Abstract
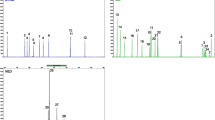
A multiplex real-time PCR method for discriminating deer and common domestic species, including cattle, goat, horse, donkey, pig, and chicken was developed. Species-specific primer pairs were designed and used to produce different size DNA fragments with diverse melting temperature (T m ) values. The specificity and sensitivity of these primer pairs were separately confirmed using simplex real-time PCR analysis. Multiplex real-time PCR analysis was performed using combined primers, yielding distinct melting curve profiles for each species. The sensitivity limit of the multiplex PCR method was evaluated. Trace DNA of other species in deer DNA could be identified. Common deer products, including blood, meat, and antler were tested using this multiplex PCR method and different species of deer and domestic animals were identified. The rapid multiplex real-time PCR assay described herein is a specific, sensitive, and reliable method for high-throughput authentication of deer and domestic animal products.
Similar content being viewed by others
References
Wang BX, Zhao XH, Qi SB, Kaneko S, Hattori M, Namba T, Nomura Y. Effects of repeated administration of deer antler extract on biochemical changes related to aging in senescence-accelerated mice. Chem. Pharm. Bull. (Tokyo) 36: 2587–2592 (1988)
Zhou QL, Guo YJ, Wang LJ, Wang Y, Liu YQ, Wang BX. Velvet antler polypeptides promoted proliferation of chondrocytes and osteoblast precursors and fracture healing. Zhongguo Yao Li Xue Bao 20: 279–282 (1999)
Kim KS, Choi YH, Kim KH, Lee YC, Kim CH, Moon SH, Kang SG, Park YG. Protective and anti-arthritic effects of deer antler aquaacupuncture (DAA), inhibiting dihydroorotate dehydrogenase, on phosphate ions-mediated chondrocyte apoptosis and rat collageninduced arthritis. Int. Immunopharmacol. 4: 963–973 (2004)
Shi B, Li G, Wang P, Yin W, Sun G, Wu Q, Yu G. Effect of antler extract on corticosteroid-induced avascular necrosis of the femoral head in rats. J. Ethnopharmacol. 127: 124–129 (2010)
Tseng SH, Sung HC, Chen LG, Lai YJ, Wang KT, Sung CH, Wang CC. Effects of velvet antler with blood on bone in ovariectomized rats. Molecules 17: 10574–10585 (2012)
Ghovvati S, Nassiri MR, Mirhoseini SZ, Moussavi AH, Javadmanesh A. Fraud identification in industrial meat products by multiplex PCR assay. Food Control 20: 696–699 (2009)
Mafra I, Ferreira I, Oliveira MB. Food authentication by PCR-based methods. Eur. Food Res. Technol. 227: 649–665 (2008)
Gachet E, Martin GG, Vigneau F, Meyer G. Detection of genetically modified organisms (GMOs) by PCR: A brief review of methodologies available. Trends Food Sci. Technol. 9: 380–388 (1998)
Zha D-M, Xing X-M, Yang F-H. Rapid identification of deer products by multiplex PCR assay. Food Chem. 129: 1904–1908 (2011)
Kim ES, Kim YH, Ko BS, Oh SE, Do EJ, Lee MY. Multiplex polymerase chain reaction (PCR) and fluorescence-based capillary electrophoresis for identification of deer species from antlers. Afr. J. Biotechnol. 11: 3179–3086 (2012)
Zha D, Xing X, Yang F. A multiplex PCR assay for fraud identification of deer products. Food Control 21: 1402–1407 (2010)
Fajardo V, Gonzalez I, Lopez-Calleja I, Martin I, Hernandez PE, Garcia T, Martin R. PCR-RFLP authentication of meats from red deer (Cervus elaphus), fallow deer (Dama dama), roe deer (Capreolus capreolus), cattle (Bos taurus), sheep (Ovis aries), and goat (Capra hircus). J. Agr. Food Chem. 54: 1144–1150 (2006)
Fajardo V, Gonzalez I, Lopez-Calleja I, Martin I, Rojas M, Hernandez PE, Garcia T, Martin R. Identification of meats from red deer (Cervus elaphus), fallow deer (Dama dama), and roe deer (Capreolus capreolus) using polymerase chain reaction targeting specific sequences from the mitochondrial 12S rRNA gene. Meat Sci. 76: 234–240 (2007)
Fajardo V, Gonzalez I, Martin I, Rojas M, Hernandez PE, Garcia T, Martin R. Real-time PCR for detection and quantification of red deer (Cervus elaphus), fallow deer (Dama dama), and roe deer (Capreolus capreolus) in meat mixtures. Meat Sci. 79: 289–298 (2008)
Mafra I, Ferreira IPLVO, Oliveira MBP. Food authentication by PCR-based methods. Eur. Food Res. Technol. 227: 649–665 (2008)
Fajardo V, Gonzalez I, Rojas M, Garcia T, Martin R. A review of current PCR-based methodologies for the authentication of meats from game animal species. Trends Food Sci. Technol. 21: 408–421 (2010)
Dooley JJ, Paine KE, Garrett SD, Brown HM. Detection of meat species using TaqMan real-time PCR assays. Meat Sci. 68: 431–438 (2004)
Kesmen Z, Gulluce A, Sahin F, Yetim H. Identification of meat species by TaqMan-based real-time PCR assay. Meat Sci. 82: 444–449 (2009)
Kesmen Z, Yetiman AE, Sahin F, Yetim H. Detection of chicken and turkey meat in meat mixtures by using real-time PCR assays. J. Food Sci. 77: C167–C173 (2012)
Sakalar E, Abasiyanik MF. The devolopment of duplex real-time PCR based on SYBR Green florescence for rapid identification of ruminant and poultry origins in foodstuff. Food Chem. 130: 1050–1054 (2012)
Soares S, Amaral JS, Oliveira MB, Mafra I. A SYBR Green realtime PCR assay to detect and quantify pork meat in processed poultry meat products. Meat Sci. 94: 115–120 (2013)
Ponchel F, Toomes C, Bransfield K, Leong F, Douglas S, Field S, Bell S, Combaret V, Puisieux A, Mighell A, Robinson P, Inglehearn C, Isaacs J, Markham A. Real-time PCR based on SYBR-Green I fluorescence: An alternative to the TaqMan assay for a relative quantification of gene rearrangements, gene amplifications and micro gene deletions. BMC Biotechnol. 3: 18 (2003)
Buh Gasparic M, Tengs T, La Paz JL, Holst-Jensen A, Pla M, Esteve T, Zel J, Gruden K. Comparison of nine different real-time PCR chemistries for qualitative and quantitative applications in GMO detection. Anal. Bioanal. Chem. 396: 2023–2029 (2010)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
You, J., Huang, L., Zhuang, J. et al. Species-specific multiplex real-time PCR assay for identification of deer and common domestic animals. Food Sci Biotechnol 23, 133–139 (2014). https://doi.org/10.1007/s10068-014-0018-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-014-0018-3