion and was confirmed by mecA-specific polymerase chain reaction as described [29]. Susceptibility to antimicrobial agents was tested by VITEK2 automated systems according to recommendations of the manufacturer. The presence of the Panton-Valentine leukocidin (PVL) encoding genes (lukS-PV, lukF-PV) was determined as described elsewhere [30]. Genes of other members of the staphylococcal bicomponent leukotoxin family (lukM, lukE, lukD) were tested as published [31]. Detection of genes of the members of the staphylococcal pyrogenic toxin superantigen (PTSAg) family including the toxic shock syndrome 1 gene (tst), classical and newly described enterotoxin and enterotoxin-like genes (sea, seb, sec 1–3 , sed, seg, seh, sei, sej, sem, sen and seo), exfoliative toxin genes (eta, etb and etd) as well as of genes of the epidermal cell differentiation inhibitor family (edinA, edinB and edinC) and the accessory gene regulatory loci (agrI, agrII, agrIII and agrIV) was carried out using previously reported methods [31, 32]. All MRSA isolates were typed by S. aureus protein A (spa) sequence-based typing as described [33]. Multilocus sequence typing (MLST) was performed on one isolate of each different spa type [34]. Typing of SCCmec elements of types I to IV was carried out for one isolate of every spa type from every farm using a PCR approach, including a combination of different PCRs [35]. To demonstrate SCCmec-elements of type V, we used primers type VF/type VR, as described by Zhang et al. [36], and primer pair ccrC9f 5′-CACTTAATCCATGTACACAG-3′ and ccrC-R [36].
MRSA spa types found on regional pig farms among patients in a tertiary-care university hospital: a case control study
The University Hospital Münster (UHM) is a 1300-bed maximum care teaching hospital in the central part of the Dutch-German border area. Its referral area mostly comprises the regions involved in the screening at pig farms. The MRSA database (Ridom StaphType, Würzburg, Germany) of the UHM includes spa typing results of every first MRSA isolate of each patient for isolates obtained from January 2005 to December 2008. The isolates were obtained from admission screening cultures from the nasopharynx and non-infected skin of patients at the UHM. Screening was performed by swabbing of defined risk patients according to the national German guideline (www.rki.de, version 1.12.99 and comment 12.11.04) between January 2005 and June 2006 and was then replaced by a general nasal screening of all patients at admission. Microbiological diagnostics were performed as described previously [29]. We searched the UHM database for MRSA isolates which were derived from screening samples and which were characterized by spa types which were identified on regional pig farms in this study (t011, t034, t108, t1451, t2510). Using a standardized questionnaire, we interviewed randomly selected patients from whom such presumptive livestock-related MRSA spa types were isolated (n = 100 patients, case group). As a control group, we randomly chose a corresponding group of patients (n = 100) from whom MRSA with other than known livestock-associated spa types were isolated from screening specimens. Randomization was performed using an internet tool (http://www.wesleyan.edu/spn/random/form.htm). The patients were asked whether they shared one or more of the following classical nosocomial risk factors for MRSA acquisition at the time when the MRSA was detected for the first time: contact to another person with MRSA carriage or infection, prescribing of any antibiotics during the previous 6 months, hospitalization for >24 h within the past 6 months, direct transfer of the patient from long-term care facilities to the admitting hospital, need for long-term care, presence of indwelling devices (urinary catheters, intravenous catheters, percutaneous gastrostomies, postsurgical drains), presence of skin lesions (ulcers, eczema, other wounds), need for haemodialysis. Furthermore, the age of the patient, contact to horses, sheep, cattle, pigs and pets (e.g. cats, dogs) were assessed. For statistical analysis with SPSS Statistics 17.0 (SPSS, Chicago, IL, USA), data were entered into an Excel database (Microsoft, Redmond, WA, USA). Univariate analysis was performed using Chi-square test or Fisher exact test for ordinal variables and Student’s t-test for continuous variables. All risk factors associated with a different distribution among the case and control groups with a P-value <0.2 were included in the initial logistic regression model. The backward stepwise method was used to create the models. Choice of the optimal regression model was based on comparison of likelihood ratios and Hosmer-Lemshow goodness-of-fit statistics. A P value <0.05 was considered significant.
Results
Screening on pig-farms
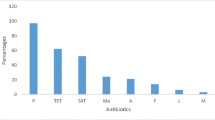
Screening of 1,600 weaned piglets at 40 pig farms in the Dutch-German border area identified nasal MRSA colonization in animals on a total of 28 farms (70%). Among these, two were rearing farms (50% of all rearing farms), seven were farrow-to-finish farms (64% of all farrow-to-finish farms) and 19 were farrowing farms (76% of all farrowing farms). On these farms, MRSA were isolated from all eight culture pools in 16 cases (57%). On four farms MRSA were isolated from 5–7 pools, on three farms from 3–4 pools and on five farms from 1–2 pools. Overall, we isolated 169 MRSA. The isolates were associated with spa types t011 (75%, n = 127), t034 (5.3%, n = 9), t108 (0.6%, n = 1), t1451 (3.6%, n = 6) and t2510 (15.4%, n = 26). MLST results showed that every spa type was associated with ST398. We typed SCCmec elements of one isolate per different spa type found on each farm (in total: n = 33 isolates; t011 (n = 24), t034 (n = 2), t108 (n = 1), t1451 (n = 1), t2510 (n = 5)). All isolates were associated with SCCmec type V with one exception which was an MRSA t011 harbouring SCCmec type IV. None of the MRSA harboured the genes encoding PVL and other leukocidins tested. All isolates were negative for the genes encoding enterotoxins, exfoliative toxins, toxic shock toxin 1 and epidermal cell differentiation inhibitors. All isolates were associated with agr type I. All porcine MRSA strains were resistant to tetracycline, 8 (52%) to erythromycin and clindamycin, 4 (26%) to gentamicin, and 4 (8%) to trimethoprim-sulfomethoxazole. All isolates were susceptible to ciprofloxacin, rifampin, fusidic acid and vancomycin.
Case control study
In the UHM database, we identified 249 different patients from whom MRSA with spa types identified on regional pig farms (t011, t034, t108, t1451, t2510) were isolated from screening samples between January 2005 and December 2008. The MRSA shared spa types t011 (n = 139, 56%), t034 (n = 98, 39%), t108 (n = 8, 3%) and t1451 (n = 4, 2%); spa type t2510 was not detected among patients. These isolates accounted for 13% of all MRSA isolated from screening samples at the UHM in 2005 (n = 16 of 121 isolates), increasing to 15% in 2006 (n = 43 of 286 isolates), 21.9% in 2007 (n = 96 of 438 isolates) and 22.4% in 2008 (n = 94 of 419 isolates). Among the patients colonized by presumptive livestock-related MRSA, 100 were available for telephone interviews and were included in the case group. The 100 patients within the control group were randomly selected out of 1,015 MRSA isolates obtained from screening between January 2005 and December 2008 and exhibiting spa types which were not found on regional pig farms. These 1,015 isolates were associated with a total of 136 (predominantly classical nosocomial) spa types among which nine spa types were outstanding: t003 (34%), t032 (28%), t004 (4%), t001, t008, t014 (each 2%), t002, t015, and t065 (each 1%). All other spa types were found in less than 1% of the isolates.
Isolates within the case group exhibited spa types t011 (n = 57), t034 (n = 39), t108 (n = 3) and t1451 (n = 1). Those within the control group were associated with 26 spa types including t003 (n = 39), t032 (n = 29), t004 (n = 5), t002, t008, t014, t061 (each n = 2), t001, t012, t018, t020, t022, t024, t038, t045, t065 t068, t069, t091, t256, t284, t578, t660, t1329, t1996, and t4111 (each n = 1).
Since the age of the patients differed between the case and control groups (P = 0.06; case group mean/median age: 48.2/48 years; control group mean/median age 53.7/56 years), age categories were built and included in the statistical analysis (Table 1). At least one of the classical nosocomial risk factors for MRSA acquisition was identified among n = 64 (64%) of patients in the case group and among n = 91 (91%) of patients in the control group (P < 0.001). Table 1 shows in detail the distribution of risk factors in the case and control groups as well as the results of univariate analysis. The final logistic regression model included the variables: age 3–18 years, contact to pigs, to cattle and to horses, chronic need for nursing care, presence of indwelling devices and chronic skin lesions. In this model, the risk factors contact with pigs (OR = 20.455; 95% confidence interval (CI) 7.831–64.386), contact with cattle (OR = 8.607; 95% CI 1.729–42.854) and age between 3 and 18 years (OR = 9.629; 95% CI 1.374–67.476) were independently associated with the case group (Table 1). The risk factor chronic need for nursing care (OR = 0.065; 95% CI 0.008–0.524) was less frequent among patients in the case group.
Discussion
In 2003, a new MRSA clone exhibiting MLST CC398 (spa types t108, t011 and relatives) emerged among patients in Dutch hospitals. Since the occurrence of this MRSA type was shown to be epidemiologically linked to regions with a high density of pig farming [37], and as screening of pigs yielded high colonization rates [9], it was assumed that pigs might be a potential zoonotic source for MRSA transmissions.
It is unknown how often pigs at German farms, which are major European pig producers (Eurostat yearbook 2008, http://epp.eurostat.ec.europa.eu), are colonized with MRSA. In this study, we identified MRSA colonization in pigs on 70% of the regional farms. This corresponds to data from The Netherlands where 23–80% of the pig farms were affected [9, 38], and it clearly reflects that porcine MRSA colonization is a common phenomenon on farms in Germany, at least in the study area. However, genotypic investigations on MRSA isolates from pigs with acute diseases (urinary tract infection, skin infection, metritis mastitis agalactia syndrome) [16] and screening of clinical specimens and cadavers [24] recently indicated that other German geographic regions may also be involved.
Molecular typing revealed that the regional pigs’ isolates were associated with a subset of spa types associated with MLST CC398, agr type I and SCCmec types V and IV, as observed in other countries [9, 18]. All isolates derived from pigs in this study did not harbour genes encoding exfoliative toxin, the toxic shock syndrome toxin 1 and enterotoxins. Moreover, the porcine isolates lacked leukocidal toxin genes lukE/lukD, lukM and the PVL-encoding lukS/lukF. These results confirm previous investigations [38] and suggest a low capacity of regionally disseminated porcine MRSA subtypes to induce toxin-mediated diseases among humans. The importance of these findings is highlighted against the background that MRSA CC398 have entered the food chain [22] and an outbreak of foodborne disease caused by enterotoxin-producing MRSA exhibiting a different (non-CC398) molecular type has been described previously [39]. Hence, a close surveillance of S. aureus from different species and from food samples should reveal the potential emergence of toxin-producing MRSA isolates.
Logistic regression modelling in our case control study yielded that, in addition to contact with pigs, contact with cattle was also independently associated with an increased risk for MRSA CC398 colonization. This finding confirms recent observations by van Loo et al. who also found a correlation of human MRSA CC398 carriage and contact to cattle [37]. Since no data on the prevalence of MRSA among cattle on German farms are published, further studies are warranted to investigate this issue.
It has previously been observed that rates of antimicrobial resistance in the commensal flora of (pig) farmers are significantly higher than in the general population [40]. The recent emergence of MRSA among farmers [17] therefore might reflect another facet of a long-known phenomenon. Nevertheless, we showed that the proportion of MRSA with presumptively livestock-associated spa types among all MRSA isolated from admission screenings at the UHM increased from 13% in 2005 to 22.4% in 2008. This proportion is higher than suggested by national German surveillance data [20, 41]. It has been confirmed that MRSA CC398 are not only found locally at the UHM, but are frequently isolated from admission screenings in an entire region of Northwestern Germany [42, 43]. The high burden of this MRSA subtype might therefore primarily reflect a regional situation, which could be explainable by the fact that the region involved is the area with the highest pig density (number of pigs/km2) in Germany (information.medien.agrar e.V., Agraratlas Jan 2005, http://www.ima-agrar.de).
In this retrospective case control study, we showed a significant association of the spa types t011 and relatives with contact to pigs and cattle. We recognize the limitations (recall bias) inherent within this study type and state that our findings, which represent correlation data, are not suitable for proving a direct transmission of MRSA between animals and humans. However, we demonstrated that 36% of presumptive livestock-related MRSA carriers lacked classical risk factors hitherto used for admission screening of patients. Moreover, carriers of MRSA CC398 in this study comprised younger patient groups who are usually less frequent carriers of MRSA [42] and less frequently associated with classical MRSA risk factors. This raises the question of whether commonly used recommendations for admission screening are convenient to detect patients harbouring livestock-associated strains. Therefore, we favour considering adding “contact to livestock” as a risk factor to the admission screening schedule for hospitals, at least in regions with a high density of farming. Otherwise the floodgates for a clonal spread of MRSA CC398 could be opened in these regions. Furthermore, efforts of molecular MRSA surveillance and reports on prevalence will fail to reflect the actual epidemiological situation. In The Netherlands, current reports show that an adaptation of the on-admission MRSA screening protocol in this manner revealed that 32% of pig-exposed patients admitted to a Dutch hospital were colonized by MRSA. This led to a three-fold increase in the local MRSA incidence in a predominantly agriculturally structured region [44]. Moreover, our observation that the increase of MRSA CC398 at the UHM was related to the introduction of a new local MRSA “search policy” performing screening in all patients, irrespective of risk factors, emphasizes the importance of the risk patient definition for the identification of these isolates. This is highlighted by the threat that continuous introduction of MRSA CC398 to the hospital setting, might cause nosocomial spread to patient groups with underlying comorbidities and thereby facilitate infections. This threat has recently been increased by the first nosocomial outbreak of MRSA ST398 in a Dutch hospital [45].
Apart from giving impetus to antimicrobial resistance import to hospitals, colonization with MRSA CC398 might set livestock-exposed persons under risk of developing severe S. aureus CC398-associated infections [21, 46]. First, this highlights the special need to consider MRSA as causative agents when treating such patient groups. Second, it raises the question as to whether the implementation of preventive measures (consequent alcoholic hand disinfection, wearing of gloves and prevention of dust inhalation) could forestall MRSA colonization of livestock-exposed workers.
In conclusion, our findings confirm that MRSA associated with spa type t011 and relatives can be found on German pig farms and indicate that livestock might represent a relevant source for MRSA imports to hospitals.
References
Deurenberg RH, Vink C, Kalenic S, Friedrich AW, Bruggeman CA, Stobberingh EE (2007) The molecular evolution of methicillin-resistant Staphylococcus aureus. Clin Microbiol Infect 13:222–235
Walther B, Monecke S, Ruscher C, Friedrich AW, Ehricht R, Slickers P, Soba A, Wleklinski CG, Wieler LH, Lubke-Becker A (2009) Comparative molecular analysis substantiates a zoonotic potential of equine methicillin-resistant Staphylococcus aureus (MRSA). J Clin Microbiol 47(3):704–710
Weese JS, Lefebvre SL (2007) Risk factors for methicillin-resistant Staphylococcus aureus colonization in horses admitted to a veterinary teaching hospital. Can Vet J 48:921–926
Cuny C, Kuemmerle J, Stanek C, Willey B, Strommenger B, Witte W (2006) Emergence of MRSA infections in horses in a veterinary hospital: strain characterisation and comparison with MRSA from humans. Euro Surveill 11:44–47
Juhasz-Kaszanyitzky E, Janosi S, Somogyi P, Dan A, van der Graaf-van Bloois L, van Duijkeren E, Wagenaar JA (2007) MRSA transmission between cows and humans. Emerg Infect Dis 13:630–632
Moodley A, Stegger M, Bagcigil AF, Baptiste KE, Loeffler A, Lloyd DH, Williams NJ, Leonard N, Abbott Y, Skov R, Guardabassi L (2006) spa typing of methicillin-resistant Staphylococcus aureus isolated from domestic animals and veterinary staff in the UK and Ireland. J Antimicrob Chemother 58:1118–1123
Walther B, Wieler LH, Friedrich AW, Hanssen AM, Kohn B, Brunnberg L, Lubke-Becker A (2007) Methicillin-resistant Staphylococcus aureus (MRSA) isolated from small and exotic animals at a university hospital during routine microbiological examinations. Vet Microbiol 127:171–178
Rich M, Roberts L (2006) MRSA in companion animals. Vet Rec 159:535–536
De Neeling AJ, van den Broek MJ, Spalburg EC, van Santen-Verheuvel MG, Dam-Deisz WD, Boshuizen HC, van de Giessen AW, van Duijkeren E, Huijsdens XW (2007) High prevalence of methicillin resistant Staphylococcus aureus in pigs. Vet Microbiol 122:366–372
Guardabassi L, Stegger M, Skov R (2007) Retrospective detection of methicillin resistant and susceptible Staphylococcus aureus ST398 in Danish slaughter pigs. Vet Microbiol 122:384–386
Lewis HC, Molbak K, Reese C, Aarestrup FM, Selchau M, Sorum M, Skov RL (2008) Pigs as source of Methicillin-resistant Staphylococcus aureus CC398 infections in humans, Denmark. Emerg Infect Dis 14:1383–1389
Armand-Lefevre L, Ruimy R, Andremont A (2005) Clonal comparison of Staphylococcus aureus isolates from healthy pig farmers, human controls, and pigs. Emerg Infect Dis 11:711–714
Khanna T, Friendship R, Dewey C, Weese JS (2007) Methicillin resistant Staphylococcus aureus colonization in pigs and pig farmers. Vet Microbiol 128:298–303
Smith TC, Male MJ, Harper AL, Kroeger JS, Tinkler GP, Moritz ED, Capuano AW, Herwaldt LA, Diekema DJ (2008) Methicillin-resistant Staphylococcus aureus (MRSA) strain ST398 is present in midwestern U.S. swine and swine workers. PLoS ONE 4:e4258
Sergio DMB, Koh TH, Hsu L, Ogden BE, Goh ALH, Chow PKH (2007) Investigation of methicillin-resistant Staphylococcus aureus in pigs used for research. J Med Microbiol 56:1107–1109
Schwarz S, Kadlec K, Strommenger B (2008) Methicillin-resistant Staphylococcus aureus and Staphylococcus pseudintermedius detected in the BfT-GermVet monitoring programme 2004–2006 in Germany. J Antimicrob Chemother 61:282–285
Voss A, Loeffen F, Bakker J, Klaassen C, Wulf M (2005) Methicillin-resistant Staphylococcus aureus in pig farming. Emerg Infect Dis 11:1965–1966
Huijsdens XW, van Dijke BJ, Spalburg E, van Santen-Verheuvel MG, Heck MEOC, Pluister GN, Voss A, Wannet WJB, de Neeling AJ (2006) Community-acquired MRSA and pig-farming. Ann Clin Microbiol Antimicrob 5:26
Wulf M, van Nes A, Eikelenboom-Boskamp A, de Vries J, Melchers W, Klaassen C, Voss A (2006) Methicillin-resistant Staphylococcus aureus in veterinary doctors and students, the Netherlands. Emerg Infect Dis 12:1939–1941
Witte W, Strommenger B, Stanek C, Cuny C (2007) Methicillin-resistant Staphylococcus aureus ST398 in humans and animals, Central Europe. Emerg Infect Dis 13:255–258
Ekkelenkamp MB, Sekkat M, Carpaij N, Troelstra A, Bonten MJM (2006) Endocarditis due to methicillin-resistant Staphylococcus aureus originating from pigs. Ned Tijdschr Geneeskd 150:2442–2447
van Loo IHM, Diederen BMW, Savelkoul PHM, Woudenberg JHC, Roosendaal R, van Belkum A, Lemmens-den Toom N, Verhulst C, van Keulen PHJ, Kluytmans JAJW (2007) Methicillin-resistant Staphylococcus aureus in meat products, the Netherlands. Emerg Infect Dis 13:1753–1755
Pu S, Han F, Ge B (2009) Isolation and characterization of methicillin-resistant Staphylococcus aureus strains from Louisiana retail meats. Appl Environ Microbiol 75:265–267
Meemken D, Cuny C, Witte W, Eichler U, Staudt R, Blaha T (2008) Occurrence of MRSA in pigs and in humans involved in pig production–preliminary results of a study in the northwest of Germany. Dtsch Tierarztl Wochenschr 115:132–139
Tiemersma EW, Bronzwaer SLAM, Lyytikainen O, Degener JE, Schrijnemakers P, Bruinsma N, Monen J, Witte W, Grundman H (2004) Methicillin-resistant Staphylococcus aureus in Europe, 1999–2002. Emerg Infect Dis 10:1627–1634
Harbarth S, Sax H, Fankhauser-Rodriguez C, Schrenzel J, Agostinho A, Pittet D (2006) Evaluating the probability of previously unknown carriage of MRSA at hospital admission. Am J Med 119(275):e15–e23
Wertheim H, Verbrugh HA, van Pelt C, de Man P, van Belkum A, Vos MC (2001) Improved detection of methicillin-resistant Staphylococcus aureus using phenyl mannitol broth containing aztreonam and ceftizoxime. J Clin Microbiol 39:2660–2662
Martineau F, Picard FJ, Roy PH, Ouellette M, Bergeron MG (1998) Species-specific and ubiquitous-DNA-based assays for rapid identification of Staphylococcus aureus. J Clin Microbiol 36:618–623
Becker K, Pagnier I, Schuhen B, Wenzelburger F, Friedrich AW, Kipp F, Peters G, von Eiff C (2006) Does nasal cocolonization by methicillin-resistant coagulase-negative staphylococci and methicillin-susceptible Staphylococcus aureus strains occur frequently enough to represent a risk of false-positive methicillin-resistant S. aureus determinations by molecular methods? J Clin Microbiol 44:229–231
Lina G, Piemont Y, Godail-Gamot F, Bes M, Peter MO, Gauduchon V, Vandenesch F, Etienne J (1999) Involvement of Panton-Valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis 29:1128–1132
von Eiff C, Friedrich AW, Peters G, Becker K (2004) Prevalence of genes encoding for members of the staphylococcal leukotoxin family among clinical isolates of Staphylococcus aureus. Diagn Microbiol Infect Dis 49:157–162
Becker K, Friedrich AW, Lubritz G, Weilert M, Peters G, von Eiff C (2003) Prevalence of genes encoding pyrogenic toxin superantigens and exfoliative toxins among strains of Staphylococcus aureus isolated from blood and nasal specimens. J Clin Microbiol 41:1434–1439
Mellmann A, Friedrich AW, Rosenkotter N, Rothganger J, Karch H, Reintjes R, Harmsen D (2006) Automated DNA sequence-based early warning system for the detection of methicillin-resistant Staphylococcus aureus outbreaks. PLoS Med 3:e33
Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG (2000) Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol 38:1008–1015
Strommenger B, Kehrenberg C, Kettlitz C, Cuny C, Verspohl J, Witte W, Schwarz S (2006) Molecular characterization of methicillin-resistant Staphylococcus aureus strains from pet animals and their relationship to human isolates. J Antimicrob Chemother 57:461–465
Zhang K, McClure JA, Elsayed S, Louie T, Conley JM (2005) Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol 43:5026–5033
van Loo I, Huijsdens X, Tiemersma E, de Neeling A, van de Sande-Bruinsma N, Beaujean D, Voss A, Kluytmans J (2007) Emergence of methicillin-resistant Staphylococcus aureus of animal origin in humans. Emerg Infect Dis 13:1834–1839
van Duijkeren E, Ikawaty R, Broekhuizen-Stins MJ, Jansen MD, Spalburg EC, de Neeling AJ, Allaart JG, van Nes A, Wagenaar JA, Fluit AC (2008) Transmission of methicillin-resistant Staphylococcus aureus strains between different kinds of pig farms. Vet Microbiol 126:383–389
Jones TF, Kellum ME, Porter SS, Bell M, Schaffner W (2002) An outbreak of community-acquired foodborne illness caused by methicillin-resistant Staphylococcus aureus. Emerg Infect Dis 8:82–84
Aubry-Damon H, Grenet K, Sall-Ndiaye P, Che D, Cordeiro E, Bougnoux M, Rigaud E, Le Strat Y, Lemanissier V, Armand-Lefevre L, Delzescaux D, Desenclos J, Lienard M (2004) Antimicrobial resistance in commensal flora of pig farmers. Emerg Infect Dis 10:873–839
Strommenger B, Braulke C, Heuck D, Schmidt C, Pasemann B, Nubel U, Witte W (2008) spa Typing of Staphylococcus aureus as a frontline tool in epidemiological typing. J Clin Microbiol 46:574–581
Köck R, Brakensiek L, Mellmann A, Kipp F, Henderikx M, Harmsen D, Daniels-Haardt I, von Eiff C, Becker K, Hendrix MG, Friedrich AW (2009) Cross-border comparison of the admission prevalence and clonal structure of methicillin-resistant Staphylococcus aureus. J Hosp Infect 71:320–326
Friedrich AW, Daniels-Haardt I, Köck R, Verhoeven F, Mellmann A, Harmsen D, van Gemert-Pijnen JE, Becker K, Hendrix MGR (2008) EUREGIO MRSA-net Twente/Munsterland—a Dutch-German cross-border network for the prevention and control of infections caused by methicillin-resistant Staphylococcus aureus. Euro Surveill 13(pii):18965
van Rijen MML, Van Keulen PH, Kluytmans JA (2008) Increase in a Dutch hospital of methicillin-resistant Staphylococcus aureus related to animal farming. Clin Infect Dis 46:261–263
Wulf MWH, Markestein A, van der Linden FT, Voss A, Klaassen C, Verduin CM (2008) First outbreak of methicillin-resistant Staphylococcus aureus ST398 in a Dutch hospital, June 2007. Euro Surveill 28(13)pii:8051
van Belkum A, Melles DC, Peeters JK, van Leeuwen WB, van Duijkeren E, Huijsdens XW, Spalburg E, de Neeling AJ, Verbrugh HA, On Behalf Of The Dutch Working Party On Surveillance And Research (2008) Methicillin-resistant and -susceptible Staphylococcus aureus sequence type 398 in pigs and humans. Emerg Infect Dis 14:479–483
Acknowledgments
This study was partly performed within the framework of the INTERREG-IIIa project “EUREGIO MRSA-net Twente/Münsterland” (reference no. 2-EUR-V-1 = 96) and the INTERREG-IVa Euregio-project Safeguard MRSA VetMed-net (reference no. 34-INTERREG IV A-II-2-05=025).
We are especially grateful to M.G. Ron Hendrix (MD, PhD), Laboratorium Microbiologie, Enschede, for fruitful discussion, and we thank Monique Henderikx, Isabell Ramminger and Ursula Keckevoet for their skilled technical assistance.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Köck, R., Harlizius, J., Bressan, N. et al. Prevalence and molecular characteristics of methicillin-resistant Staphylococcus aureus (MRSA) among pigs on German farms and import of livestock-related MRSA into hospitals. Eur J Clin Microbiol Infect Dis 28, 1375–1382 (2009). https://doi.org/10.1007/s10096-009-0795-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-009-0795-4