Abstract
Unlike the 1p36 microdeletion syndrome, which has been extensively described, 1p36.3 microduplications have rarely been reported. We report the two siblings of familial 1p36.3 microduplication, presenting with a severe global developmental delay, epilepsy, and a few dysmorphic features. They were referred to moderate-to-severe developmental delay (DD) and intellectual disability (ID). Both were considered eyelid myoclonus with absence of epilepsy (Jeavons syndrome). The EEG is characterized by widespread 2.5–3.5 Hz spikes and spike slow complex wave, eye closure sensitivity, and photosensitivity. The children has same dysmorphic features, including mild bitemporal narrowing and sloping forehead, sparse eyebrows, hypertelorism, ptosis, strabismus, infraorbital creases, wide nasal bridge with bulbous nasal tip, dystaxia, hallux valgus, and flat feet. Family exome sequencing revealed a maternally inherited 3.2-Mb microduplication of chromosomal band 1p36.3p36.2. However, DNA purified from blood samples of either parent did not find evidence for a microduplication of 1p36 in somatic tissue, indicating that such a mutation might be carried in the germline of the parents as gonadal mosaicism. No other family members of the affected siblings’ parents were reported to be affected by the symptoms found.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The 1p36.3 chromosomal region is prone to microrearrangements leading to an abnormal phenotype. The 1p36.3 microdeletion syndrome is one of the most frequent microdeletion syndromes and its phenotypical spectrum is well known. Several critical regions have been defined as causative for several phenotypical features associated with the syndrome, and positional candidate genes have thus been identified. Conversely, isolated 1p36.3 microduplications have rarely been reported. This can be at least partly explained by an ascertainment bias, chromosomal duplications being classically considered to have a milder phenotypical effect than corresponding deletions. Indeed most of the described patients with a 1p36.3 microduplication also carry another chromosomal rearrangement, whether a 1p36 microdeletion or another deletion or duplication. Thus, it is very difficult to delineate a phenotypical spectrum for 1p36.3 microduplications.
Clinical report
Description of the family
We describe a family consisting of a 35-year-old mother (individual II in Fig. 1), a 36-year-old father (individual II in Fig. 1), and their two children. The elder daughter (individual III in Fig. 1) was 1 year old and her brother (individual III in Fig. 1) was 2 years old at first examination. Both children presenting with moderate-to-severe developmental delay (DD) and intellectual disability (ID), another important feature is epilepsy. The mother (individual II in Fig. 1) and father were normal. The mother has two sisters and one brother (individuals II in Fig. 1), who had their children had a normal development. The father has one sister and one brother (individuals II in Fig. 1), and their children had normal development (Fig. 2).
Clinical examination
Proband
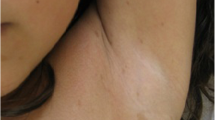
The first child of this family, a young girl, was referred to moderate-to-severe developmental delay (DD) and intellectual disability (ID). She was born at 39 weeks of gestation, denied history of hypoxia and trauma, birth weight, height, and OFC were normal, and poor feeding in the neonatal period. She was able to sit after 1 year of age and could not walk without support at the age of 12 months. Her language is also delayed, consisting of a few simple words at the age of 18 months. Her medical history was marked by eyelid myoclonus with absence of epilepsy (Jeavons syndrome), treated effectively with sodium valproate since the age of 2. EEG (Figs. 3, 4, 5, and 6): widespread 2.5–3.5 Hz spikes and spike slow complex wave, eye closure sensitivity, and photosensitivity. Wechsler Intelligence Scale score is 72. MRI showed bilateral hippocampal sclerosis. She has an unusual behavior with some autistic signs: she is motionless and impassive with poor interactions and reduced interest in other people or toys. The child has dysmorphic features, including mild bitemporal narrowing and sloping forehead, sparse eyebrows, hypertelorism, ptosis, strabismus, infraorbital creases, wide nasal bridge with bulbous nasal tip, dystaxia, hallux valgus, and flat feet (Fig. 2).
Eyelid myoclonus with absence in the sister at the age of 5 years, with rapid blinking during play with daze and slight loss of consciousness lasting approximately 5 s in the waking period, and spike-and-slow wave bursts of 2.5–3.5 Hz visible in the full EEG conduction (synchronized with action termination)
Proband’s younger brother
The second child of the family, a boy, was born at term after an uneventful pregnancy. Birth weight, height, and OFC were normal. His medical history was marked by eyelid myoclonus with absence of epilepsy (Jeavons syndrome), and treated effectively with sodium valproate since the age of 4. Like his elder brother, she has a mild developmental delay. At physical examination, mild bitemporal narrowing and sloping forehead, sparse eyebrows, hypertelorism, ptosis, strabismus, infraorbital creases, wide nasal bridge with bulbous nasal tip, dystaxia, hallux valgus, and flat feet were noted (Fig. 2).
Family exome sequencing results
Materials and methods
Genomic DNA from peripheral blood was extracted using the Whole Blood Genomic DNA Extraction Kit (Beijing Makino Medical Laboratory Institute, Beijing, China) according to the standard operating procedure (SOP). The DNA library was generated by polymerase chain reaction (PCR) using the KAPA Library Preparation Kit (KAPA Biosystems, Boston, USA) according to the SOP. The whole exome was captured using the SureSelect Human All Exon Kit V6 (Agilent Technologies, CA, USA). The target region was sequenced in high throughput using the NovaSeq 6000 platform (Illumina, CA, USA). CNV-sequencing was performed to confirm the pathogenic mutations detected by WES.
Results
The siblings were tested for the same mutation: seq[hg19]dup(1)(p36.33p36.32) chrl: g.621095–3816818dup. The younger brother was sick and also carried the duplication mutation in this segment. However, the variant was not detected in the blood samples of the patient's parents (Fig. 7). The presence of gonadal mosaicism in one of the parents was highly suspected.
There were 73 protein-coding genes in the CNV (score = 0.9) (Fig. 8). On the basis of these considerations, we are concerned that the evidence is insufficient for pathogenicity. We searched the DECIPHER database for cases of shorter CNV that were covered by the duplication found in this study. This duplication is associated with 1p36.3 microduplication syndrome, which is consistent; this duplication is associated with 1p36.3 microduplication syndrome, which is consistent with the clinical manifestations of two patients (Table 1). The patients’ genetic information was not applicable and phenotypes were, but consistent with that described in similar cases (score = 0.1), so the ACMG score was pathogenic (P). And we wanted to define the possible pathogenic gene.
Identifcation of variants in GNB1
In addition, we found that GNB1 may be a repeat intolerance gene (pTriplo score = 1.0); both LoF and DN effects have been described for patients carrying pathogenic GNB1 variants, which would be in line with the hypothesis that this gene has a strong contribution to the phenotypes presented for the siblings.

Discussion
We report a family in which two members carry the same 1p36.3 microduplication without any other chromosomal imbalance [1,2,3,4,5]. The first published case of familial pure 1p36.3 microduplication was reported by [6]; its additional particularity is that the duplicated segment is located at the end of an otherwise intact chromosome 9. The four members of the family harboring the duplication presented with global developmental delay, particular facial appearance including broad nasal bridge, bulbous nose and infraorbital creases, hypertelorism, and epilepsy for the mother and sister of the proband. We report the two siblings of familial 1p36.3 microduplication; considering that the variant was not detected in the blood samples of the patient’s parents, it is highly suspected that one of the parents was gonadal mosaicism.
The phenotypical spectrum of the 1p36.3 microduplication is quite similar to that of the 1p36 microdeletion syndrome, which is the most common terminal deletion in human beings. The hearing loss has been found to be affected in the patients of 1p36 deletion syndrome, but our duplication patients has no hearing deo hea.
Greco et al. review 34 scientific articles that described 315 patients with 1p36 deletion syndrome. It is difficult to delineate the age of seizure onset because most reports do not specify it. However, from the available data, it seems that epilepsy usually started in infancy, often during the first 6 months of life, but rarely in the neonatal period. Although seizure types are highly variable, infantile spasms are the most common seizure type detected [7]. However, in particular, many authors described the presence of seizure in 1p36 microduplication; it seems that epilepsy usually started after 3 years old [4,5,6]. The age of onset in our cases at 2–3 years. We may need more cases to clarify the age of onset in 1p36 microduplication.
In addition, among the 73 protein-coding genes, the GNB1 gene is considered to be associated with autosomal dominant mental retardation type 42. Germline de novo point mutations in the GNB1 gene have been associated with a new neurodevelopmental disorder named the GNB1 syndrome. It is a rare disease, with just over 80 known cases, first identified in 2016. Phenotypic features include global developmental delay (100%), hypotonia (78%), seizures (50%), ophthalmological anomalies (70%), absence of speech (60%), abnormal vision (58%), inability to walk independently (48%), growth delays (46%), speech delays (40%), autism (18%), dystonia (17%), and cardiovascular defects (11%) [8,9,10,11,12,13,14]. GNB1-related pathogenic mechanisms have been reported to include loss of function (LOF) and dominant negative (DN) effects. In vitro, D76G and I80T resulted in a loss of function, whereas K57E, K78E, and K89E exhibited a dominant negative effect, causing changes in baseline calcium signaling[14]. Studies of germline de novo mutations in GNB1 demonstrated a gain-of-function effect due to constitutive activation of G protein downstream signaling cascades for some of the affected residues [11]. However, Schultz-Rogers et al. [15] reported two patients with functionally confirmed loss of function variants in GNB1. Their results suggested haploinsufficiency of GNB1 was a mechanism for neurodevelopmental disorders in humans [15].
Protein kinase C(PKC)-zeta is an atypical member of the PKC family. Martin et al. found that the loss of Prkcz in mice impaired signaling through the B-cell receptor, resulting in inhibition of cell proliferation and survival, as well as defects in the activation of Erk and the transcription of NF-kappa-B-dependent genes. Furthermore, Prkcz-null mice were unable to mount an optimal T cell-dependent immune response [16].
In conclusion, we report a family in which two members carry the same 1p36.3 microduplication without any other chromosomal imbalance. It is highly suspected that one of the parents was gonadal mosaicism.
Data availability
The datasets used or analyzed during the current case reports are available from the corresponding author on reasonable request.
Change history
07 October 2023
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s10048-023-00735-7
References
Heilstedt HA, Shapira SK, Gregg AR, Shaffer LG (1999) Molecular and clinical characterization of a patient with duplication of 1p36.3 and metopic synostosis. Clin Genet 56(2):123–128. https://doi.org/10.1034/j.1399-0004.1999.560205.x
Giannikou K, Fryssira H, Oikonomakis V, Syrmou A, Kosma K, Tzetis M, Kitsiou-Tzeli S, Kanavakis E (2012) Further delineation of novel 1p36 rear rangements by array-cgh analysis: narrowing the breakpoints and clarifying the “extended” phenotype. Gene 506(2):360–368. https://doi.org/10.1016/j.gene.2012.06.060
Gajecka M, Yu W, Ballif BC, Glotzbach CD, Bailey KA, Shaw CA, Kashork CD, Heilstedt HA, Ansel DA, Theisen A, Rice R, Rice DP, Shaffer LG (2005) Delineation of mechanisms and regions of dosage imbalance in complex rearrangements of 1p36 leads to a putative gene for regulation of cranial suture closure. Eur J Hum Genet 13(2):139–149. https://doi.org/10.1038/sj.ejhg.5201302
Isidor B, Le Cunff M, Boceno M, Boisseau P, Thomas C, Rival JM, David A, Le Caignec C (2008) Complex constitutional subtelomeric 1p36.3 deletion/duplication in a mentally retarded child with neonatal neuroblastoma. Eur J Med Genet 51(6):679–684. https://doi.org/10.1016/j.ejmg.2008.06.004
Tonk VS, Wilson GN, Yatsenko SA, Stankiewicz P, Lupski JR, Schutt RC, Northup JK, Velagaleti GV (2005) Molecular cytogenetic characterization of a familial der(1)del(1)(p36.33)dup(1)(p36.33p36.22) with variable phenotype. Am J Med Genet A 139A(2):136–40. https://doi.org/10.1002/ajmg.a.30958
Marquet V, Bourthoumieu S, Dobrescu A, Laroche-Raynaud C, Yardin C (2017) Familial 1p36.3 microduplication resulting from a 1p–9q non-reciprocal translocation. Eur J Med Genet 60(11):583–588. https://doi.org/10.1016/j.ejmg.2017.08.009
Greco M, Ferrara P, Farello G, Striano P, Verrotti A (2018) Electroclinical features of epilepsy associated with 1p36 deletion syndrome: a review. Epilepsy Res 139:92–101. https://doi.org/10.1016/j.eplepsyres.2017.11.016
Brett M, Lai AH, Ting TW, Tan AM, Foo R, Jamuar S, Tan EC (2017) Acute lymphoblastic leukemia in a child with a de novo germline gnb1 mutation. Am J Med Genet A 173(2):550–552. https://doi.org/10.1002/ajmg.a.38026
Hemati P, Revah-Politi A, Bassan H, Petrovski S, Bilancia CG, Ramsey K, Griffin NG, Bier L, Cho MT, Rosello M, Lynch SA, Colombo S, Weber A, Haug M, Heinzen EL, Sands TT, Narayanan V, Primiano M, Aggarwal VS, Millan F, Sattler-Holtrop SG, Caro-Llopis A, Pillar N, Baker J, Freedman R, Kroes HY, Sacharow S, Stong N, Lapunzina P, Schneider MC, Mendelsohn NJ, Singleton A, Loik Ramey V, Wou K, Kuzminsky A, Monfort S, Weiss M, Doyle S, Iglesias A, Martinez F, Mckenzie F, Orellana C, van Gassen KLI, Palomares M, Bazak L, Lee A, Bircher A, Basel-Vanagaite L, Hafström M, Houge G, C4RCD Research Group, DDD study, Goldstein DB, Anyane-Yeboa K (2018) Refining the phenotype associated with GNB1 mutations: clinical data on 18 newly identified patients and review of the literature. Am J Med Genet A 176(11):2259–2275. https://doi.org/10.1002/ajmg.a.40472
Lohmann K, Masuho I, Patil DN, Baumann H, Hebert E, Steinrücke S, Trujillano D, Skamangas NK, Dobricic V, Hüning I, Gillessen-Kaesbach G, Westenberger A, Savic-Pavicevic D, Münchau A, Oprea G, Klein C, Rolfs A, Martemyanov KA (2017) Novel GNB1 mutations disrupt assembly and function of G protein heterotrimers and cause global developmental delay in humans. Hum Mol Genet 26(6):1078–1086. https://doi.org/10.1093/hmg/ddx018
Petrovski S, Küry S, Myers CT, Anyane-Yeboa K, Cogné B, Bialer M, Xia F, Hemati P, Riviello J, Mehaffey M, Besnard T, Becraft E, Wadley A, Politi AR, Colombo S, Zhu X, Ren Z, Andrews I, Dudding-Byth T, Schneider AL, Wallace G, University of Washington Center for Mendelian Genomics, Rosen ABI, Schelley S, Enns GM, Corre P, Dalton J, Mercier S, Latypova X, Schmitt S, Guzman E, Moore C, Bier L, Heinzen EL, Karachunski P, Shur N, Grebe T, Basinger A, Nguyen JM, Bézieau S, Wierenga K, Bernstein JA, Scheffer IE, Rosenfeld JA, Mefford HC, Isidor B, Goldstein DB (2016) Germline de novo mutations in GNB1 cause severe neurodevelopmental disability, hypotonia, and seizures. Am J Hum Genet 98(5):1001–1010. https://doi.org/10.1016/j.ajhg.2016.03.011
Steinrücke S, Lohmann K, Domingo A, Rolfs A, Bäumer T, Spiegler J, Hartmann C, Münchau A (2016) Novel GNB1 missense mutation in a patient with generalized dystonia, hypotonia, and intellectual disability. Neurol Genet 2(5):e106. https://doi.org/10.1212/NXG.0000000000000106
Szczałuba K, Biernacka A, Szymańska K, Gasperowicz P, Kosińska J, Rydzanicz M, Płoski R (2018) Novel GNB1 de novo mutation in a patient with neurodevelopmental disorder and cutaneous mastocytosis: Clinical report and literature review. Eur J Med Genet 61(3):157–160. https://doi.org/10.1016/j.ejmg.2017.11.010
Pirvulescu I (2019) Functional study of disease-causing GNB1[M]. McGill University Libraries, Canada
Schultz-Rogers L, Masuho I, Pinto E, Vairo F, Schmitz CT, Schwab TL, Clark KJ, Gunderson L, Pichurin PN, Wierenga K, Martemyanov KA, Klee EW (2020) Haploinsufficiency as a disease mechanism in GNB1-associated neurodevelopmental disorder. Mol Genet Genomic Med 8(11):e1477. https://doi.org/10.1002/mgg3.1477
Martin P, Duran A, Minguet S, Gaspar ML, Diaz-Meco MT, Rennert P, Leitges M, Moscat J (2002) Role of zeta PKC in B-cell signaling and function. EMBO J 21(15):4049–4057. https://doi.org/10.1093/emboj/cdf407
Acknowledgements
The authors are grateful to the two patients and their families.
Author information
Authors and Affiliations
Contributions
JPJ and YPW analyzed and interpreted data from the two patients. JPJ and YH conducted the literature research. CG collected clinical data of the patients. JPJ and SJT reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Institutional Review Board of the First Hospital of Hebei Medical University.
Consent for publication
We obtained verbal and written informed consent for the use of patient data. Images work for the publication of the cases for purely educational and research purposes, to which the patients and families agreed.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1007/s10048-023-00735-7
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiao, J., Wang, Y., Hou, Y. et al. RETRACTED ARTICLE: Clinical characterization of familial 1p36.3 microduplication. Neurogenetics 24, 201–208 (2023). https://doi.org/10.1007/s10048-023-00722-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-023-00722-y