Abstract
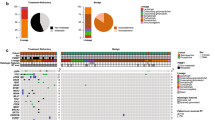
Oncogenic gene fusions have been reported in diffuse gliomas and may serve as potential therapeutic targets. Here, using next-generation sequencing analysis (Illumina TruSight Tumor 170 panel), we analyzed a total of 356 diffuse gliomas collected from 2017 to 2019 to evaluate clinical, pathological, and genetic features of gene fusion. We found 53 cases of glioblastomas harboring the following oncogenic gene fusions: MET (n = 18), EGFR (n = 14), FGFR (n = 12), NTRK (n = 5), RET (n = 2), AKT3 (n = 1), and PDGFRA fusions (n = 1). Gene fusions were consistently observed in both IDH-wildtype and IDH-mutant glioblastomas (8.8% and 9.4%, p = 1.000). PTPRZ1–MET fusion was the only fusion that genetically resembled secondary glioblastomas (i.e., high frequency of IDH mutation, ATRX loss, TP53 mutation, and absence of EGFR amplification), whereas other gene fusion types were similar to primary glioblastomas (i.e., high frequency of IDH-wildtype, TERT mutation, EGFR amplification, and PTEN mutation). In IDH-wildtype glioblastoma patients, multivariable analysis revealed that the PTPRZ1–MET fusion was associated with poor progression-free survival (HR [95% CI]: 5.42 (1.72–17.05), p = 0.004). Additionally, we described two novel cases of CCDC6–RET fusion in glioma. Collectively, our findings indicate that targetable gene fusions are associated with aggressive biological behavior and can aid the clinical treatment strategy for glioma patients.


Similar content being viewed by others
References
Gao Q, Liang WW, Foltz SM, Mutharasu G, Jayasinghe RG, Cao S, Liao WW, Reynolds SM, Wyczalkowski MA, Yao L, Yu L, Sun SQ, Chen K, Lazar AJ, Fields RC, Wendl MC, Van Tine BA, Vij R, Chen F, Nykter M, Shmulevich I, Ding L (2018) Driver fusions and their implications in the development and treatment of human cancers. Cell Rep 23(1):227–238.e223. https://doi.org/10.1016/j.celrep.2018.03.050
Mitelman F, Johansson B, Mertens F (2007) The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer 7(4):233–245. https://doi.org/10.1038/nrc2091
Mertens F, Johansson B, Fioretos T, Mitelman F (2015) The emerging complexity of gene fusions in cancer. Nat Rev Cancer 15(6):371–381. https://doi.org/10.1038/nrc3947
Yoshihara K, Wang Q, Torres-Garcia W, Zheng S, Vegesna R, Kim H, Verhaak RG (2015) The landscape and therapeutic relevance of cancer-associated transcript fusions. Oncogene 34(37):4845–4854. https://doi.org/10.1038/onc.2014.406
Lobato MN, Metzler M, Drynan L, Forster A, Pannell R, Rabbitts TH (2008) Modeling chromosomal translocations using conditional alleles to recapitulate initiating events in human leukemias. J Natl Cancer Inst Monogr 39:58–63. https://doi.org/10.1093/jncimonographs/lgn022
Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K, Hatanaka H, Bando M, Ohno S, Ishikawa Y, Aburatani H, Niki T, Sohara Y, Sugiyama Y, Mano H (2007) Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448(7153):561–566. https://doi.org/10.1038/nature05945
Druker BJ (2004) Imatinib as a paradigm of targeted therapies. Adv Cancer Res 91:1–30. https://doi.org/10.1016/s0065-230x(04)91001-9
Shaw AT, Yeap BY, Solomon BJ, Riely GJ, Gainor J, Engelman JA, Shapiro GI, Costa DB, Ou SH, Butaney M, Salgia R, Maki RG, Varella-Garcia M, Doebele RC, Bang YJ, Kulig K, Selaru P, Tang Y, Wilner KD, Kwak EL, Clark JW, Iafrate AJ, Camidge DR (2011) Effect of crizotinib on overall survival in patients with advanced non-small-cell lung cancer harbouring ALK gene rearrangement: a retrospective analysis. Lancet Oncol 12(11):1004–1012. https://doi.org/10.1016/s1470-2045(11)70232-7
Ostrom QT, Gittleman H, Farah P, Ondracek A, Chen Y, Wolinsky Y, Stroup NE, Kruchko C, Barnholtz-Sloan JS (2013) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2006–2010. Neuro Oncol 15(Suppl 2):ii1–56. https://doi.org/10.1093/neuonc/not151
(2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068. https://doi.org/10.1038/nature07385
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O'Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. https://doi.org/10.1016/j.cell.2013.09.034
Louis DNOH, Wiestler OD, Cavenee WK (2016) WHO classification of tumours of the central nervous system, revised. 4th edn. International Agency for Research on Cancer, Lyon
Touat M, Idbaih A, Sanson M, Ligon KL (2017) Glioblastoma targeted therapy: updated approaches from recent biological insights. Ann Oncol 28(7):1457–1472. https://doi.org/10.1093/annonc/mdx106
Aldape K, Zadeh G, Mansouri S, Reifenberger G, von Deimling A (2015) Glioblastoma: pathology, molecular mechanisms and markers. Acta Neuropathol 129(6):829–848. https://doi.org/10.1007/s00401-015-1432-1
Charest A, Lane K, McMahon K, Park J, Preisinger E, Conroy H, Housman D (2003) Fusion of FIG to the receptor tyrosine kinase ROS in a glioblastoma with an interstitial del(6)(q21q21). Genes Chromosomes Cancer 37(1):58–71. https://doi.org/10.1002/gcc.10207
Kim J, Lee Y, Cho HJ, Lee YE, An J, Cho GH, Ko YH, Joo KM, Nam DH (2014) NTRK1 fusion in glioblastoma multiforme. PLoS ONE 9(3):e91940. https://doi.org/10.1371/journal.pone.0091940
Ozawa T, Brennan CW, Wang L, Squatrito M, Sasayama T, Nakada M, Huse JT, Pedraza A, Utsuki S, Yasui Y, Tandon A, Fomchenko EI, Oka H, Levine RL, Fujii K, Ladanyi M, Holland EC (2010) PDGFRA gene rearrangements are frequent genetic events in PDGFRA-amplified glioblastomas. Genes Dev 24(19):2205–2218. https://doi.org/10.1101/gad.1972310
Parker BC, Annala MJ, Cogdell DE, Granberg KJ, Sun Y, Ji P, Li X, Gumin J, Zheng H, Hu L, Yli-Harja O, Haapasalo H, Visakorpi T, Liu X, Liu CG, Sawaya R, Fuller GN, Chen K, Lang FF, Nykter M, Zhang W (2013) The tumorigenic FGFR3-TACC3 gene fusion escapes miR-99a regulation in glioblastoma. J Clin Invest 123(2):855–865. https://doi.org/10.1172/jci67144
Singh D, Chan JM, Zoppoli P, Niola F, Sullivan R, Castano A, Liu EM, Reichel J, Porrati P, Pellegatta S, Qiu K, Gao Z, Ceccarelli M, Riccardi R, Brat DJ, Guha A, Aldape K, Golfinos JG, Zagzag D, Mikkelsen T, Finocchiaro G, Lasorella A, Rabadan R, Iavarone A (2012) Transforming fusions of FGFR and TACC genes in human glioblastoma. Science 337(6099):1231–1235. https://doi.org/10.1126/science.1220834
Fisher KE, Zhang L, Wang J, Smith GH, Newman S, Schneider TM, Pillai RN, Kudchadkar RR, Owonikoko TK, Ramalingam SS, Lawson DH, Delman KA, El-Rayes BF, Wilson MM, Sullivan HC, Morrison AS, Balci S, Adsay NV, Gal AA, Sica GL, Saxe DF, Mann KP, Hill CE, Khuri FR, Rossi MR (2016) Clinical validation and implementation of a targeted next-generation sequencing assay to detect somatic variants in non-small cell lung, melanoma, and gastrointestinal malignancies. J Mol Diagn 18(2):299–315. https://doi.org/10.1016/j.jmoldx.2015.11.006
Di Stefano AL, Fucci A, Frattini V, Labussiere M, Mokhtari K, Zoppoli P, Marie Y, Bruno A, Boisselier B, Giry M, Savatovsky J, Touat M, Belaid H, Kamoun A, Idbaih A, Houillier C, Luo FR, Soria J-C, Tabernero J, Eoli M, Paterra R, Yip S, Petrecca K, Chan JA, Finocchiaro G, Lasorella A, Sanson M, Iavarone A (2015) Detection, characterization, and inhibition of FGFR-TACC fusions in IDH wild-type glioma. Clin Cancer Res 21(14):3307–3317. https://doi.org/10.1158/1078-0432.CCR-14-2199
Heyer EE, Deveson IW, Wooi D, Selinger CI, Lyons RJ, Hayes VM, O'Toole SA, Ballinger ML, Gill D, Thomas DM, Mercer TR, Blackburn J (2019) Diagnosis of fusion genes using targeted RNA sequencing. Nat Commun 10(1):1388. https://doi.org/10.1038/s41467-019-09374-9
Ferguson SD, Zhou S, Huse JT, de Groot JF, Xiu J, Subramaniam DS, Mehta S, Gatalica Z, Swensen J, Sanai N, Spetzler D, Heimberger AB (2018) Targetable gene fusions associate with the IDH wild-type astrocytic lineage in adult gliomas. J Neuropathol Exp Neurol 77(6):437–442. https://doi.org/10.1093/jnen/nly022
Bao ZS, Chen HM, Yang MY, Zhang CB, Yu K, Ye WL, Hu BQ, Yan W, Zhang W, Akers J, Ramakrishnan V, Li J, Carter B, Liu YW, Hu HM, Wang Z, Li MY, Yao K, Qiu XG, Kang CS, You YP, Fan XL, Song WS, Li RQ, Su XD, Chen CC, Jiang T (2014) RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome Res 24(11):1765–1773. https://doi.org/10.1101/gr.165126.113
Hu H, Mu Q, Bao Z, Chen Y, Liu Y, Chen J, Wang K, Wang Z, Nam Y, Jiang B, Sa JK, Cho HJ, Her NG, Zhang C, Zhao Z, Zhang Y, Zeng F, Wu F, Kang X, Liu Y, Qian Z, Wang Z, Huang R, Wang Q, Zhang W, Qiu X, Li W, Nam DH, Fan X, Wang J, Jiang T (2018) Mutational landscape of secondary glioblastoma guides MET-targeted trial in brain tumor. Cell 175(6):1665–1678.e1618. https://doi.org/10.1016/j.cell.2018.09.038
Cheng F, Guo D (2019) MET in glioma: signaling pathways and targeted therapies. J Exp Clin Cancer Res 38(1):270. https://doi.org/10.1186/s13046-019-1269-x
Maire CL, Ligon KL (2014) Molecular pathologic diagnosis of epidermal growth factor receptor. Neuro Oncol 16(Suppl 8):viii1–6. https://doi.org/10.1093/neuonc/nou294
Frattini V, Trifonov V, Chan JM, Castano A, Lia M, Abate F, Keir ST, Ji AX, Zoppoli P, Niola F, Danussi C, Dolgalev I, Porrati P, Pellegatta S, Heguy A, Gupta G, Pisapia DJ, Canoll P, Bruce JN, McLendon RE, Yan H, Aldape K, Finocchiaro G, Mikkelsen T, Privé GG, Bigner DD, Lasorella A, Rabadan R, Iavarone A (2013) The integrated landscape of driver genomic alterations in glioblastoma. Nat Genet 45(10):1141–1149. https://doi.org/10.1038/ng.2734
Li AY, McCusker MG, Russo A, Scilla KA, Gittens A, Arensmeyer K, Mehra R, Adamo V, Rolfo C (2019) RET fusions in solid tumors. Cancer Treat Rev 81:101911. https://doi.org/10.1016/j.ctrv.2019.101911
Spina R, Voss DM, Asnaghi L, Sloan A, Bar EE (2016) Atracurium Besylate and other neuromuscular blocking agents promote astroglial differentiation and deplete glioblastoma stem cells. Oncotarget 7(1):459–472. https://doi.org/10.18632/ONCOTARGET.6314
Kumar-Sinha C, Kalyana-Sundaram S, Chinnaiyan AM (2015) Landscape of gene fusions in epithelial cancers: seq and ye shall find. Genome Med 7:129. https://doi.org/10.1186/s13073-015-0252-1
Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C (2014) The landscape of kinase fusions in cancer. Nat Commun 5(1):4846. https://doi.org/10.1038/ncomms5846
Davare MA, Tognon CE (2015) Detecting and targetting oncogenic fusion proteins in the genomic era. Biol Cell 107(5):111–129. https://doi.org/10.1111/boc.201400096
Acknowledgements
S.H.K. was supported by grants from the Brain Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science, ICT & Future Planning (Grant No. 2016M3C7A1913844). The funding source had no role in the design, practice, or analysis of this study. The authors would like to gratefully thank Won Young Park and Yi Rang Kim for their dedicated effort in NGS testing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Woo, H.Y., Na, K., Yoo, J. et al. Glioblastomas harboring gene fusions detected by next-generation sequencing. Brain Tumor Pathol 37, 136–144 (2020). https://doi.org/10.1007/s10014-020-00377-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-020-00377-9