Abstract
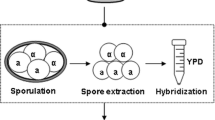
Fermentation properties of top-fermenting yeast are under the control of multiple genes difficult to manipulate directly by classical breeding, metabolic engineering, or other genetic methods with specific genes or pathways as target. Here, genome shuffling is introduced to improve fermentation performance (such as the viability of the yeast, flavor of beer, and the fermentation time) by improving wort and ethanol tolerance of top-fermenting yeast. The strategy was performed not based on polyethylene glycol (PEG)-mediated protoplast fusion but using yeast sexual and asexual reproduction by itself. The best performing strain W3-8 was selected on the selective plates after 3 rounds of genome shuffling. The fermentation time of W3-8 was not only markedly shortened, but also, most flavor compounds were distinctly improved. In particular, ethanol yield was increased by up to 67% after the 3rd pitching compared with the control. Furthermore, W3-8 promoted desired amounts of esters and higher alcohols, in accordance with specific consumer preferences. Significant improvement in the fermentation traits of the top-fermenting yeast was achieved using genome shuffling.
Similar content being viewed by others
References
Saerens SMG, Verbelen PJ, Vanbeneden N, Thevelen JM, Delvaux FR. Monitoring yeast physiology during very high gravity wort fermentations by frequent analysis of gene expression. Appl. Microbiol. Biot. 24: 741–760 (2007)
Gibson BR, Lawrence SJ, Leclaire JPR, Powell CD, Smart KA. Yeast responses to stresses associated with industrial brewery handling. FEMS Microbiol. Rev. 31: 535–569 (2007)
Zhang YX, Perry K, Vinci VA, Powell K, Stemmer WPC, Cardayre S. Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415: 644–646 (2002)
Patnaik R, Louie S, Gavrilovic V, Stemmer WPC, Ryan CM, Cardayre S. Genome shuffling of lactobacillus for improved acid tolerance. Nat. Biotechnol. 20: 707–712 (2002)
Dai MH, Copley SD. Genome shuffling improves degradation of the anthropogenic pesticide pentachlorophenol by Sphingobium chlorophenolicum ATCC 39723. Appl. Environ. Microb. 70: 2391–2397 (2004)
Hida H, Yamada T, Yamada Y. Genome shuffling of Streptomyces sp. U121 for improved production of hydroxycitric acid. Appl. Microbiol. Biot. 73: 1387–1393 (2007)
Yu L, Pei X, Lei T, Wang Y, Feng Y. Genome shuffling enhanced Llactic acid production by improving glucose tolerance of lactobacillus rhamnosus. J. Biotechnol. 134: 154–159 (2008)
Sigler K, Matoulková D, Dienstbier M, Gabriel P. Net effect of wort osmotic pressure on fermentation course, yeast vitality, beer flavor, and haze. Appl. Microbiol. Biot. 82: 1027–1035 (2009)
Lawrence CW. Methods in Enzymology. Part A. 194. Elesevier Academic Press, NY, New York, USA. pp. 277–278 (2004)
Herman PK, Rine J. Yeast spore germination: A requirement for ras protein activity during re-entry into the cell cycle. EMBO J. 16: 6171–6181 (1997)
Hou L. Novel methods of genome shuffling used in Saccharomyces cerevisiae. Biotechnol. Lett. 31: 671–677 (2009)
Farahnak F, Seki T, Ryu DD, Ogrydziak D. Construction of lactoseassimilating and high-ethanol-producing yeasts by protoplast fusion. Appl. Environ. Microb. 51: 362–367 (1986)
Sigler K, Mikyka A, Kosa K, Gabriel P, Diensbier M. Factors affecting the outcome of the acidification power test of yeast quality: Critical reappraisal. Folia Microbiol. 51: 525–534 (2006)
Gabriel P, Dienstbier M, Matoulková D, Kosa Optimized acidification power test of yeast vitality and its use in brewing practice. J. I. Brewing 114: 270–276 (2008)
Opekarová M, Sigler K. Acidification power: Indicator of metabolic activity and autolytic changes in Saccharomyces cerevisiae. Folia Microbiol. 27: 395–403 (1982)
Gong J, Zheng H, Wu Z, Chen T, Zhao X. Genome shuffling: Progress and applications for phenotype improvement. Biotechnol. Adv. 27: 996–1005 (2009)
Kubota S, Takeo I, Kume K, Kani M, Shitamukai A, Mizunuma M, Miyakawa T, Shimoi H, Iefuji H, Hirata D. Effect of ethanol on cell growth of budding yeast: Genes that are important for cell growth in the presence of ethanol. Biosci. Biotech. Bioch. 68: 968–972 (2004)
Carlson CR, Grallert B, Bernander R, Stokke T, Boye E. Measurement of nuclear DNA content in fission yeast by flow cytometry. Yeast 13: 1329–1335 (1997)
Wei P, Li Z, He P, Lin Y, Jiang N. Genome shuffling of ethanologenic yeast Candida krusei for improved acetic acid tolerance. Biotechnol. Appl. Bioc. 49: 113–128 (2008)
Sigler K, Matoulková D, Dienstbier M, Gabriel P. Net effect of wort osmotic pressure on fermentation course, yeast vitality, beer flavor, and haze. Appl. Microbiol. Biot. 82: 1027–1035 (2009)
Casey GP, Magnus CA, Ingledew WM. High-gravity brewing: Effects of nutrition on yeast composition, fermentative ability, and alcohol production. Appl. Environ. Microb. 48: 639–646 (1984)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, H., Hou, L. Genome shuffling to improve fermentation properties of top-fermenting yeast by the improvement of stress tolerance. Food Sci Biotechnol 19, 145–150 (2010). https://doi.org/10.1007/s10068-010-0020-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-010-0020-3